Abstract
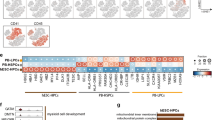
Understanding how differentiation programs originate from the gene-expression 'landscape' of hematopoietic stem cells (HSCs) is crucial for the development of new clinical therapies. We mapped the transcriptional dynamics underlying the first steps of commitment by tracking transcriptome changes in human HSCs and eight early progenitor populations. We found that transcriptional programs were extensively shared, extended across lineage-potential boundaries and were not strictly lineage affiliated. Elements of stem, lymphoid and myeloid programs were retained in multilymphoid progenitors (MLPs), which reflected a hybrid transcriptional state. By functional single cell analysis, we found that the transcription factors Bcl-11A, Sox4 and TEAD1 (TEF1) governed transcriptional networks in MLPs, which led to B cell specification. Overall, we found that integrated transcriptome approaches can be used to identify previously unknown regulators of multipotency and show additional complexity in lymphoid commitment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Orkin, S.H. & Zon, L.I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
Enver, T., Pera, M., Peterson, C. & Andrews, P.W. Stem cell states, fates, and the rules of attraction. Cell Stem Cell 4, 387–397 (2009).
Zandi, S., Bryder, D. & Sigvardsson, M. Load and lock: the molecular mechanisms of B-lymphocyte commitment. Immunol. Rev. 238, 47–62 (2010).
Nutt, S.L. & Kee, B.L. The transcriptional regulation of B cell lineage commitment. Immunity 26, 715–725 (2007).
Georgopoulos, K. Haematopoietic cell-fate decisions, chromatin regulation and ikaros. Nat. Rev. Immunol. 2, 162–174 (2002).
Ji, H. et al. Comprehensive methylome map of lineage commitment from haematopoietic progenitors. Nature 467, 338–342 (2010).
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011).
Chambers, S.M. et al. Hematopoietic fingerprints: an expression database of stem cells and their progeny. Cell Stem Cell 1, 578–591 (2007).
Månsson, R. et al. Molecular Evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity 26, 407–419 (2007).
Ng, S.Y.-M., Yoshida, T., Zhang, J. & Georgopoulos, K. Genome-wide lineage-specific transcriptional networks underscore Ikaros-dependent lymphoid priming in hematopoietic stem cells. Immunity 30, 493–507 (2009).
Hu, M. et al. Multilineage gene expression precedes commitment in the hemopoietic system. Genes Dev. 11, 774–785 (1997).
Akashi, K., Traver, D., Miyamoto, T. & Weissman, I.L. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature 404, 193–197 (2000).
Doulatov, S. et al. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat. Immunol. 11, 585–593 (2010).
Luc, S. et al. The earliest thymic T cell progenitors sustain B cell and myeloid lineage potential. Nat. Immunol. 13, 412–419 (2012).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005).
Kohn, L.A. et al. Lymphoid priming in human bone marrow begins before expression of CD10 with upregulation of L-selectin. Nat. Immunol. 13, 963–971 (2012).
Notta, F. et al. Isolation of single human hematopoietic stem cells capable of long-term multilineage engraftment. Science 333, 218–221 (2011).
Hao, Q.-L. et al. Human intrathymic lineage commitment is marked by differential CD7 expression: identification of CD7− lympho-myeloid thymic progenitors. Blood 111, 1318–1326 (2008).
Doulatov, S., Notta, F., Laurenti, E. & Dick, J.E. Hematopoiesis: a human perspective. Cell Stem Cell 10, 120–136 (2012).
Zhang, Q., Iida, R., Shimazu, T. & Kincade, P.W. Replenishing B lymphocytes in health and disease. Curr. Opin. Immunol. 24, 196–203 (2012).
Ernst, J., Nau, G.J. & Bar-Joseph, Z. Clustering short time series gene expression data. Bioinformatics 21, i159–i168 (2005).
Ernst, J., Vainas, O., Harbison, C.T., Simon, I. & Bar-Joseph, Z. Reconstructing dynamic regulatory maps. Mol. Syst. Biol. 3 (2007).
Itoh, K. et al. Reproducible establishment of hemopoietic supportive stromal cell lines from murine bone marrow. Exp. Hematol. 17, 145–153 (1989).
Rossi, M.I. et al. Relatively normal human lymphopoiesis but rapid turnover of newly formed B cells in transplanted nonobese diabetic/SCID mice. J. Immunol. 167, 3033–3042 (2001).
Lin, Y.C. et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat. Immunol. 11, 635–643 (2010).
Kawamoto, H., Ikawa, T., Masuda, K., Wada, H. & Katsura, Y. A map for lineage restriction of progenitors during hematopoiesis: the essence of the myeloid-based model. Immunol. Rev. 238, 23–36 (2010).
Zhang, J.A., Mortazavi, A., Williams, B.A., Wold, B.J. & Rothenberg, E.V. Dynamic transformations of genome-wide epigenetic marking and transcriptional control establish T cell identity. Cell 149, 467–482 (2012).
Santos, P.M. & Borghesi, L. Molecular resolution of the B cell landscape. Curr. Opin. Immunol. 23, 163–170 (2011).
Basso, K. & Dalla-Favera, R. Roles of BCL6 in normal and transformed germinal center B cells. Immunol. Rev. 247, 172–183 (2012).
Duy, C. et al. BCL6 is critical for the development of a diverse primary B cell repertoire. J. Exp. Med. 207, 1209–1221 (2010).
Schilham, M.W. et al. Defects in cardiac outflow tract formation and pro-B-lymphocyte expansion in mice lacking Sox-4. Nature 380, 711–714 (1996).
Liu, P. et al. Bcl11a is essential for normal lymphoid development. Nat. Immunol. 4, 525–532 (2003).
Yu, Y. et al. Bcl11a is essential for lymphoid development and negatively regulates p53. J. Exp. Med. 209, 2467–2483 (2012).
Zhao, B., Tumaneng, K. & Guan, K.-L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat. Cell Biol. 13, 877–883 (2011).
Jansson, L. & Larsson, J. Normal hematopoietic stem cell function in mice with enforced expression of the Hippo signaling effector YAP1. PLoS ONE 7, e32013 (2012).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012).
Fan, J.-B. et al. Highly parallel genome-wide expression analysis of single mammalian cells. PLoS ONE 7, e30794 (2012).
April, C. et al. Whole-genome gene expression profiling of formalin-fixed, paraffin-embedded tissue samples. PLoS ONE 4, e8162 (2009).
Yeung, K.Y., Haynor, D.R. & Ruzzo, W.L. Validating clustering for gene expression data. Bioinformatics 17, 309–318 (2001).
Kel, A.E. et al. MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 31, 3576–3579 (2003).
Chekmenev, D.S., Haid, C. & Kel, A.E. P-Match: transcription factor binding site search by combining patterns and weight matrices. Nucleic Acids Res. 33, W432–W437 (2005).
Ci, W. et al. The BCL6 transcriptional program features repression of multiple oncogenes in primary B cells and is deregulated in DLBCL. Blood 113, 5536–5548 (2009).
Basso, K. et al. Integrated biochemical and computational approach identifies BCL6 direct target genes controlling multiple pathways in normal germinal center B cells. Blood 115, 975–984 (2010).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Mazurier, F., Gan, O.I., McKenzie, J.L., Doedens, M. & Dick, J.E. Lentivector-mediated clonal tracking reveals intrinsic heterogeneity in the human hematopoietic stem cell compartment and culture-induced stem cell impairment. Blood 103, 545–552 (2004).
Acknowledgements
We thank the obstetrics unit of Trillium Hospital for cord blood samples; P.A. Penttilä, A. Khandi, L. Jamieson and S. Zhao at the SickKids Flow Cytometry Facility for cell sorting; V. Voisin and G. Bader for advice on bioinformatics and critical review of the manuscript; S. Bashir for statistical advice; A. Neumann for help with cloning and plasmids; M. Doedens for help with intrafemoral injections; and O. Gan for critical review of the manuscript. Supported by the Swiss National Science Foundation (E.L.), Roche (E.L.), the Fondation Suisse pour les Bourses en Médecine et Biologie (E.L.), the Swedish Research Council (S.Z.), Genome Canada (through the Ontario Genomics Institute), Ontario Institute for Cancer Research with funds from the province of Ontario, the Canadian Institutes for Health Research, Canada Research Chairs Program, the Princess Margaret Hospital Foundation, the Terry Fox Research Institute, Canadian Cancer Society Research Institute and the Ontario Ministry of Health and Long Term Care. The views expressed do not necessarily reflect those of the Ontario Ministry of Health and Long Term Care.
Author information
Authors and Affiliations
Contributions
E.L. designed and did experiments, analyzed data and wrote the manuscript; S.D. designed and did experiments, analyzed data and edited the manuscript; S.Z. designed and did experiments and edited the manuscript; I.P. cloned lentiviral vectors plasmids and did RT-PCR; J.C. and C.A. did gene-expression profiling experiments; J.-B.F. supervised gene-expression profile experiments, and J.E.D. supervised the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3 and 5 and Supplementary Note 1 (PDF 4384 kb)
Supplementary Table 4
Gene expression profiles determined with the STEM algorithm and K-means. (XLSX 1216 kb)
Supplementary Table 6
Transcription factors differentially expressed throughout the hierarchy. (XLSX 35 kb)
Supplementary Table 7
Gene-lists used for transcription factor binding motif enrichment. (XLSX 93 kb)
Supplementary Table 8
Data for MS5-MBN assay. (XLSX 72 kb)
Supplementary Table 9
Human chimerism and proportions of B and myeloid cells in all mice analyzed. (XLSX 63 kb)
Rights and permissions
About this article
Cite this article
Laurenti, E., Doulatov, S., Zandi, S. et al. The transcriptional architecture of early human hematopoiesis identifies multilevel control of lymphoid commitment. Nat Immunol 14, 756–763 (2013). https://doi.org/10.1038/ni.2615
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2615
This article is cited by
-
Single-cell RNA sequencing of a new transgenic t(8;21) preleukemia mouse model reveals regulatory networks promoting leukemic transformation
Leukemia (2024)
-
miR-126 identifies a quiescent and chemo-resistant human B-ALL cell subset that correlates with minimal residual disease
Leukemia (2023)
-
Differentiation Epitopes Define Hematopoietic Stem Cells and Change with Cell Cycle Passage
Stem Cell Reviews and Reports (2022)
-
Phenotypically-defined stages of leukemia arrest predict main driver mutations subgroups, and outcome in acute myeloid leukemia
Blood Cancer Journal (2022)
-
A transcriptomic continuum of differentiation arrest identifies myeloid interface acute leukemias with poor prognosis
Leukemia (2021)