Abstract
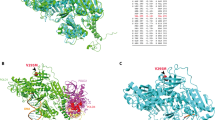
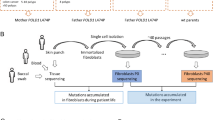
Many individuals with multiple or large colorectal adenomas or early-onset colorectal cancer (CRC) have no detectable germline mutations in the known cancer predisposition genes. Using whole-genome sequencing, supplemented by linkage and association analysis, we identified specific heterozygous POLE or POLD1 germline variants in several multiple-adenoma and/or CRC cases but in no controls. The variants associated with susceptibility, POLE p.Leu424Val and POLD1 p.Ser478Asn, have high penetrance, and POLD1 mutation was also associated with endometrial cancer predisposition. The mutations map to equivalent sites in the proofreading (exonuclease) domain of DNA polymerases ɛ and δ and are predicted to cause a defect in the correction of mispaired bases inserted during DNA replication. In agreement with this prediction, the tumors from mutation carriers were microsatellite stable but tended to acquire base substitution mutations, as confirmed by yeast functional assays. Further analysis of published data showed that the recently described group of hypermutant, microsatellite-stable CRCs is likely to be caused by somatic POLE mutations affecting the exonuclease domain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
08 May 2013
In the version of this article initially published, the name of author Estrella Guarino was incorrectly listed as Estrella Guarino Almeida. The error has been corrected in the HTML and PDF versions of this article.
References
Lunter, G. & Goodson, M. Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 21, 936–939 (2011).
Kemp, Z. et al. Evidence for a colorectal cancer susceptibility locus on chromosome 3q21-q24 from a high-density SNP genome-wide linkage scan. Hum. Mol. Genet. 15, 2903–2910 (2006).
Papaemmanuil, E. et al. Deciphering the genetics of hereditary non-syndromic colorectal cancer. Eur. J. Hum. Genet. 16, 1477–1486 (2008).
Laurent-Puig, P., Béroud, C. & Soussi, T. APC gene: database of germline and somatic mutations in human tumors and cell lines. Nucleic Acids Res. 26, 269–270 (1998).
Pursell, Z.F., Isoz, I., Lundstrom, E.B., Johansson, E. & Kunkel, T.A. Yeast DNA polymerase ɛ participates in leading-strand DNA replication. Science 317, 127–130 (2007).
Swan, M.K., Johnson, R.E., Prakash, L., Prakash, S. & Aggarwal, A.K. Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase δ. Nat. Struct. Mol. Biol. 16, 979–986 (2009).
Jin, Y.H., Ayyagari, R., Resnick, M.A., Gordenin, D.A. & Burgers, P.M. Okazaki fragment maturation in yeast. II. Cooperation between the polymerase and 3′-5′-exonuclease activities of Pol δ in the creation of a ligatable nick. J. Biol. Chem. 278, 1626–1633 (2003).
Murphy, K., Darmawan, H., Schultz, A., Fidalgo da Silva, E. & Reha-Krantz, L.J. A method to select for mutator DNA polymerase deltas in Saccharomyces cerevisiae. Genome 49, 403–410 (2006).
Burgers, P.M. Polymerase dynamics at the eukaryotic DNA replication fork. J. Biol. Chem. 284, 4041–4045 (2009).
Hindges, R. & Hubscher, U. DNA polymerase δ, an essential enzyme for DNA transactions. Biol. Chem. 378, 345–362 (1997).
Bellacosa, A. Functional interactions and signaling properties of mammalian DNA mismatch repair proteins. Cell Death Differ. 8, 1076–1092 (2001).
Mitra, S., Boldogh, I., Izumi, T. & Hazra, T.K. Complexities of the DNA base excision repair pathway for repair of oxidative DNA damage. Environ. Mol. Mutagen. 38, 180–190 (2001).
Fleck, O., Lehmann, E., Schar, P. & Kohli, J. Involvement of nucleotide-excision repair in msh2 pms1-independent mismatch repair. Nat. Genet. 21, 314–317 (1999).
da Costa, L.T. et al. Polymerase δ variants in RER colorectal tumours. Nat. Genet. 9, 10–11 (1995).
Flohr, T. et al. Detection of mutations in the DNA polymerase δ gene of human sporadic colorectal cancers and colon cancer cell lines. Int. J. Cancer 80, 919–929 (1999).
Yoshida, R. et al. Concurrent genetic alterations in DNA polymerase proofreading and mismatch repair in human colorectal cancer. Eur. J. Hum. Genet. 19, 320–325 (2011).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
Wang, L. et al. MYH mutations in patients with attenuated and classic polyposis and with young-onset colorectal cancer without polyps. Gastroenterology 127, 9–16 (2004).
Lanspa, S.J. et al. Colorectal adenomas in the Lynch syndromes. Results of a colonoscopy screening program. Gastroenterology 98, 1117–1122 (1990).
Wijnen, J. et al. Familial endometrial cancer in female carriers of MSH6 germline mutations. Nat. Genet. 23, 142–144 (1999).
Dunlop, M.G. et al. Common variation near CDKN1A, POLD3 and SHROOM2 influences colorectal cancer risk. Nat. Genet. 44, 770–776 (2012).
Albertson, T.M. et al. DNA polymerase ɛ and δ proofreading suppress discrete mutator and cancer phenotypes in mice. Proc. Natl. Acad. Sci. USA 106, 17101–17104 (2009).
Preston, B.D., Albertson, T.M. & Herr, A.J. DNA replication fidelity and cancer. Semin. Cancer Biol. 20, 281–293 (2010).
Goldsby, R.E. et al. Defective DNA polymerase-δ proofreading causes cancer susceptibility in mice. Nat. Med. 7, 638–639 (2001).
Goldsby, R.E. et al. High incidence of epithelial cancers in mice deficient for DNA polymerase δ proofreading. Proc. Natl. Acad. Sci. USA 99, 15560–15565 (2002).
Venkatesan, R.N., Hsu, J.J., Lawrence, N.A., Preston, B.D. & Loeb, L.A. Mutator phenotypes caused by substitution at a conserved motif A residue in eukaryotic DNA polymerase δ. J. Biol. Chem. 281, 4486–4494 (2006).
Venkatesan, R.N. et al. Mutation at the polymerase active site of mouse DNA polymerase δ increases genomic instability and accelerates tumorigenesis. Mol. Cell. Biol. 27, 7669–7682 (2007).
Jones, A.M. et al. Analysis of copy number changes suggests chromosomal instability in a minority of large colorectal adenomas. J. Pathol. 213, 249–256 (2007).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat. Genet. 39, 984–988 (2007).
Penegar, S. et al. National study of colorectal cancer genetics. Br. J. Cancer 97, 1305–1309 (2007).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Pendlebury, S., Duchesne, F., Reed, K.A., Smith, J.L. & Kerr, D.J. A trial of adjuvant therapy in colorectal cancer: the VICTOR trial. Clin. Colorectal Cancer 3, 58–60 (2003).
Power, C. & Elliott, J. Cohort profile: 1958 British birth cohort (National Child Development Study). Int. J. Epidemiol. 35, 34–41 (2006).
Ouwehand, W.H. Platelet genomics and the risk of atherothrombosis. J. Thromb. Haemost. 5 (suppl. 1), 188–195 (2007).
Spurdle, A.B. et al. Genome-wide association study identifies a common variant associated with risk of endometrial cancer. Nat. Genet. 43, 451–454 (2011).
Cuppen, E. Genotyping by Allele-Specific Amplification (KASPar). CSH Protoc. 2007, pdb.prot4841 (2007).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Bähler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951 (1998).
Marti, T.M., Mansour, A.A., Lehmann, E. & Fleck, O. Different frameshift mutation spectra in non-repetitive DNA of MutSα- and MutLα-deficient fission yeast cells. DNA Repair (Amst.) 2, 571–580 (2003).
Hall, B.M., Ma, C.X., Liang, P. & Singh, K.K. Fluctuation analysis CalculatOR: a web tool for the determination of mutation rate using Luria-Delbruck fluctuation analysis. Bioinformatics 25, 1564–1565 (2009).
Acknowledgements
We are grateful to all the affected individuals and their relatives in the families studied and to those who have provided their medical care. We acknowledge the help of many colleagues who are part of the teams who work on the CORGI study. We acknowledge the use of DNA from the British 1958 Birth Cohort collection. This work was principally funded by Cancer Research UK (C6199/A10417) and the Oxford NIHR Comprehensive Biomedical Research Centre (to I.T.). We also acknowledge core funding to the Wellcome Trust Centre for Human Genetics from the Wellcome Trust (090532/Z/09/Z). Work in the laboratory of R.S.H. is supported by funding from Cancer Research UK (C1298/A8362, supported by the Bobby Moore Fund). Work in the laboratory of S.E.K. was supported by project grants from Cancer Research UK and the John Fell Oxford University Press (OUP) Research Fund. We thank O. Fleck (University of Copenhagen) for fission yeast strains. I.S. is supported by a fellowship from the Junta de Extremadura, Spain (Consejería de Economía, Comercio e Innovación). R.S.H. and I.T. acknowledge funding from the European Union Seventh Framework Programme (FP7/207-2013) under grant 258236, FP7 collaborative project SYSCOL.
Author information
Authors and Affiliations
Consortia
Contributions
C. Palles, K.M.H., E.D., A.M.J., P.B., A.S., D.C., Z.K., S.L.S., C. Petridis, E.J.S., L.G.C.-C., Y.M., K.K., S.D., E.G., I.S. and S.H. performed laboratory experiments and/or analyzed the data. J.-B.C. analyzed whole-genome sequencing data, with assistance from M.B.K., and supervised other bioinformatics data analysis. J.T., S.E.K. and I.T. supervised laboratory experiments. G.M., P.D. and D.B. oversaw WGS500 analysis, and C.C.H. provided additional statistical advice. L.M., E.B., M.G., A.L., C. Petridis, R.R., E.J.S., D.J.K., S.C., H.J.W.T., R.S.H. and I.T. obtained samples. J.G. undertook structural analysis. R.S.H. and I.T. provided sequencing data and oversaw the study. I.T. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of contributing members is provided at the end of the manuscript
A list of contributing members is provided at the end of the manuscript
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–8 and Supplementary Note (PDF 1486 kb)
Rights and permissions
About this article
Cite this article
Palles, C., Cazier, JB., Howarth, K. et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat Genet 45, 136–144 (2013). https://doi.org/10.1038/ng.2503
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2503
This article is cited by
-
Curative resection via right hemicolectomy and regional lymph node dissection for colonic adenomatous polyposis of unknown etiology with adenocarcinomas localized in the right side of the colon: a case report
Surgical Case Reports (2024)
-
Founder pathogenic variants in colorectal neoplasia susceptibility genes in Ashkenazi Jews undergoing colonoscopy
BJC Reports (2024)
-
Heterogeneous expression of ARID1A in colorectal cancer indicates distinguish immune landscape and efficacy of immunotherapy
Discover Oncology (2024)
-
Multiple duodenal epithelial tumors in a patient with polymerase proofreading-associated polyposis in POLE variant
Clinical Journal of Gastroenterology (2024)
-
Human Autosomal Recessive DNA Polymerase Delta 3 Deficiency Presenting as Omenn Syndrome
Journal of Clinical Immunology (2024)