Abstract
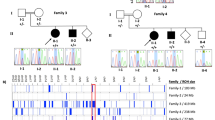
Deletions of the PAFAH1B1 gene (encoding LIS1) in 17p13.3 result in isolated lissencephaly sequence, and extended deletions including the YWHAE gene (encoding 14-3-3ε) cause Miller-Dieker syndrome. We identified seven unrelated individuals with submicroscopic duplication in 17p13.3 involving the PAFAH1B1 and/or YWHAE genes, and using a 'reverse genomics' approach, characterized the clinical consequences of these duplications. Increased PAFAH1B1 dosage causes mild brain structural abnormalities, moderate to severe developmental delay and failure to thrive. Duplication of YWHAE and surrounding genes increases the risk for macrosomia, mild developmental delay and pervasive developmental disorder, and results in shared facial dysmorphologies. Transgenic mice conditionally overexpressing LIS1 in the developing brain showed a decrease in brain size, an increase in apoptotic cells and a distorted cellular organization in the ventricular zone, including reduced cellular polarity but preserved cortical cell layer identity. Collectively, our results show that an increase in LIS1 expression in the developing brain results in brain abnormalities in mice and humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Lupski, J.R. Genomic rearrangements and sporadic disease. Nat. Genet. 39, S43–S47 (2007).
Reiner, O. et al. Isolation of a Miller-Dieker lissencephaly gene containing G protein beta-subunit-like repeats. Nature 364, 717–721 (1993).
Barkovich, A.J., Kuzniecky, R.I., Jackson, G.D., Guerrini, R. & Dobyns, W.B. A developmental and genetic classification for malformations of cortical development. Neurology 65, 1873–1887 (2005).
Harding, B. in Dysplasias of Cerebral Cortex and Epilepsy (ed. Guerrini, R.) 81–88 (Lippincott-Raven, Philadelphia, 1996).
Kamiya, A. et al. A schizophrenia-associated mutation of Drosoph. Inf. Serv.C1 perturbs cerebral cortex development. Nat. Cell Biol. 7, 1167–1178 (2005).
Schumacher, J. et al. Strong genetic evidence of DCDC2 as a susceptibility gene for dyslexia. Am. J. Hum. Genet. 78, 52–62 (2006).
Walsh, T. et al. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 320, 539–543 (2008).
Xu, B. et al. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 40, 880–885 (2008).
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008).
International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 455, 237–241 (2008).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Weiss, L.A. et al. Association between microdeletion and microduplication at 16p11.2 and autism. N. Engl. J. Med. 358, 667–675 (2008).
Kumar, R.A. et al. Recurrent 16p11.2 microdeletions in autism. Hum. Mol. Genet. 17, 628–638 (2008).
Cardoso, C. et al. Refinement of a 400-kb critical region allows genotypic differentiation between isolated lissencephaly, Miller-Dieker syndrome, and other phenotypes secondary to deletions of 17p13.3. Am. J. Hum. Genet. 72, 918–930 (2003).
Toyo-oka, K. et al. 14–3-3ε is important for neuronal migration by binding to NUDEL: a molecular explanation for Miller-Dieker syndrome. Nat. Genet. 34, 274–285 (2003).
Mikhail, F.M. et al. Complete trisomy 17p syndrome in a girl with der(14)t(14;17)(p11.2;p11.2). Am. J. Med. Genet. A. 140, 1647–1654 (2006).
Morelli, S.H., Deubler, D.A., Brothman, L.J., Carey, J.C. & Brothman, A.R. Partial trisomy 17p detected by spectral karyotyping. Clin. Genet. 55, 372–375 (1999).
Cahana, A. et al. Targeted mutagenesis of Lis1 disrupts cortical development and LIS1 homodimerization. Proc. Natl. Acad. Sci. USA 98, 6429–6434 (2001).
Hirotsune, S. et al. Graded reduction of Pafah1b1 (Lis1) activity results in neuronal migration defects and early embryonic lethality. Nat. Genet. 19, 333–339 (1998).
Shu, T. et al. Ndel1 operates in a common pathway with LIS1 and cytoplasmic dynein to regulate cortical neuronal positioning. Neuron 44, 263–277 (2004).
Tsai, J.W., Bremner, K.H. & Vallee, R.B. Dual subcellular roles for LIS1 and dynein in radial neuronal migration in live brain tissue. Nat. Neurosci. 10, 970–979 (2007).
Tsai, J.W., Chen, Y., Kriegstein, A.R. & Vallee, R.B. LIS1 RNA interference blocks neural stem cell division, morphogenesis, and motility at multiple stages. J. Cell Biol. 170, 935–945 (2005).
Peiffer, D.A. et al. High-resolution genomic profiling of chromosomal aberrations using Infinium whole-genome genotyping. Genome Res. 16, 1136–1148 (2006).
Lee, J.A., Carvalho, C.M. & Lupski, J.R.A. DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell 131, 1235–1247 (2007).
Dobyns, W.B., Reiner, O., Carrozzo, R. & Ledbetter, D.H. Lissencephaly. A human brain malformation associated with deletion of the LIS1 gene located at chromosome 17p13. J. Am. Med. Assoc. 270, 2838–2842 (1993).
Pilz, D.T. et al. LIS1 and XLIS (DCX) mutations cause most classical lissencephaly, but different patterns of malformation. Hum. Mol. Genet. 7, 2029–2037 (1998).
Chenn, A., Zhang, Y.A., Chang, B.T. & McConnell, S.K. Intrinsic polarity of mammalian neuroepithelial cells. Mol. Cell. Neurosci. 11, 183–193 (1998).
Tamamaki, N. et al. Green fluorescent protein expression and colocalization with calretinin, parvalbumin, and somatostatin in the GAD67-GFP knock-in mouse. J. Comp. Neurol. 467, 60–79 (2003).
Lupski, J.R. Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 14, 417–422 (1998).
Feller, S.M. Crk family adaptors-signalling complex formation and biological roles. Oncogene 20, 6348–6371 (2001).
Assadi, A.H. et al. Interaction of reelin signaling and Lis1 in brain development. Nat. Genet. 35, 270–276 (2003).
Ballif, B.A. et al. Activation of a Dab1/CrkL/C3G/Rap1 pathway in Reelin-stimulated neurons. Curr. Biol. 14, 606–610 (2004).
Chen, K. et al. Interaction between Dab1 and CrkII is promoted by Reelin signaling. J. Cell Sci. 117, 4527–4536 (2004).
Wall, M.A., Socolich, M. & Ranganathan, R. The structural basis for red fluorescence in the tetrameric GFP homolog DsRed. Nat. Struct. Biol. 7, 1133–1138 (2000).
Ligon, L.A., Karki, S., Tokito, M. & Holzbaur, E.L. Dynein binds to β-catenin and may tether microtubules at adherens junctions. Nat. Cell Biol. 3, 913–917 (2001).
Yingling, J. et al. Neuroepithelial stem cell proliferation requires LIS1 for precise spindle orientation and symmetric division. Cell 132, 474–486 (2008).
Hirokawa, N. & Takemura, R. Molecular motors in neuronal development, intracellular transport and diseases. Curr. Opin. Neurobiol. 14, 564–573 (2004).
Reiner, O., Sapoznik, S. & Sapir, T. Lissencephaly 1 linking to multiple diseases: mental retardation, neurodegeneration, schizophrenia, male sterility, and more. Neuromolecular Med. 8, 547–565 (2006).
Cappello, S. et al. The Rho-GTPase cdc42 regulates neural progenitor fate at the apical surface. Nat. Neurosci. 9, 1099–1107 (2006).
Chen, L. et al. Cdc42 deficiency causes Sonic hedgehog-independent holoprosencephaly. Proc. Natl. Acad. Sci. USA 103, 16520–16525 (2006).
Kholmanskikh, S.S., Dobrin, J.S., Wynshaw-Boris, A., Letourneau, P.C. & Ross, M.E. Disregulated RhoGTPases and actin cytoskeleton contribute to the migration defect in Lis1-deficient neurons. J. Neurosci. 23, 8673–8681 (2003).
Kholmanskikh, S.S. et al. Calcium-dependent interaction of Lis1 with IQGAP1 and Cdc42 promotes neuronal motility. Nat. Neurosci. 9, 50–57 (2006).
Shen, Y. et al. Nudel binds Cdc42GAP to modulate Cdc42 activity at the leading edge of migrating cells. Dev. Cell 14, 342–353 (2008).
Cheung, S.W. et al. Development and validation of a CGH microarray for clinical cytogenetic diagnosis. Genet. Med. 7, 422–432 (2005).
Lu, X. et al. Clinical implementation of chromosomal microarray analysis: summary of 2513 postnatal cases. PLoS ONE 2, e327 (2007).
Ou, Z. et al. BAC-emulation oligonucleotide arrays for targeted clinical array CGH analyses. Genet. Med. 10, 278–289 (2008).
Lobe, C.G. et al. Z/AP, a double reporter for cre-mediated recombination. Dev. Biol. 208, 281–292 (1999).
Coquelle, F.M. et al. LIS1, CLIP-170's key to the dynein/dynactin pathway. Mol. Cell. Biol. 22, 3089–3102 (2002).
Hebert, J.M. & McConnell, S.K. Targeting of cre to the Foxg1 (BF-1) locus mediates loxP recombination in the telencephalon and other developing head structures. Dev. Biol. 222, 296–306 (2000).
Benard, V., Bohl, B.P. & Bokoch, G.M. Characterization of rac and cdc42 activation in chemoattractant-stimulated human neutrophils using a novel assay for active GTPases. J. Biol. Chem. 274, 13198–13204 (1999).
Acknowledgements
We thank the participating families for their cooperation in the study, the members of the Chromosomal Microarray Analysis and Cytogenetic/FISH laboratories for technical assistance, G. Eichele for help with the in situ hybridization experiments, E. Arama and S. Haiderleu for useful comments and advice, S. McConnell for the Foxg1(Cre) mice and M. O'Gorman (Children's Memorial Hospital, Chicago) for assistance with specimen collection. The work was supported in part by the Israeli Science Foundation (grant no. 270/04 to O.R. and an equipment grant), the Foundation Jérôme Lejeune, the Minerva Foundation with funding from the Federal German Ministry for Education and Research, German-Israeli collaboration grant Gr-1905, March of Dimes grant 6-FY07-388, collaborative BSF grant 2007081 (to O.R. and J.R.L.), a grant from the Paul Godfrey Research Foundation in Children's Diseases, the Benoziyo Center for Neurological Diseases, the Kekst Center, the Forcheimer Center, a Weizmann-Pasteur collaborative grant, a research grant from the Michigan Women of Wisdom Fund to support Weizmann Women scientists, support from Maurice Janin, the Jewish Communal Fund, Albert Einstein College of Medicine of Yeshiva University, the David and Fela Shapell Family Center research grant for Genetic Disorders Research, grants DIGESIC-MEC BFU2005-09085 and Ingenio 2010 MEC-CONSOLIDER CSD2007-00023 (to S.M.), support from EU grant LSHG-CT-2004-512003, the Baylor Medical Genetics Laboratories, the Mental Retardation Developmental Disabilities Research Center (HD024064) and a Program Project grant (P01 HD39420) from the National Institute of Child Health and Human Development (to J.R.L.). O.R. is an Incumbent of the Bernstein-Mason professorial chair of Neurochemistry.
Author information
Authors and Affiliations
Contributions
W.B. coordinated human studies and conducted real time RT-PCR assays. T.S. produced transgenic mice and conducted mouse studies. O.A.S. recruited patients and reviewed clinical data. F.Z. conducted high-density array CGH and breakpoint analyses. M.A.W. carried out cell culture. J.V.H. reviewed the MRI data. T.L., V.S. and S.M. assisted in mouse analyses. Y.Y. provided GAD67-GFP mice. D.A.P. and K.L.G. conducted SNP genotyping. M.M.N., V.A.S., S.S.A., S.K.S., D.J.H., D.-L.D.-S., M.H. and A.L.B. recruited and clinically characterized patients. S.W.C., X.-Y.L. and T.S. were involved in cytogenetic and clinical array CGH studies. J.R.L. and O.R. were involved in research design and data analyses. W.B., T.S., O.A.S., O.R. and J.R.L. prepared the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Methods, Supplementary Table 1 and Supplementary Figures 1–5 (PDF 760 kb)
Supplementary Movie 1
Organotypic slice cultures prepared from brains of E13.5 control mice carrying a silent transgene (Cre negative). (MOV 1461 kb)
Supplementary Movie 2
Organotypic slice cultures prepared from brains of E13.5 LIS1 overexpressing embryos (LIS1::Foxg1(cre)). (MOV 1627 kb)
Rights and permissions
About this article
Cite this article
Bi, W., Sapir, T., Shchelochkov, O. et al. Increased LIS1 expression affects human and mouse brain development. Nat Genet 41, 168–177 (2009). https://doi.org/10.1038/ng.302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.302
This article is cited by
-
Clinical findings and genetic analysis of patients with copy number variants involving 17p13.3 using a single nucleotide polymorphism array: a single-center experience
BMC Medical Genomics (2022)
-
A new case of 17p13.3p13.1 microduplication resulted from unbalanced translocation: clinical and molecular cytogenetic characterization
Molecular Cytogenetics (2021)
-
Neuronal migration genes and a familial translocation t (3;17): candidate genes implicated in the phenotype
BMC Medical Genetics (2020)
-
Split hand/foot malformation with long bone deficiency associated with BHLHA9 gene duplication: a case report and review of literature
BMC Medical Genetics (2019)
-
Modeling the quantitative nature of neurodevelopmental disorders using Collaborative Cross mice
Molecular Autism (2018)