Abstract
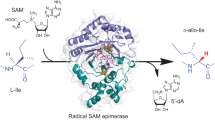
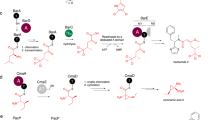
The radical S-adenosylmethionine (S-AdoMet) superfamily contains thousands of proteins that catalyze highly diverse conversions, most of which are poorly understood, owing to a lack of information regarding chemical products and radical-dependent transformations. We here report that NosL, involved in forming the indole side ring of the thiopeptide nosiheptide (NOS), is a radical S-AdoMet 3-methyl-2-indolic acid (MIA) synthase. NosL catalyzed an unprecedented carbon chain reconstitution of L-tryptophan to give MIA, showing removal of the Cα-N unit and shift of the carboxylate to the indole ring. Dissection of the enzymatic process upon the identification of products and a putative glycyl intermediate uncovered a radical-mediated, unusual fragmentation-recombination reaction. This finding unveiled a key step in radical S-AdoMet enzyme–catalyzed structural rearrangements during complex biotransformations. Additionally, NosL tolerated fluorinated L-tryptophan as the substrate, allowing for production of a regiospecifically halogenated thiopeptide that has not been found among the more than 80 members of the naturally occurring thiopeptide family.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bagley, M.C., Dale, J.W., Merritt, E.A. & Xiong, X. Thiopeptide antibiotics. Chem. Rev. 105, 685–714 (2005).
Arndt, H.-D., Schoof, S. & Lu, J.-Y. Thiopeptide antibiotic biosynthesis. Angew. Chem. Int. Ed. Engl. 48, 6770–6773 (2009).
Li, C. & Kelly, W.L. Recent advances in thiopeptide antibiotic biosynthesis. Nat. Prod. Rep. 27, 153–164 (2010).
Nicolaou, K.C., Chen, J.S., Edmonds, D.J. & Estrada, A.A. Recent advances in the chemistry and biology of naturally occurring antibiotics. Angew. Chem. Int. Ed. Engl. 48, 660–719 (2009).
Wieland Brown, L.C., Acker, M.G., Clardy, J., Walsh, C.T. & Fischbach, M.A. Thirteen posttranslational modifications convert a 14-residue peptide into the antibiotic thiocillin. Proc. Natl. Acad. Sci. USA 106, 2549–2553 (2009).
Liao, R. et al. Thiopeptide biosynthesis featuring ribosomally synthesized precursor peptides and conserved posttranslational modifications. Chem. Biol. 16, 141–147 (2009).
Kelly, W.L., Pan, L. & Li, C. Thiostrepton biosynthesis: prototype for a new family of bacteriocins. J. Am. Chem. Soc. 131, 4327–4334 (2009).
Morris, R.P. et al. Ribosomally synthesized thiopeptide antibiotics targeting elongation factor Tu. J. Am. Chem. Soc. 131, 5946–5955 (2009).
Yu, Y. et al. Nosiheptide biosynthesis featuring a unique indole side ring formation on the characteristic thiopeptide framework. ACS Chem. Biol. 4, 855–864 (2009).
Wang, J. et al. Identification and analysis of the biosynthetic gene cluster encoding the thiopeptide antibiotic cyclothiazomycin in Streptomyces hygroscopicus 10–22. Appl. Environ. Microbiol. 76, 2335–2344 (2010).
Ding, Y. et al. Moving posttranslational modifications forward to biosynthesize the glycosylated thiopeptide nocathiacin I in Nocardia sp. ATCC202099. Mol. Biosyst. 6, 1180–1185 (2010).
Houck, D.R., Chen, L.-C., Keller, P.J., Beale, J.M. & Floss, H.G. Biosynthesis of the modified peptide antibiotic nosiheptide in Streptomyces actuosus. J. Am. Chem. Soc. 110, 5800–5806 (1988).
Mocek, U. et al. Biosynthesis of the modified peptide antibiotic nosiheptide in Streptomyces actuosus. J. Am. Chem. Soc. 115, 7557–7568 (1993).
Mocek, U. et al. Biosynthesis of the modified peptide antibiotic thiostrepton in Streptomyces azureus and Streptomyces laurentii. J. Am. Chem. Soc. 115, 7992–8001 (1993).
Smith, T.M., Priestley, N.D., Knaggs, A.R., Nguyen, T. & Floss, H.G. 3,4-Dimethylindole-2-carboxylate and 4-(1-Hydroxyethyl)-2-quinolinecarboxylate activating enzymes from the nosiheptide and thiostrepton producers, Streptomyces actuosus and Streptomyces laurentii. J. Chem. Soc. Chem. Commun. 21, 1612–1614 (1993).
Priestley, N.D. et al. Studies on the biosynthesis of thiostrepton: 4-(1-hydroxyethyl)-quinoline-2-carboxylate as a free intermediate on the pathway to the quinaldic acid moiety. Bioorg. Med. Chem. 4, 1135–1147 (1996).
Frey, P.A., Hegeman, A.D. & Ruzika, F.J. The radical SAM superfamily. Crit. Rev. Biochem. Mol. Biol. 43, 63–88 (2008).
Chih, H.W. & Marsh, E.N.G. Pre-steady-state kinetic investigation of intermediates in the reaction catalyzed by adenosylcobalamin-dependent glutamate mutase. Biochemistry 38, 13684–13691 (1999).
Chih, H.-W. & March, N.G. Mechanism of glutamate mutase: Identification and kinetic competence of acrylate and glycyl radical as intermediates in the rearrangment of glutamate to methylaspartate. J. Am. Chem. Soc. 122, 10732–10733 (2000).
Banerjee, R. Radical carbon skeleton rearrangements: catalysis by coenzyme B12-dependent mutases. Chem. Rev. 103, 2083–2094 (2003).
Sofia, H.J., Chen, G., Hetzler, B.G., Reyes-Spindola, J.F. & Miller, N.E. Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res. 29, 1097–1106 (2001).
Nicolet, Y. & Drennan, C.L. AdoMet radical proteins–from structure to evolution–alignment of divergent protein sequences reveals strong secondary structure element conservation. Nucleic Acids Res. 32, 4015–4025 (2004).
Wang, S.C. & Frey, P.A. S-adenosylmethionine as an oxidant: the radical SAM superfamily. Trends Biochem. Sci. 32, 101–110 (2007).
Marsh, E.N.G., Patterson, D.P. & Li, L. Adenosyl radical: reagent and catalyst in enzyme reactions. ChemBioChem 11, 604–621 (2010).
Bonifačić, M., Stefanic, I., Hug, G.I., Armstrong, D.A. & Asmus, K.-D. Glycine decarboxylation: the free radical mechanism. J. Am. Chem. Soc. 120, 9930–9940 (1998).
Kriek, M., Martins, F., Challand, M.R., Croft, A. & Roach, P.L. Thiamine biosynthesis in Escherichia coli: identification of the intermediate and by-product derived from tyrosine. Angew. Chem. Int. Ed. Engl. 46, 9223–9226 (2007).
Driesener, R.C. et al. [FeFe]-hydrogenase cyanide ligands derived from S-adenosylmethionine-dependent cleavage of tyrosine. Angew. Chem. Int. Ed. Engl. 49, 1687–1690 (2010).
Chatterjee, A. et al. Reconstitution of ThiC in thiamine pyrimidine biosynthesis expands the radical SAM superfamily. Nat. Chem. Biol. 4, 758–765 (2008).
Acker, M.G., Bowers, A.A. & Walsh, C.T. Generation of thiocillin variants by prepeptide gene replacement and in vivo processing by Bacillus cereus. J. Am. Chem. Soc. 131, 17563–17565 (2009).
Bowers, A.A., Acker, M.G., Koglin, A. & Walsh, C.T. Thiazolyl peptide antibiotic biosynthesis: a cascade of posttranslational modifications on ribosomal nascent proteins. J. Am. Chem. Soc. 132, 7519–7527 (2010).
Kriek, M. et al. Thiazole synthase from Escherichia coli - an investigation of the substrates and purified proteins required for activity in vitro. J. Biol. Chem. 282, 17413–17423 (2007).
Acknowledgements
We thank H.G. Floss (University of Washington) for providing S. actuosus ATCC25421 and for his pioneering work on NOS biosynthesis and Y. Zhang and W. Tong, (High Magnetic Field Laboratory, Chinese Academy of Sciences) for assistance with EPR analysis. This work was supported in part by grants from US National Institutes of Health (CA094426 to B.S.), Chinese National Natural Science Foundation (20832009, 30525001, 90713012 and 20921091), Chinese Ministry of Science and Technology (2009ZX09501-008), Chinese National Basic Research Program (“973 program,” 2010CB833200), Chinese Academy of Sciences (KJCX2-YW-H08 and KSCX2-YW-G-06) and Science and Technology Commission of Shanghai Municipality (09QH1402700) of China (all to W.L).
Author information
Authors and Affiliations
Contributions
Q.Z., D.C., Y.Y. and L.D. carried out the experiments; Y.L. performed the theoretical calculations; Q.Z., B.S. and W.L. analyzed the data and wrote the paper; and W.L. designed the research. All authors discussed results and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–19 and Supplementary Tables 1–5 (PDF 2161 kb)
Rights and permissions
About this article
Cite this article
Zhang, Q., Li, Y., Chen, D. et al. Radical-mediated enzymatic carbon chain fragmentation-recombination. Nat Chem Biol 7, 154–160 (2011). https://doi.org/10.1038/nchembio.512
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.512
This article is cited by
-
L-tyrosine-bound ThiH structure reveals C–C bond break differences within radical SAM aromatic amino acid lyases
Nature Communications (2022)
-
Structure–function relationships of radical SAM enzymes
Nature Catalysis (2020)
-
Radical-mediated C-S bond cleavage in C2 sulfonate degradation by anaerobic bacteria
Nature Communications (2019)
-
The hidden enzymology of bacterial natural product biosynthesis
Nature Reviews Chemistry (2019)
-
Biosynthesis of the nosiheptide indole side ring centers on a cryptic carrier protein NosJ
Nature Communications (2017)