Abstract
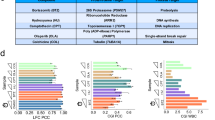
Bioactive compounds are widely used to modulate protein function and can serve as important leads for drug development. Identifying the in vivo targets of these compounds remains a challenge. Using yeast, we integrated three genome-wide gene-dosage assays to measure the effect of small molecules in vivo. A single TAG microarray was used to resolve the fitness of strains derived from pools of (i) homozygous deletion mutants, (ii) heterozygous deletion mutants and (iii) genomic library transformants. We demonstrated, with eight diverse reference compounds, that integration of these three chemogenomic profiles improves the sensitivity and specificity of small-molecule target identification. We further dissected the mechanism of action of two protein phosphatase inhibitors and in the process developed a framework for the rational design of multidrug combinations to sensitize cells with specific genotypes more effectively. Finally, we applied this platform to 188 novel synthetic chemical compounds and identified both potential targets and structure-activity relationships.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
17 September 2008
In the version of this article initially published, there was a space missing in the author name Robert P St Onge. The error has been corrected in the HTML and PDF versions of the article.
References
Giaever, G. et al. Chemogenomic profiling: identifying the functional interactions of small molecules in yeast. Proc. Natl. Acad. Sci. USA 101, 793–798 (2004).
Lum, P.Y. et al. Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes. Cell 116, 121–137 (2004).
Lee, W. et al. Genome-wide requirements for resistance to functionally distinct DNA-damaging agents. PLoS Genet. 1, 235–246 (2005).
Parsons, A.B. et al. Exploring the mode-of-action of bioactive compounds by chemical-genetic profiling in yeast. Cell 126, 611–625 (2006).
Rine, J., Hansen, W., Hardeman, E. & Davis, R.W. Targeted selection of recombinant clones through gene dosage effects. Proc. Natl. Acad. Sci. USA 80, 6750–6754 (1983).
Luesch, H. et al. A genome-wide overexpression screen in yeast for small-molecule target identification. Chem. Biol. 12, 55–63 (2005).
Butcher, R.A. et al. Microarray-based method for monitoring yeast overexpression strains reveals small-molecule targets in TOR pathway. Nat. Chem. Biol. 2, 103–109 (2006).
Parsons, A.B. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat. Biotechnol. 22, 62–69 (2004).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Kung, C. et al. Chemical genomic profiling to identify intracellular targets of a multiplex kinase inhibitor. Proc. Natl. Acad. Sci. USA 102, 3587–3592 (2005).
Lamb, J. et al. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Pierce, S.E. et al. A unique and universal molecular barcode array. Nat. Methods 3, 601–603 (2006).
Pierce, S.E., Davis, R.W., Nislow, C. & Giaever, G. Genome-wide analysis of barcoded Saccharomyces cerevisiae gene-deletion mutants in pooled cultures. Nat. Protoc. 2, 2958–2974 (2007).
Brenner, C. Chemical genomics in yeast. Genome Biol. 5, 240 (2004).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Li, X. et al. Multicopy suppressors for novel antibacterial compounds reveal targets and drug efflux susceptibility. Chem. Biol. 11, 1423–1430 (2004).
Breitling, R., Armengaud, P. & Amtmann, A. Vector analysis as a fast and easy method to compare gene expression responses between different experimental backgrounds. BMC Bioinformatics 6, 181 (2005).
Myers, C.E., Lippman, M.E., Elliot, H.M. & Chabner, B.A. Competitive protein binding assay for methotrexate. Proc. Natl. Acad. Sci. USA 72, 3683–3686 (1975).
Kontoyiannis, D.P., Sagar, N. & Hirschi, K.D. Overexpression of Erg11p by the regulatable GAL1 promoter confers fluconazole resistance in Saccharomyces cerevisiae . Antimicrob. Agents Chemother. 43, 2798–2800 (1999).
Zheng, X.F., Florentino, D., Chen, J., Crabtree, G.R. & Schreiber, S.L. TOR kinase domains are required for two distinct functions, only one of which is inhibited by rapamycin. Cell 82, 121–130 (1995).
Koltin, Y. et al. Rapamycin sensitivity in Saccharomyces cerevisiae is mediated by a peptidyl-prolyl cis-trans isomerase related to human FK506-binding protein. Mol. Cell. Biol. 11, 1718–1723 (1991).
Honkanen, R.E. Cantharidin, another natural toxin that inhibits the activity of serine/threonine protein phosphatases types 1 and 2A. FEBS Lett. 330, 283–286 (1993).
Li, Y.M. & Casida, J.E. Cantharidin-binding protein: identification as protein phosphatase 2A. Proc. Natl. Acad. Sci. USA 89, 11867–11870 (1992).
Ishihara, H. et al. Calyculin A and okadaic acid: inhibitors of protein phosphatase activity. Biochem. Biophys. Res. Commun. 159, 871–877 (1989).
Fujiki, H. & Suganuma, M. Tumor promotion by inhibitors of protein phosphatases 1 and 2A: the okadaic acid class of compounds. Adv. Cancer Res. 61, 143–194 (1993).
MacKintosh, C., Beattie, K.A., Klumpp, S., Cohen, P. & Codd, G.A. Cyanobacterial microcystin-LR is a potent and specific inhibitor of protein phosphatases 1 and 2A from both mammals and higher plants. FEBS Lett. 264, 187–192 (1990).
Walsh, E.P., Lamont, D.J., Beattie, K.A. & Stark, M.J. Novel interactions of Saccharomyces cerevisiae type 1 protein phosphatase identified by single-step affinity purification and mass spectrometry. Biochemistry 41, 2409–2420 (2002).
Krogan, N.J. et al. High-definition macromolecular composition of yeast RNA-processing complexes. Mol. Cell 13, 225–239 (2004).
Andrews, P.D. & Stark, M.J. Type 1 protein phosphatase is required for maintenance of cell wall integrity, morphogenesis and cell cycle progression in Saccharomyces cerevisiae . J. Cell Sci. 113, 507–520 (2000).
Baker, S.H., Frederick, D.L., Bloecher, A. & Tatchell, K. Alanine-scanning mutagenesis of protein phosphatase type 1 in the yeast Saccharomyces cerevisiae . Genetics 145, 615–626 (1997).
Hsu, J.Y. et al. Mitotic phosphorylation of histone H3 is governed by Ipl1/aurora kinase and Glc7/PP1 phosphatase in budding yeast and nematodes. Cell 102, 279–291 (2000).
Bloecher, A. & Tatchell, K. Defects in Saccharomyces cerevisiae protein phosphatase type I activate the spindle/kinetochore checkpoint. Genes Dev. 13, 517–522 (1999).
Hillenmeyer, M.E. et al. The chemical genomic portrait of yeast: uncovering a phenotype for all genes. Science 320, 362–365 (2008).
Martin, J.L. & McMillan, F.M. SAM (dependent) I AM: the S-adenosylmethionine-dependent methyltransferase fold. Curr. Opin. Struct. Biol. 12, 783–793 (2002).
Arnaud, M.B. et al. Sequence resources at the Candida Genome Database. Nucleic Acids Res. 35, D452–D456 (2007).
Bliss, C. The toxicity of poisons applied jointly. Ann. Appl. Biol. 26, 585–615 (1939).
Lehar, J. et al. Chemical combination effects predict connectivity in biological systems. Mol. Syst. Biol. 3, 80 (2007).
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Young, D.W. et al. Integrating high-content screening and ligand-target prediction to identify mechanism of action. Nat. Chem. Biol. 4, 59–68 (2008).
Baguley, B.C. & Nash, R. Antitumour activity of substituted 9-anilinoacridines–comparison of in vivo and in vitro testing systems. Eur. J. Cancer 17, 671–679 (1981).
Nelson, E.M., Tewey, K.M. & Liu, L.F. Mechanism of antitumor drug action: poisoning of mammalian DNA topoisomerase II on DNA by 4′-(9-acridinylamino)-methanesulfon-m-anisidide. Proc. Natl. Acad. Sci. USA 81, 1361–1365 (1984).
Denny, W.A. et al. Potential antitumor agents. 36. Quantitative relationships between experimental antitumor activity, toxicity, and structure for the general class of 9-anilinoacridine antitumor agents. J. Med. Chem. 25, 276–315 (1982).
Terstappen, G.C., Schlupen, C., Raggiaschi, R. & Gaviraghi, G. Target deconvolution strategies in drug discovery. Nat. Rev. Drug Discov. 6, 891–903 (2007).
Zimmermann, G.R., Lehar, J. & Keith, C.T. Multi-target therapeutics: when the whole is greater than the sum of the parts. Drug Discov. Today 12, 34–42 (2007).
Borisy, A.A. et al. Systematic discovery of multicomponent therapeutics. Proc. Natl. Acad. Sci. USA 100, 7977–7982 (2003).
Bishop, A.C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000).
Gietz, R.D., Schiestl, R.H., Willems, A.R. & Woods, R.A. Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11, 355–360 (1995).
Kadosh, D. & Johnson, A.D. Rfg1, a protein related to the Saccharomyces cerevisiae hypoxic regulator Rox1, controls filamentous growth and virulence in Candida albicans . Mol. Cell. Biol. 21, 2496–2505 (2001).
Lan, C.Y. et al. Metabolic specialization associated with phenotypic switching in Candida albicans . Proc. Natl. Acad. Sci. USA 99, 14907–14912 (2002).
Breitling, R., Armengaud, P., Amtmann, A. & Herzyk, P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 573, 83–92 (2004).
Acknowledgements
We thank K. Tatchell (Louisiana State University) for sharing the glc-7 alleles and M. Cyert (Stanford University) for providing the S. cerevisiae genomic library. We thank H. Ng and S. Lockey for critically reading the manuscript and members of the chemogenomics lab at the Stanford Genome Technology Center for discussions. S.H. is supported by a graduate fellowship from the Agency for Science Technology and Research (Singapore). R.P.S. was supported by a postdoctoral fellowship from the Canadian Institutes of Health Research. K.M.S. and R.W.D. are supported by grants from the US National Institutes of Health; G.G. and C.N. are supported by grants from the US National Institutes of Health and the Canadian Institutes of Health Research (MOP-81340 to G.G. and MOP-84305 to C.N.).
Author information
Authors and Affiliations
Contributions
C.N., R.P.S. and S.H. designed the study, analyzed the data and wrote the paper; S.H. and R.P.S. did the experiments. G.G. analyzed the data and helped write the paper. A.S. did the glc7 allele analysis; I.M.W. did the cheminformatic analysis. M.P. designed the robotic assay platform; E.F. designed the database infrastructure. K.M.S. and C.Z. designed the cdc28-as experiment. R.W.D. provided valuable advice. M.M. and S.S. helped with genome-wide screens.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Methods (PDF 4803 kb)
Rights and permissions
About this article
Cite this article
Hoon, S., M Smith, A., Wallace, I. et al. An integrated platform of genomic assays reveals small-molecule bioactivities. Nat Chem Biol 4, 498–506 (2008). https://doi.org/10.1038/nchembio.100
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.100
This article is cited by
-
An inducible CRISPR interference library for genetic interrogation of Saccharomyces cerevisiae biology
Communications Biology (2020)
-
Chemogenomic study of gemcitabine using Saccharomyces cerevisiae as model cell—molecular insights about chemoresistance
Brazilian Journal of Microbiology (2020)
-
Hypoculoside, a sphingoid base-like compound from Acremonium disrupts the membrane integrity of yeast cells
Scientific Reports (2019)
-
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Nature Communications (2019)
-
Chemical-genetic profiling reveals limited cross-resistance between antimicrobial peptides with different modes of action
Nature Communications (2019)