Abstract
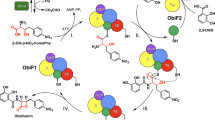
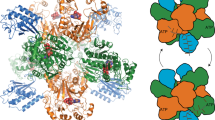
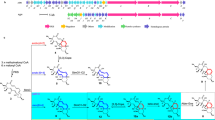
Oligomerization and macrocyclization reactions are key steps in the biosynthesis of many bioactive natural products. Important macrocycles include the antibiotic daptomycin (1; ref. 1), the immunosuppressant FK-506 (2; ref. 2), the anthelmintic avermectin B1a (3; ref. 3) and the insecticide spinosyn A (4; ref. 4); important oligomeric macrocycles include the siderophores enterobactin (5; ref. 5) and desferrioxamine E (6; ref. 6). Biosynthetic oligomerization and macrocyclization reactions typically involve covalently tethered intermediates and are catalyzed by thioesterase domains of polyketide synthase and nonribosomal peptide synthetase multienzymes7. Here we report that the purified recombinant desferrioxamine siderophore synthetase DesD from Streptomyces coelicolor M145 catalyzes ATP-dependent trimerization-macrocyclization of a chemically synthesized 10-aminocarboxylic acid substrate via noncovalently bound intermediates. DesD is dissimilar to other known synthetase families but is similar to other enzymes known or proposed to be required for the biosynthesis of ω-aminocarboxylic acid–derived cyclodimeric siderophores8,9. This suggests that DesD is the first biochemically characterized member of a new family of oligomerizing and macrocyclizing synthetases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
18 March 2009
In the version of this article initially published, an extra methyl group was inadvertently added to the structure of desferrioxamine G1 and related compounds. The error has been corrected in the HTML and PDF versions of the article.
References
Debono, M. et al. Enzymatic and chemical modifications of lipopeptide antibiotic A21978C: the synthesis and evaluation of daptomycin (LY146032). J. Antibiot. (Tokyo) 41, 1093–1105 (1988).
Tanaka, H. et al. Structure of FK506, a novel immunosuppressant isolated from Streptomyces. J. Am. Chem. Soc. 109, 5031–5033 (1987).
Springer, J.P., Arison, B.H., Hirshfield, J.M. & Hoogsteen, K. The absolute stereochemistry and conformation of avermectin B2a aglycone and avermectin B1a. J. Am. Chem. Soc. 103, 4221–4224 (1981).
Kirst, H.A. et al. A83543A-D, unique fermentation-derived tetracyclic macrolides. Tetrahedr. Lett. 32, 4839–4842 (1991).
Pollack, J.R. & Neilands, J.B. Enterobactin, an iron transport compound from Salmonella typhimurium. Biochem. Biophys. Res. Commun. 38, 989–992 (1970).
Bickel, H. et al. Metabolic products of Actinomycetaceae. XXVI. Isolation and properties of ferrioxamines A to F, representing new sideramine compounds. Helv. Chim. Acta 43, 2118–2128 (1960).
Grünewald, J. & Marahiel, M.A. Chemoenzymatic and template-directed synthesis of bioactive macrocyclic peptides. Microbiol. Mol. Biol. Rev. 70, 121–146 (2006).
Challis, G.L. A widely distributed bacterial pathway for siderophore biosynthesis independent of nonribosomal peptide synthetases. ChemBioChem 6, 601–611 (2005).
Kang, H.Y., Brickman, T.J., Beaumont, F.C. & Armstrong, S.K. Identification and characterization of iron-regulated Bordetella pertussis alcaligin siderophore biosynthesis genes. J. Bacteriol. 178, 4877–4884 (1996).
ShawReid, C.A. et al. Assembly line enzymology by multimodular nonribosomal peptide synthetases: the thioesterase domain of E. coli EntF catalyzes both elongation and cyclolactonization. Chem. Biol. 6, 385–400 (1999).
Trauger, J.W., Kohli, R.M., Mootz, H.D., Marahiel, M.A. & Walsh, C.T. Peptide cyclization catalysed by the thioesterase domain of tyrocidine synthetase. Nature 407, 215–218 (2000).
Hoyer, K.M., Mahlert, C. & Marahiel, M.A. The iterative gramicidin S thioesterase catalyzes peptide ligation and cyclization. Chem. Biol. 14, 13–22 (2007).
Tsai, S.-C. et al. Crystal structure of the macrocycle-forming thioesterase domain of the erythromycin polyketide synthase: versatility from a unique substrate channel. Proc. Natl. Acad. Sci. USA 98, 14808–14813 (2001).
Bruner, S.D. et al. Structural basis for the cyclization of the lipopeptide antibiotic surfactin by the thioesterase domain SrfTE. Structure 10, 301–310 (2002).
Tsai, S.-C., Lu, H., Cane, D.E., Khosla, C. & Stroud, R.M. Insights into channel architecture and substrate specificity from crystal structures of two macrocycle-forming thioesterases of modular polyketide synthases. Biochemistry 41, 12598–12606 (2002).
Samel, S.A., Wagner, B., Marahiel, M.A. & Essen, L.-O. The thioesterase domain of the fengycin biosynthesis cluster: a structural base for the macrocyclization of a non-ribosomal lipopeptide. J. Mol. Biol. 359, 876–889 (2006).
Akey, D.L. et al. Structural basis for macrolactonization by the pikromycin thioesterase. Nat. Chem. Biol. 2, 537–542 (2006).
Giraldes, J.W. et al. Structural and mechanistic insights into polyketide macrolactonization from polyketide-based affinity labels. Nat. Chem. Biol. 2, 531–536 (2006).
Schmidt, E.W. et al. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 102, 7315–7320 (2005).
Chatterjee, C. et al. Lacticin 481 synthetase phosphorylates its substrate during lantibiotic production. J. Am. Chem. Soc. 127, 15332–15333 (2005).
Keller-Schierlein, W. & Prelog, V. Metabolic products of actinomycetes. XXXIV. Ferrioxamine G. Helv. Chim. Acta 45, 590–595 (1962).
Takahashi, A. et al. Bisucaberin, a new siderophore, sensitizing tumor cells to macrophage-mediated cytolysis. II. Physico-chemical properties and structure determination. J. Antibiot. (Tokyo) 40, 1671–1676 (1987).
Nishio, T. et al. Isolation and structure of the novel dihydroxamate siderophore alcaligin. J. Am. Chem. Soc. 110, 8733–8734 (1988).
Ledyard, K.M. & Butler, A. Structure of putrebactin, a new dihydroxamate siderophore produced by Shewanella putrefaciens. J. Biol. Inorg. Chem. 2, 93–97 (1997).
Anderegg, G., L'Eplattenier, F. & Schwarzenbach, G. Hydroxamate complexes. III. Iron(III) exchange between sideramines and complexones. A discussion of the formation constants of the hydroxamate complexes. Helv. Chim. Acta 46, 1409–1422 (1963).
Hossain, M.B., Van der Helm, D. & Poling, M. The structure of deferriferrioxamine E (nocardamin), a cyclic trihydroxamate. Acta Crystallogr. B 39, 258–263 (1983).
Barona-Gomez, F. et al. Multiple biosynthetic and uptake systems mediate siderophore-dependent iron acquisition in Streptomyces coelicolor A3(2) and Streptomyces ambofaciens ATCC 23877. Microbiology 152, 3355–3366 (2006).
Barona-Gomez, F., Wong, U., Giannakopulos, A.E., Derrick, P.J. & Challis, G.L. Identification of a cluster of genes that directs desferrioxamine biosynthesis in Streptomyces coelicolor M145. J. Am. Chem. Soc. 126, 16282–16283 (2004).
Bergeron, R.J., Liu, Z.-R., McManis, J.S. & Wiegand, J. Structural alterations in desferrioxamine compatible with iron clearance in animals. J. Med. Chem. 35, 4739–4744 (1992).
Parkin, K.L. in Enzyme Kinetics: a Modern Approach (ed. Marangoni, A.G.) 174–192 (Jon Wiley and Sons, Hoboken, New Jersey, USA, 2003).
Acknowledgements
We thank L. Song for assistance with the LC-MS and MS/MS analyses and T.D.H. Bugg for helpful discussions. This research was funded by grants to G.L.C. from the Biotechnology and Biological Sciences Research Council SPoRT initiative (Grant ref. BBS/B/14450) and exploiting genomics initiative (Grant Ref. EGH16081).
Author information
Authors and Affiliations
Contributions
N.K. designed experiments, performed experiments, interpreted data and wrote the paper; D.O.-C. designed experiments, performed experiments, interpreted data and wrote the paper; F.B.-G. designed experiments, performed experiments and interpreted data; G.L.C. designed experiments, interpreted data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Methods (PDF 165 kb)
Rights and permissions
About this article
Cite this article
Kadi, N., Oves-Costales, D., Barona-Gomez, F. et al. A new family of ATP-dependent oligomerization-macrocyclization biocatalysts. Nat Chem Biol 3, 652–656 (2007). https://doi.org/10.1038/nchembio.2007.23
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2007.23
This article is cited by
-
Analogues of desferrioxamine B (DFOB) with new properties and new functions generated using precursor-directed biosynthesis
BioMetals (2019)
-
Novel desferrioxamine derivatives synthesized using the secondary metabolism-specific nitrous acid biosynthetic pathway in Streptomyces davawensis
The Journal of Antibiotics (2018)
-
The chemical biology and coordination chemistry of putrebactin, avaroferrin, bisucaberin, and alcaligin
JBIC Journal of Biological Inorganic Chemistry (2018)
-
Decoding and reprogramming fungal iterative nonribosomal peptide synthetases
Nature Communications (2017)
-
Anti-MRSA and anti-TB metabolites from marine-derived Verrucosispora sp. MS100047
Applied Microbiology and Biotechnology (2016)