Abstract
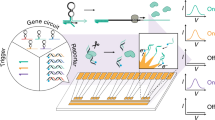
Synthetic biology advances the rational engineering of mammalian cells to achieve cell-based therapy goals. Synthetic gene networks have nearly reached the complexity of digital electronic circuits and enable single cells to perform programmable arithmetic calculations or to provide dynamic remote control of transgenes through electromagnetic waves. We designed a synthetic multilayered gaseous-fragrance-programmable analog-to-digital converter (ADC) allowing for remote control of digital gene expression with 2-bit AND-, OR- and NOR-gate logic in synchronized cell consortia. The ADC consists of multiple sampling-and-quantization modules sensing analog gaseous fragrance inputs; a gas-to-liquid transducer converting fragrance intensity into diffusible cell-to-cell signaling compounds; a digitization unit with a genetic amplifier circuit to improve the signal-to-noise ratio; and recombinase-based digital expression switches enabling 2-bit processing of logic gates. Synthetic ADCs that can remotely control cellular activities with digital precision may enable the development of novel biosensors and may provide bioelectronic interfaces synchronizing analog metabolic pathways with digital electronics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brophy, J.A.N. & Voigt, C.A. Principles of genetic circuit design. Nat. Methods 11, 508–520 (2014).
Moon, T.S., Lou, C., Tamsir, A., Stanton, B.C. & Voigt, C.A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Gitzinger, M., Kemmer, C., El-Baba, M.D., Weber, W. & Fussenegger, M. Controlling transgene expression in subcutaneous implants using a skin lotion containing the apple metabolite phloretin. Proc. Natl. Acad. Sci. USA 106, 10638–10643 (2009).
Gitzinger, M. et al. The food additive vanillic acid controls transgene expression in mammalian cells and mice. Nucleic Acids Res. 40, e37 (2012).
Weber, W., Bacchus, W., Daoud-El Baba, M. & Fussenegger, M. Vitamin H-regulated transgene expression in mammalian cells. Nucleic Acids Res. 35, e116 (2007).
Weber, W. et al. Conditional human VEGF-mediated vascularization in chicken embryos using a novel temperature-inducible gene regulation (TIGR) system. Nucleic Acids Res. 31, e69 (2003).
Boorsma, M. et al. A temperature-regulated replicon-based DNA expression system. Nat. Biotechnol. 18, 429–432 (2000).
Ausländer, D. et al. A synthetic multifunctional mammalian pH sensor and CO2 transgene-control device. Mol. Cell 55, 397–408 (2014).
Stanley, S.A. et al. Radio-wave heating of iron oxide nanoparticles can regulate plasma glucose in mice. Science 336, 604–608 (2012).
Folcher, M. et al. Mind-controlled transgene expression by a wireless-powered optogenetic designer cell implant. Nat. Commun. 5, 5392 (2014).
Weber, W. & Fussenegger, M. Inducible product gene expression technology tailored to bioprocess engineering. Curr. Opin. Biotechnol. 18, 399–410 (2007).
Weber, W. & Fussenegger, M. Emerging biomedical applications of synthetic biology. Nat. Rev. Genet. 13, 21–35 (2011).
Tigges, M., Marquez-Lago, T.T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Benenson, Y. Biomolecular computing systems: principles, progress and potential. Nat. Rev. Genet. 13, 455–468 (2012).
Ausländer, S., Ausländer, D., Müller, M., Wieland, M. & Fussenegger, M. Programmable single-cell mammalian biocomputers. Nature 487, 123–127 (2012).
Ausländer, S., Wieland, M. & Fussenegger, M. Smart medication through combination of synthetic biology and cell microencapsulation. Metab. Eng. 14, 252–260 (2012).
Kemmer, C. et al. Self-sufficient control of urate homeostasis in mice by a synthetic circuit. Nat. Biotechnol. 28, 355–360 (2010).
Rössger, K., Charpin-El-Hamri, G. & Fussenegger, M. Bile acid-controlled transgene expression in mammalian cells and mice. Metab. Eng. 21, 81–90 (2014).
Saxena, P., Charpin-El Hamri, G., Folcher, M., Zulewski, H. & Fussenegger, M. Synthetic gene network restoring endogenous pituitary-thyroid feedback control in experimental Graves' disease. Proc. Natl. Acad. Sci. USA 113, 1244–1249 (2016).
Schukur, L., Geering, B., Charpin-El Hamri, G. & Fussenegger, M. Implantable synthetic cytokine converter cells with AND-gate logic treat experimental psoriasis. Sci. Transl. Med. 7, 318ra201 (2015).
Riccione, K.A., Smith, R.P., Lee, A.J. & You, L. A synthetic biology approach to understanding cellular information processing. ACS Synth. Biol. 1, 389–402 (2012).
Regot, S. et al. Distributed biological computation with multicellular engineered networks. Nature 469, 207–211 (2011).
Tamsir, A., Tabor, J.J. & Voigt, C.A. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires'. Nature 469, 212–215 (2011).
Tubio, M.R. et al. Expression of a G protein-coupled receptor (GPCR) leads to attenuation of signaling by other GPCRs: experimental evidence for a spontaneous GPCR constitutive inactive form. J. Biol. Chem. 285, 14990–14998 (2010).
Bushdid, C., Magnasco, M.O., Vosshall, L.B. & Keller, A. Humans can discriminate more than 1 trillion olfactory stimuli. Science 343, 1370–1372 (2014).
Nern, A., Pfeiffer, B.D., Svoboda, K. & Rubin, G.M. Multiple new site-specific recombinases for use in manipulating animal genomes. Proc. Natl. Acad. Sci. USA 108, 14198–14203 (2011).
Fenno, L.E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014).
Yang, L. et al. Permanent genetic memory with >1-byte capacity. Nat. Methods 11, 1261–1266 (2014).
Friedland, A.E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Lapique, N. & Benenson, Y. Digital switching in a biosensor circuit via programmable timing of gene availability. Nat. Chem. Biol. 10, 1020–1027 (2014).
Schönhuber, N. et al. A next-generation dual-recombinase system for time- and host-specific targeting of pancreatic cancer. Nat. Med. 20, 1340–1347 (2014).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011).
Morel, M., Shtrahman, R., Rotter, V., Nissim, L. & Bar-Ziv, R.H. Cellular heterogeneity mediates inherent sensitivity-specificity tradeoff in cancer targeting by synthetic circuits. Proc. Natl. Acad. Sci. USA 113, 8133–8138 (2016).
Nissim, L. & Bar-Ziv, R.H. A tunable dual-promoter integrator for targeting of cancer cells. Mol. Syst. Biol. 6, 444 (2010).
Toh-e, A. & Utatsu, I. Physical and functional structure of a yeast plasmid, pSB3, isolated from Zygosaccharomyces bisporus. Nucleic Acids Res. 13, 4267–4283 (1985).
Bonnet, J., Yin, P., Ortiz, M.E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Hartwell, L.H., Hopfield, J.J., Leibler, S. & Murray, A.W. From molecular to modular cell biology. Nature 402 Suppl: C47–C52 (1999).
Sprinzak, D. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors. Cell 164, 780–791 (2016).
Clark, B. & Häusser, M. Neural coding: hybrid analog and digital signalling in axons. Curr. Biol. 16, R585–R588 (2006).
Dreosti, E., Esposti, F., Baden, T. & Lagnado, L. In vivo evidence that retinal bipolar cells generate spikes modulated by light. Nat. Neurosci. 14, 951–952 (2011).
Hahnloser, R.H., Sarpeshkar, R., Mahowald, M.A., Douglas, R.J. & Seung, H.S. Digital selection and analogue amplification coexist in a cortex-inspired silicon circuit. Nature 405, 947–951 (2000).
Firestein, S. How the olfactory system makes sense of scents. Nature 413, 211–218 (2001).
Roybal, K.T. et al. Precision tumor recognition by T cells with combinatorial antigen-sensing circuits. Cell 164, 770–779 (2016).
Chen, Y.Y., Jensen, M.C. & Smolke, C.D. Genetic control of mammalian T-cell proliferation with synthetic RNA regulatory systems. Proc. Natl. Acad. Sci. USA 107, 8531–8536 (2010).
Kemmer, C. et al. A designer network coordinating bovine artificial insemination by ovulation-triggered release of implanted sperms. J. Control. Release 150, 23–29 (2011).
Ye, H., Daoud-El Baba, M., Peng, R.W. & Fussenegger, M. A synthetic optogenetic transcription device enhances blood-glucose homeostasis in mice. Science 332, 1565–1568 (2011).
Roquet, N., Soleimany, A.P., Ferris, A.C., Aaronson, S. & Lu, T.K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
Siuti, P., Yazbek, J. & Lu, T.K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Rubens, J.R., Selvaggio, G. & Lu, T.K. Synthetic mixed-signal computation in living cells. Nat. Commun. 7, 11658 (2016).
Saito, H., Kubota, M., Roberts, R.W., Chi, Q. & Matsunami, H. RTP family members induce functional expression of mammalian odorant receptors. Cell 119, 679–691 (2004).
Acknowledgements
We thank T. Horn, E. Montani, T. Lopes and V. Jaeggin for their support with microscopy and flow cytometry. We thank F. Sedlmayer, V. Viswam, S. Bürgel and P. Buchmann for their generous advice and O. Chepurny and G. Holz (Department of Physiology and Neuroscience, New York University School of Medicine, New York, USA), H. Matsunami (Department of Molecular Genetics and Microbiology, Duke University Medical Center, North Carolina, USA), H. Ye (Shanghai Key Laboratory of Regulatory Biology, Institute of Biomedical Sciences and School of Life Sciences, East China Normal University, Shanghai, China) and W. Bacchus (Department of Biosystems Science and Engineering, ETH Zurich, Basel, Switzerland) for providing plasmids and genetic components. This work was supported by a European Research Council (ERC) advanced grant (ProNet, no. 321381) and in part by funding from the National Centre of Competence in Research (NCCR) for Molecular Systems Engineering awarded to M.F.
Author information
Authors and Affiliations
Contributions
M.M., S.A., D.A., M. Folcher and M. Fussenegger designed the project and analyzed the results; M.M., J.S. and A.S. performed the experimental work; and M.M. and M. Fussenegger wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3 and Supplementary Figures 1–13 (PDF 3812 kb)
Rights and permissions
About this article
Cite this article
Müller, M., Ausländer, S., Spinnler, A. et al. Designed cell consortia as fragrance-programmable analog-to-digital converters. Nat Chem Biol 13, 309–316 (2017). https://doi.org/10.1038/nchembio.2281
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2281
This article is cited by
-
An electrogenetic interface to program mammalian gene expression by direct current
Nature Metabolism (2023)
-
Synthetic memory circuits for stable cell reprogramming in plants
Nature Biotechnology (2022)
-
Synthetic neuromorphic computing in living cells
Nature Communications (2022)
-
Quantitative characterization of recombinase-based digitizer circuits enables predictable amplification of biological signals
Communications Biology (2021)
-
Protease circuits for processing biological information
Nature Communications (2020)