Abstract
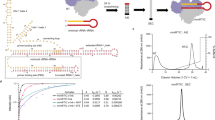
The anti-AIDS drug rilpivirine undergoes conformational changes to bind HIV-1 reverse transcriptase (RT), which is an essential enzyme for the replication of HIV. These changes allow it to retain potency against mutations that otherwise would render the enzyme resistant. Here we report that water molecules play an essential role in this binding process. Femtosecond experiments and theory expose the molecular level dynamics of rilpivirine bound to HIV-1 RT. Two nitrile substituents, one on each arm of the drug, are used as vibrational probes of the structural dynamics within the binding pocket. Two-dimensional vibrational echo spectroscopy reveals that one nitrile group is unexpectedly hydrogen-bonded to a mobile water molecule, not identified in previous X-ray structures. Ultrafast nitrile–water dynamics are confirmed by simulations. A higher (1.51 Å) resolution X-ray structure also reveals a water–drug interaction network. Maintenance of a crucial anchoring hydrogen bond may help retain the potency of rilpivirine against pocket mutations despite the structural variations they cause.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Das, K. et al. High-resolution structures of HIV-1 reverse transcriptase/TMC278 complexes: strategic flexibility explains potency against resistance mutations. Proc. Natl Acad. Sci. USA 105, 1466–1471 (2008).
Bollini, M. et al. Computationally-guided optimization of a docking hit to yield catechol diethers as potent anti-HIV agents. J. Med. Chem. 54, 8582–8591 (2011).
Mccoy, S. & Caughey, W. S. Infrared studies of azido, cyano, and other derivatives of metmyoglobin, methemoglobin, and hemins. Biochemistry 9, 2387–2393 (1970).
Getahun, Z. et al. Using nitrile-derivatized amino acids as infrared probes of local environment. J. Am. Chem. Soc. 125, 405–411 (2003).
Ghosh, A., Remorino, A., Tucker, M. J. & Hochstrasser, R. M. 2D-IR photon echo spectroscopy reveals hydrogen bond dynamics of aromatic nitriles. Chem. Phys. Lett. 469, 325–330 (2009).
Kim, Y. S. & Hochstrasser, R. M. Chemical exchange 2D-IR of hydrogen-bond making and breaking. Proc. Natl Acad. Sci. USA 102, 11185–11190 (2005).
Fang, C. et al. Two-dimensional infrared spectra reveal relaxation of the nonnucleoside inhibitor TMC278 complexed with HIV-1 reverse transcriptase. Proc. Natl Acad. Sci. USA 105, 1472–1477 (2008).
Hamm, P., Lim, M. H. & Hochstrasser, R. M. Structure of the amide I band of peptides measured by femtosecond nonlinear-infrared spectroscopy. J. Phys. Chem. B 102, 6123–6138 (1998).
Hochstrasser, R. M. Two-dimensional spectroscopy at infrared and optical frequencies. Proc. Natl Acad. Sci. USA 104, 14190–14196 (2007).
Finkelstein, I. J. et al. Probing dynamics of complex molecular systems with ultrafast 2D-IR vibrational echo spectroscopy. Phys. Chem. Chem. Phys. 9, 1533–1549 (2007).
Ganim, Z. et al. Amide I two-dimensional infrared spectroscopy of proteins. Acc. Chem. Res. 41, 432–441 (2008).
Andrews, S. S. & Boxer, S. G. Vibrational Stark effects of nitriles I. Methods and experimental results. J. Phys. Chem. A 104, 11853–11863 (2000).
Andrews, S. S. & Boxer, S. G. Vibrational Stark effects of nitriles II. Physical origins of Stark effects from experiment and perturbation models. J. Phys. Chem. A 106, 469–477 (2002).
Cho, M. H. et al. The integrated photon echo and solvation dynamics. J. Phys. Chem. 100, 11944–11953 (1996).
DeBoeij, W. P., Pshenichnikov, M. S. & Wiersma, D. A. On the relation between the echo-peak shift and Brownian-oscillator correlation function. Chem. Phys. Lett. 253, 53–60 (1996).
Kuo, C. H. & Hochstrasser, R. M. Two dimensional infrared spectroscopy and relaxation of aqueous cyanide. Chem. Phys. 341, 21–28 (2007).
Choi, J. H. et al. Nitrile and thiocyanate IR probes: quantum chemistry calculation studies and multivariate least-square fitting analysis. J. Chem. Phys. 128, 134506 (2008).
Kim, Y. S., Liu, L., Axelsen, P. H. & Hochstrasser, R. M. 2D-IR provides evidence for mobile water molecules in beta-amyloid fibrils. Proc. Natl Acad. Sci. USA 106, 17751–17756 (2009).
Ghosh, A., Qiu, J., DeGrado, W. F. & Hochstrasser, R. M. Tidal surge in the M2 proton channel, sensed by 2D-IR spectroscopy. Proc. Natl Acad. Sci. USA 108, 6115–6120 (2011).
Tucker, M. J. et al. 2D-IR photon echo of azido-probes for biomolecular dynamics. Phys. Chem. Chem. Phys. 13, 2237–2241 (2011).
Kuroda, D. G., Vorobyev, D. Y. & Hochstrasser, R. M. Ultrafast relaxation and 2D-IR of the aqueous trifluorocarboxylate ion. J. Chem. Phys. 132, 044501 (2010).
Bonner, G. M. et al. Probing the effect of the solution environment on the vibrational dynamics of an enzyme model system with ultrafast 2D-IR spectroscopy. Faraday Discuss. 145, 429–442 (2010).
Schultz, K. C. et al. A genetically encoded infrared probe. J. Am. Chem. Soc. 128, 13984–13985 (2006).
Kuroda, D. G. & Hochstrasser, R. M. Dynamic structures of aqueous oxalate and the effects of counterions seen by 2D-IR. Phys. Chem. Chem. Phys. 14, 6219–6224 (2012).
Corcelli, S. A., Lawrence, C. P. & Skinner, J. L. Combined electronic structure/molecular dynamics approach for ultrafast infrared spectroscopy of dilute HOD in liquid H2O and D2O. J. Chem. Phys. 120, 8107–8117 (2004).
Moller, K. B., Rey, R. & Hynes, J. T. Hydrogen bond dynamics in water and ultrafast infrared spectroscopy: a theoretical study. J. Phys. Chem. A 108, 1275–1289 (2004).
Ge, N. H., Zanni, M. T. & Hochstrasser, R. M. Effects of vibrational frequency correlations on two-dimensional infrared spectra. J. Phys. Chem. A 106, 962–972 (2002).
Bauman, J. D. et al. Crystal engineering of HIV-1 reverse transcriptase for structure-based drug design. Nucleic Acids Res. 36, 5083–5092 (2008).
Kim, Y. S., Wang, J. P. & Hochstrasser, R. M. Two-dimensional infrared spectroscopy of the alanine dipeptide in aqueous solution. J. Phys. Chem. B 109, 7511–7521 (2005).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D 66, 486–501 (2010).
Case, D. A. et al. AMBER 11 (University of California, San Francisco, 2010).
Wang, J. M., Cieplak, P. & Kollman, P. A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 21, 1049–1074 (2000).
Wang, J. M. et al. Development and testing of a general AMBER force field. J. Comput. Chem. 25, 1157–1174 (2004).
Jorgensen, W. L. et al. comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Acknowledgements
We thank A. Roitberg for advice on the MD simulations. This research was supported by National Institutes of Health (NIH) Grants GM12592 (to R.M.H.) and NIH MERIT Award R37 AI27690 (to E.A.) by using instrumentation developed in the Ultrafast Optical Process Laboratory NIH Grant P41RR001348/9P41GM104605. We acknowledge the Brookhaven National Laboratory X29 beamline facility for X-ray data collection and R.S.K. Vijayan for helpful discussions.
Author information
Authors and Affiliations
Contributions
E.A. and R.M.H. conceived and designed the experiments. J.R.C. and T.T. performed the infrared experiments. D.G.K, J.R.C. and T.T. analysed the data. D.G.K. performed theoretical calculations and simulations. J.D.B. expressed and purified the protein samples and grew the crystals. J.D.B. and D.P. performed the X-ray data collection. J.D.B. and K.D. analysed and refined the crystal structure. D.G.K. and R.M.H. co-wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Kuroda, D., Bauman, J., Challa, J. et al. Snapshot of the equilibrium dynamics of a drug bound to HIV-1 reverse transcriptase. Nature Chem 5, 174–181 (2013). https://doi.org/10.1038/nchem.1559
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1559
This article is cited by
-
In silico induction of missense mutation in NNRTI protein: computational modelling and stability study of modelled proteins
Journal of Mathematical Chemistry (2023)
-
Design, antihuman immunodeficiency activity and molecular docking studies of synthesized 2-aryl and 2-pyrimidinyl pyrrolidines
Molecular Diversity (2021)
-
Ultrafast structural molecular dynamics investigated with 2D infrared spectroscopy methods
Topics in Current Chemistry (2017)
-
How intermolecular geometrical disorder affects the molecular doping of donor–acceptor copolymers
Nature Communications (2015)
-
Effects of the protonation state in the interaction of an HIV-1 reverse transcriptase (RT) amino acid, Lys101, and a non nucleoside RT inhibitor, GW420867X
Journal of Molecular Modeling (2014)