Abstract
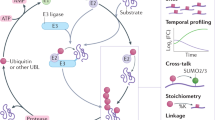
Proteolysis via the ubiquitin–proteasome system (UPS) is a rapid and effective method of degrading a specific protein at a specific time, and in many cases a protein is degraded only in response to a particular cellular signal or event. However, an added dimension to the control of protein degradation is possible because the ubiquitin system can be spatially regulated. Controlling where a protein is degraded can enhance the specificity and timing of proteolysis, generate asymmetry and maintain sub-compartments even in the mitotic cell. Here, we discuss this aspect of the UPS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Baumeister, W., Walz, J., Zuhl, F. & Seemuller, E. The proteasome: paradigm of a self-compartmentalizing protease. Cell 92, 367–380 (1998).
Voges, D., Zwickl, P. & Baumeister, W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (1999).
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Glickman, M. H. & Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 82, 373–428 (2002).
Passmore, L. A. & Barford, D. Getting into position: the catalytic mechanisms of protein ubiquitylation. Biochem. J. 379, 513–525 (2004).
Pintard, L., Willems, A. & Peter, M. Cullin-based ubiquitin ligases: Cul3–BTB complexes join the family. EMBO J. 23, 1681–1687 (2004).
Kile, B. T. et al. The SOCS box: a tale of destruction and degradation. Trends Biochem. Sci. 27, 235–241 (2002).
Hartmann-Petersen, R., Seeger, M. & Gordon, C. Transferring substrates to the 26S proteasome. Trends Biochem. Sci. 28, 26–31 (2003).
Koegl, M. et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96, 635–644 (1999).
Kleijnen, M. F. et al. The hPLIC proteins may provide a link between the ubiquitination machinery and the proteasome. Mol. Cell 6, 409–419 (2000).
Wilkinson, K. D. Ubiquitination and deubiquitination: targeting of proteins for degradation by the proteasome. Semin. Cell Dev. Biol. 11, 141–148 (2000).
Peters, J. M. The anaphase-promoting complex: proteolysis in mitosis and beyond. Mol. Cell 9, 931–943 (2002).
Vodermaier, H. C. APC/C and SCF: controlling each other and the cell cycle. Curr. Biol. 14, R787–R796 (2004).
Melchior, F. SUMO — nonclassical ubiquitin. Annu. Rev. Cell Dev. Biol. 16, 591–626 (2000).
Verma, R. et al. Proteasomal proteomics: identification of nucleotide-sensitive proteasome-interacting proteins by mass spectrometric analysis of affinity-purified proteasomes. Mol. Biol. Cell 11, 3425–3439 (2000).
Xie, Y. & Varshavsky, A. Physical association of ubiquitin ligases and the 26S proteasome. Proc. Natl Acad. Sci. USA 97, 2497–2502 (2000).
Leggett, D. S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002).
Glotzer, M., Murray, A. W. & Kirschner, M. W. Cyclin is degraded by the ubiquitin pathway. Nature 349, 132–138 (1991).
Verma, R. et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 298, 611–615 (2002).
Kostova, Z. & Wolf, D. H. For whom the bell tolls: protein quality control of the endoplasmic reticulum and the ubiquitin-proteasome connection. EMBO J. 22, 2309–2317 (2003).
Mayer, T. U., Braun, T. & Jentsch, S. Role of the proteasome in membrane extraction of a short-lived ER-transmembrane protein. EMBO J. 17, 3251–3257 (1998).
Lee, R. J. et al. Uncoupling retro-translocation and degradation in the ER-associated degradation of a soluble protein. EMBO J. 23, 2206–2215 (2004).
Swanson, R., Locher, M. & Hochstrasser, M. A conserved ubiquitin ligase of the nuclear envelope/endoplasmic reticulum that functions in both ER-associated and Matα2 repressor degradation. Genes Dev. 15, 2660–2674 (2001).
Blondel, M. et al. Nuclear-specific degradation of Far1 is controlled by the localization of the F-box protein Cdc4. EMBO J. 19, 6085–6097 (2000).
Freedman, D. A. & Levine, A. J. Nuclear export is required for degradation of endogenous p53 by MDM2 and human papillomavirus E6. Mol. Cell. Biol. 18, 7288–7293 (1998).
Yu, Z. K., Geyer, R. K. & Maki, C. G. MDM2-dependent ubiquitination of nuclear and cytoplasmic p53. Oncogene 19, 5892–5897 (2000).
Xirodimas, D. P., Stephen, C. W. & Lane, D. P. Cocompartmentalization of p53 and Mdm2 is a major determinant for Mdm2-mediated degradation of p53. Exp. Cell Res. 270, 66–77 (2001).
Li, M. et al. Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 302, 1972–1975 (2003).
Hara, T., Kamura, T., Nakayama, K., Oshikawa, K. & Hatakeyama, S. Degradation of p27(Kip1) at the G(0)–G(1) transition mediated by a Skp2-independent ubiquitination pathway. J. Biol. Chem. 276, 48937–48943 (2001).
Connor, M. K. et al. CRM1/Ran-mediated nuclear export of p27(Kip1) involves a nuclear export signal and links p27 export and proteolysis. Mol. Biol. Cell 14, 201–213 (2003).
Kamura, T. et al. Cytoplasmic ubiquitin ligase KPC regulates proteolysis of p27(Kip1) at G1 phase. Nature Cell Biol. 6, 1229–1235 (2004).
Carrano, A. C., Eytan, E., Hershko, A. & Pagano, M. SKP2 is required for ubiquitin-mediated degradation of the CDK inhibitor p27. Nature Cell Biol. 1, 193–199 (1999).
Tsvetkov, L. M., Yeh, K. H., Lee, S. J., Sun, H. & Zhang, H. p27(Kip1) ubiquitination and degradation is regulated by the SCF(Skp2) complex through phosphorylated Thr187 in p27. Curr. Biol. 9, 661–664 (1999).
Furstenthal, L., Swanson, C., Kaiser, B. K., Eldridge, A. G. & Jackson, P. K. Triggering ubiquitination of a CDK inhibitor at origins of DNA replication. Nature Cell Biol. 3, 715–722. (2001).
Mendez, J. et al. Human origin recognition complex large subunit is degraded by ubiquitin-mediated proteolysis after initiation of DNA replication. Mol. Cell 9, 481–491 (2002).
Lew, D. J. The morphogenesis checkpoint: how yeast cells watch their figures. Curr. Opin. Cell Biol. 15, 648–653 (2003).
McMillan, J. N. et al. The morphogenesis checkpoint in Saccharomyces cerevisiae: cell cycle control of Swe1p degradation by Hsl1p and Hsl7p. Mol. Cell. Biol. 19, 6929–6939 (1999).
Longtine, M. S. et al. Septin-dependent assembly of a cell cycle-regulatory module in Saccharomyces cerevisiae. Mol. Cell. Biol. 20, 4049–4061 (2000).
Bartholomew, C. R., Woo, S. H., Chung, Y. S., Jones, C. & Hardy, C. F. Cdc5 interacts with the Wee1 kinase in budding yeast. Mol. Cell. Biol. 21, 4949–4959 (2001).
McMillan, J. N., Theesfeld, C. L., Harrison, J. C., Bardes, E. S. & Lew, D. J. Determinants of Swe1p degradation in Saccharomyces cerevisiae. Mol. Biol. Cell 13, 3560–3575 (2002).
Musacchio, A. & Hardwick, K. G. The spindle checkpoint: structural insights into dynamic signalling. Nature Rev. Mol. Cell Biol. 3, 731–741 (2002).
Huang, J. Y. & Raff, J. W. The dynamic localisation of the Drosophila APC/C: evidence for the existence of multiple complexes that perform distinct functions and are differentially localised. J. Cell Sci. 115, 2847–2856 (2002).
Acquaviva, C., Herzog, F., Kraft, C. & Pines, J. The anaphase promoting complex/cyclosome is recruited to centromeres by the spindle assembly checkpoint. Nature Cell Biol. 6, 892–898 (2004).
Vigneron, S. et al. Kinetochore localization of spindle checkpoint proteins: who controls whom? Mol. Biol. Cell 15, 4584–4596 (2004).
Clute, P. & Pines, J. Temporal and spatial control of cyclin B1 destruction in metaphase. Nature Cell Biol. 1, 82–87 (1999).
Huang, J. & Raff, J. W. The disappearance of cyclin B at the end of mitosis is regulated spatially in Drosophila cells. EMBO J. 18, 2184–2195 (1999).
Wakefield, J. G., Huang, J. & Raff, J. W. Centrosomes have a role in regulating the destruction of cyclin B in early Drosophila embryos. Curr. Biol. 10, 1367–1370 (2000).
Raff, J. W., Jeffers, K. & Huang, J. Y. The roles of Fzy/Cdc20 and Fzr/Cdh1 in regulating the destruction of cyclin B in space and time. J. Cell Biol. 157, 1139–1149 (2002).
Mathe, E. et al. The E2-C vihar is required for the correct spatiotemporal proteolysis of cyclin B and itself undergoes cyclical degradation. Curr. Biol. 14, 1723–1733 (2004).
Rieder, C. L. et al. Mitosis in vertebrate somatic cells with two spindles: implications for the metaphase/anaphase transition checkpoint and cleavage. Proc. Natl Acad. Sci. USA 94, 5107–5112 (1997).
Lindon, C. & Pines, J. Ordered proteolysis in anaphase inactivates Plk1 to contribute to proper mitotic exit in human cells. J. Cell Biol. 164, 233–241 (2004).
Amsterdam, A., Pitzer, F. & Baumeister, W. Changes in intracellular localization of proteasomes in immortalized ovarian granulosa cells during mitosis associated with a role in cell cycle control. Proc. Natl Acad. Sci. USA 90, 99–103 (1993).
Tugendreich, S., Tomkiel, J., Earnshaw, W. & Hieter, P. CDC27Hs colocalizes with CDC16Hs to the centrosome and mitotic spindle and is essential for the metaphase to anaphase transition. Cell 81, 261–268 (1995).
Kraft, C. et al. Mitotic regulation of the human anaphase-promoting complex by phosphorylation. EMBO J. 22, 6598–6609 (2003).
Blondel, M. et al. Degradation of Hof1 by SCF(Grr1) is important for actomyosin contraction during cytokinesis in yeast. EMBO J. 24, 1440–1452 (2005).
Ehlers, M. D. Eppendorf 2003 prize-winning essay. Ubiquitin and the deconstruction of synapses. Science 302, 800–801 (2003).
Riechmann, V., Gutierrez, G. J., Filardo, P., Nebreda, A. R. & Ephrussi, A. Par-1 regulates stability of the posterior determinant Oskar by phosphorylation. Nature Cell Biol. 4, 337–342 (2002).
Raftopoulou, M. & Hall, A. Cell migration: Rho GTPases lead the way. Dev. Biol. 265, 23–32 (2004).
Wang, H. R. et al. Regulation of cell polarity and protrusion formation by targeting RhoA for degradation. Science 302, 1775–1779 (2003).
Campbell, D. S. & Holt, C. E. Chemotropic responses of retinal growth cones mediated by rapid local protein synthesis and degradation. Neuron 32, 1013–1026 (2001).
Acknowledgements
We are very grateful to our colleagues O. Coux, D. Lew, M. Scheffner and M. Peter for their helpful advice and to Cancer Research UK for financial support.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Pines, J., Lindon, C. Proteolysis: anytime, any place, anywhere?. Nat Cell Biol 7, 731–735 (2005). https://doi.org/10.1038/ncb0805-731
Issue Date:
DOI: https://doi.org/10.1038/ncb0805-731
This article is cited by
-
Endoplasmic reticulum–localized UBC34 interaction with lignin repressors MYB221 and MYB156 regulates the transactivity of the transcription factors in Populus tomentosa
BMC Plant Biology (2019)
-
Investigation of Novel Regulation of N-myristoyltransferase by Mammalian Target of Rapamycin in Breast Cancer Cells
Scientific Reports (2018)
-
A mammalian nervous-system-specific plasma membrane proteasome complex that modulates neuronal function
Nature Structural & Molecular Biology (2017)
-
Targeting of Fzr/Cdh1 for timely activation of the APC/C at the centrosome during mitotic exit
Nature Communications (2016)
-
BubR1 acetylation at prometaphase is required for modulating APC/C activity and timing of mitosis
The EMBO Journal (2009)