Abstract
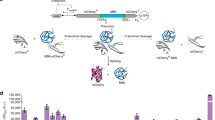
Segmental isotopic labeling of proteins using protein ligation is a recently established in vitro method for incorporating isotopes into one domain or region of a protein to reduce the complexity of NMR spectra, thereby facilitating the NMR analysis of larger proteins1,2,3. Here we demonstrate that segmental isotopic labeling of proteins can be conveniently achieved in Escherichia coli using intein-based protein ligation. Our method is based on a dual expression system that allows sequential expression of two precursor fragments in media enriched with different isotopes. Using this in vivo approach, unlabeled protein tags can be incorporated into isotopically labeled target proteins to improve protein stability and solubility for study by solution NMR spectroscopy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yamazaki, T. et al. Segmental isotope labeling for protein NMR using peptide splicing. J. Am. Chem. Soc. 120, 5591–5592 (1998).
Xu, R., Ayers, B., Cowburn, D. & Muir, T.W. Chemical ligation of folded recombinant proteins: segmental isotopic labeling of domains for NMR studies. Proc. Natl. Acad. Sci. USA 96, 388–393 (1999).
Otomo, T., Ito, N., Kyogoku, Y. & Yamazaki, T. NMR observation of selected segments in a larger protein: central-segment isotope labeling through intein-mediated ligation. Biochemistry 38, 16040–16044 (1999).
McIntosh, L.P. & Dahlquist, F.W. Biosynthetic incorporation of 15N and 13C for assignment and interpretation of nuclear magnetic resonance spectra of proteins. Q. Rev. Biophys. 23, 1–38 (1990).
Pervushin, K., Riek, R., Wider, G. & Wüthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Yu, H. Extending the size limit of protein nuclear magnetic resonance. Proc. Natl. Acad. Sci. USA 96, 332–334 (1999).
Wider, G. & Wüthrich, K. NMR spectroscopy of large molecules and multimolecular assemblies in solution. Curr. Opin. Struct. Biol. 9, 594–601 (1999).
Otomo, T., Teruya, K., Uegaki, K., Yamazaki, T. & Kyogoku, Y. Improved segmental isotope labeling of proteins and application to a larger protein. J. Biomol. NMR 14, 105–114 (1999).
Scott, C.P., Abel-Santos, E., Wall, M., Wahnon, D.C. & Benkovic, S.J. Production of cyclic peptides and proteins in vivo. Proc. Natl. Acad. Sci. USA 96, 13638–13643 (1999).
Evans, T.C., Jr. et al. Protein trans-splicing and cyclization by a naturally split intein from the dnaE gene of Synechocystis species PCC6803. J. Biol. Chem. 275, 9091–9094 (2000).
Iwai, H., Lingel, A. & Plückthun, A. Cyclic green fluorescent protein produced in vivo using an artificially split PI-PfuI intein from Pyrococcus furiosus. J. Biol. Chem. 276, 16548–16554 (2001).
Ozawa, T., Nogami, S., Sato, M., Ohya, Y. & Umezawa, Y. A fluorescent indicator for detecting protein-protein interactions in vivo based on protein splicing. Anal. Chem. 72, 5151–5157 (2000).
Giriat, I. & Muir, T.W. Protein semi-synthesis in living cells. J. Am. Chem. Soc. 125, 7180–7181 (2003).
Makrides, S.C. Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol. Rev. 60, 512–538 (1996).
Wu, H., Hu, Z.M. & Liu, X.Q. Protein trans-splicing by a split intein encoded in a split DnaE gene of Synechocystis sp. PCC6803. Proc. Natl. Acad. Sci. USA 95, 9226–9231 (1998).
Guzman, L.M., Belin, D., Carson, M.J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
King, C.Y. et al. Prion-inducing domain 2–114 of yeast Sup35 protein transforms in vitro into amyloid-like filaments. Proc. Natl. Acad. Sci. USA 94, 6618–6622 (1997).
King, C.Y. & Diaz-Avalos, R. Protein-only transmission of three yeast prion strains. Nature 428, 319–323 (2004).
Zhou, P., Lugovskoy, A.A. & Wagner, G. A solubility-enhancement tag (SET) for NMR studies of poorly behaving proteins. J. Biomol. NMR 20, 11–14 (2001).
Orban, J., Alexander, P., Bryan, P. & Khare, D. Assessment of stability differences in the protein-G B1 and B2 domains from hydrogen-deuterium exchange - comparison with calorimetric data. Biochemistry 34, 15291–15300 (1995).
Ikegami, T. et al. Solution structure of the chitin-binding domain of Bacillus circulans WL-12 chitinase A1. J. Biol. Chem. 275, 13654–13661 (2000).
Boomershine, W.P., Raj, M.L., Gopalan, V. & Foster, M.P. Preparation of uniformly labeled NMR samples in Escherichia coli under the tight control of the araBAD promoter: expression of an archaeal homolog of the RNase P Rpp29 protein. Protein Expr. Purif. 28, 246–251 (2003).
Christendat, D. et al. Structural proteomics of an archaeon. Nat. Struct. Biol. 7, 903–909 (2000).
Serber, Z. & Dötsch, V. In-cell NMR spectroscopy. Biochemistry 40, 14317–14323 (2001).
Mills, K.V. & Paulus, H. Reversible inhibition of protein splicing by zinc ion. J. Biol. Chem. 276, 10832–10838 (2001).
Inoue, Y., Kishimoto, A., Hirao, J., Yoshida, M. & Taguchi, H. Strong growth polarity of yeast prion fiber revealed by single fiber imaging. J. Biol. Chem. 276, 35227–35230 (2001).
Piotto, M., Saudek, V. & Sklenar, V. Gradient-tailored excitation for single-quantum NMR-spectroscopy of aqueous-solutions. J. Biomol. NMR 2, 661–665 (1992).
Wittekind, M. & Mueller, L. HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha-carbon and beta-carbon resonances in proteins. J. Magn. Reson. B 101, 201–205 (1993).
Güntert, P., Dötsch, V., Wider, G. & Wüthrich, K. Processing of multidimensional NMR data with the new software PROSA. J. Biomol. NMR 2, 619–629 (1992).
Bartels, C., Xia, T.H., Billeter, M., Güntert, P. & Wüthrich, K. The program XEASY for computer-supported NMR spectral-analysis of biological macromolecules. J. Biomol. NMR 6, 1–10 (1995).
Acknowledgements
We thank Henry Paulus, Hideki Taguchi and Stephen F. Marino for providing the plasmids containing Ssp DnaE intein, Sup35NM and GB1, respectively. We also thank Pui Hang Tam, Jennifer Jin, Sarah Figley for their technical assistance and the Saskatchewan Structural Science Center for the use of NMR instruments. H.I. thanks Peter Güntert for his continuous support. This study is supported by the National Science and Engineering Research Council of Canada (298493-4) and the Biomolecular Structure Teaching and Research Program at the University of Saskatchewan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Strips from the HNCACB spectrum of [13C, 15N]-GB1-CBD (PDF 260 kb)
Rights and permissions
About this article
Cite this article
Züger, S., Iwai, H. Intein-based biosynthetic incorporation of unlabeled protein tags into isotopically labeled proteins for NMR studies. Nat Biotechnol 23, 736–740 (2005). https://doi.org/10.1038/nbt1097
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1097
This article is cited by
-
Nature-inspired protein ligation and its applications
Nature Reviews Chemistry (2023)
-
Sparse pseudocontact shift NMR data obtained from a non-canonical amino acid-linked lanthanide tag improves integral membrane protein structure prediction
Journal of Biomolecular NMR (2023)
-
Kinetics study of the natural split Npu DnaE intein in the generation of bispecific IgG antibodies
Applied Microbiology and Biotechnology (2022)
-
Segmental isotopic labeling by asparaginyl endopeptidase-mediated protein ligation
Journal of Biomolecular NMR (2018)
-
Efficient generation of bispecific IgG antibodies by split intein mediated protein trans-splicing system
Scientific Reports (2017)