Abstract
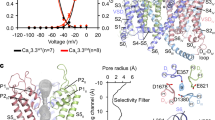
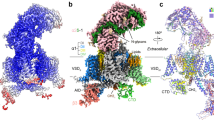
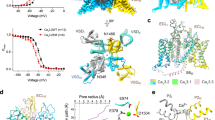
Among the ten subtypes of mammalian voltage-gated calcium (Cav) channels, Cav3.1–Cav3.3 constitute the T-type, or the low-voltage-activated, subfamily, the abnormal activities of which are associated with epilepsy, psychiatric disorders and pain1,2,3,4,5. Here we report the cryo-electron microscopy structures of human Cav3.1 alone and in complex with a highly Cav3-selective blocker, Z9446,7, at resolutions of 3.3 Å and 3.1 Å, respectively. The arch-shaped Z944 molecule reclines in the central cavity of the pore domain, with the wide end inserting into the fenestration on the interface between repeats II and III, and the narrow end hanging above the intracellular gate like a plug. The structures provide the framework for comparative investigation of the distinct channel properties of different Cav subfamilies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates and EM maps for Cav3.1 alone and in complex with Z944 have been deposited in the PDB with the accession codes 6KZO and 6KZP, and the EMDB with the codes EMD-0791 and EMD-0792, respectively. Source Data for Fig. 3e and Extended Data Figs. 2a and 6a are available in the online version of the paper. All other data are available from the corresponding author upon reasonable request.
References
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007).
Nowycky, M. C., Fox, A. P. & Tsien, R. W. Three types of neuronal calcium channel with different calcium agonist sensitivity. Nature 316, 440–443 (1985).
Ertel, E. A. et al. Nomenclature of voltage-gated calcium channels. Neuron 25, 533–535 (2000).
Zamponi, G. W., Striessnig, J., Koschak, A. & Dolphin, A. C. The physiology, pathology, and pharmacology of voltage-gated calcium channels and their future therapeutic potential. Pharmacol. Rev. 67, 821–870 (2015).
Dolphin, A. C. Voltage-gated calcium channels and their auxiliary subunits: physiology and pathophysiology and pharmacology. J. Physiol. 594, 5369–5390 (2016).
Tringham, E. et al. T-type calcium channel blockers that attenuate thalamic burst firing and suppress absence seizures. Sci. Transl. Med. 4, 121ra19 (2012).
Casillas-Espinosa, P. M. et al. Z944, a novel selective T-type calcium channel antagonist delays the progression of seizures in the amygdala kindling model. PLoS One 10, e0130012 (2015).
Tanabe, T. et al. Primary structure of the receptor for calcium channel blockers from skeletal muscle. Nature 328, 313–318 (1987).
Perez-Reyes, E. et al. Molecular characterization of a neuronal low-voltage-activated T-type calcium channel. Nature 391, 896–900 (1998).
Yang, J., Ellinor, P. T., Sather, W. A., Zhang, J. F. & Tsien, R. W. Molecular determinants of Ca2+ selectivity and ion permeation in L-type Ca2+ channels. Nature 366, 158–161 (1993).
Ellinor, P. T., Yang, J., Sather, W. A., Zhang, J. F. & Tsien, R. W. Ca2+ channel selectivity at a single locus for high-affinity Ca2+ interactions. Neuron 15, 1121–1132 (1995).
Talavera, K. et al. Aspartate residues of the Glu-Glu-Asp-Asp (EEDD) pore locus control selectivity and permeation of the T-type Ca2+ channel α1G. J. Biol. Chem. 276, 45628–45635 (2001).
Perez-Reyes, E. Molecular physiology of low-voltage-activated T-type calcium channels. Physiol. Rev. 83, 117–161 (2003).
Deschênes, M., Paradis, M., Roy, J. P. & Steriade, M. Electrophysiology of neurons of lateral thalamic nuclei in cat: resting properties and burst discharges. J. Neurophysiol. 51, 1196–1219 (1984).
Zhan, X. J., Cox, C. L., Rinzel, J. & Sherman, S. M. Current clamp and modeling studies of low-threshold calcium spikes in cells of the cat’s lateral geniculate nucleus. J. Neurophysiol. 81, 2360–2373 (1999).
Aizenman, C. D. & Linden, D. J. Regulation of the rebound depolarization and spontaneous firing patterns of deep nuclear neurons in slices of rat cerebellum. J. Neurophysiol. 82, 1697–1709 (1999).
Burlhis, T. M. & Aghajanian, G. K. Pacemaker potentials of serotonergic dorsal raphe neurons: contribution of a low-threshold Ca2+ conductance. Synapse 1, 582–588 (1987).
Carbone, E. & Lux, H. D. A low voltage-activated, fully inactivating Ca channel in vertebrate sensory neurones. Nature 310, 501–502 (1984).
Catterall, W. A. Structure and regulation of voltage-gated Ca2+ channels. Annu. Rev. Cell Dev. Biol. 16, 521–555 (2000).
Zamponi, G. W. Targeting voltage-gated calcium channels in neurological and psychiatric diseases. Nat. Rev. Drug Discov. 15, 19–34 (2016).
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015).
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 at 3.6 Å resolution. Nature 537, 191–196 (2016).
Shen, H. et al. Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution. Science 355, eaal4326 (2017).
Shen, H., Liu, D., Wu, K., Lei, J. & Yan, N. Structures of human Nav1.7 channel in complex with auxiliary subunits and animal toxins. Science 363, 1303–1308 (2019).
Pan, X. et al. Structure of the human voltage-gated sodium channel Nav1.4 in complex with β1. Science 362, eaau2486 (2018).
Pan, X. et al. Molecular basis for pore blockade of human Na+ channel Nav1.2 by the μ-conotoxin KIIIA. Science 363, 1309–1313 (2019).
Shcheglovitov, A. et al. Alternative splicing within the I-II loop controls surface expression of T-type Cav3.1 calcium channels. FEBS Lett. 582, 3765–3770 (2008).
Todorovic, S. M. et al. Redox modulation of T-type calcium channels in rat peripheral nociceptors. Neuron 31, 75–85 (2001).
Zhao, Y. et al. Molecular basis for ligand modulation of a mammalian voltage-gated Ca2+ channel. Cell 177, 1495–1506 (2019).
Cataldi, M., Perez-Reyes, E. & Tsien, R. W. Differences in apparent pore sizes of low and high voltage-activated Ca2+ channels. J. Biol. Chem. 277, 45969–45976 (2002).
Lipkind, G. M. & Fozzard, H. A. Molecular modeling of local anesthetic drug binding by voltage-gated sodium channels. Mol. Pharmacol. 68, 1611–1622 (2005).
Ahern, C. A., Eastwood, A. L., Dougherty, D. A. & Horn, R. Electrostatic contributions of aromatic residues in the local anesthetic receptor of voltage-gated sodium channels. Circ. Res. 102, 86–94 (2008).
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA 101, 16–22 (2004).
Gong, D. et al. Modulation of cardiac ryanodine receptor 2 by calmodulin. Nature 572, 347–351 (2019).
Lee, M. Z944: a first in class T-type calcium channel modulator for the treatment of pain. J. Peripher. Nerv. Syst. 19, S11–S12 (2014).
LeBlanc, B. W. et al. T-type calcium channel blocker Z944 restores cortical synchrony and thalamocortical connectivity in a rat model of neuropathic pain. Pain 157, 255–263 (2016).
Nam, G. T-type calcium channel blockers: a patent review (2012-2018). Expert Opin. Ther. Pat. 28, 883–901 (2018).
Marks, W. N. et al. The T-type calcium channel blocker Z944 reduces conditioned fear in Genetic Absence Epilepsy Rats from Strasbourg and the non-epileptic control strain. Eur. J. Neurosci. 50, 3046–3059 (2019).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
DeLano, W. L. The PyMOL Molecular Graphics System. http://www.pymol.org (2002).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360, 376 (1996).
Larkin, M. A. et al. Clustal W and clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Bourinet, E. & Zamponi, G. W. Block of voltage-gated calcium channels by peptide toxins. Neuropharmacology 127, 109–115 (2017).
IUPHAR/BPS. Guide to Pharmacology. https://www.guidetopharmacology.org/ (2019).
Weiss, N., Black, S. A., Bladen, C., Chen, L. & Zamponi, G. W. Surface expression and function of Cav3.2 T-type calcium channels are controlled by asparagine-linked glycosylation. Pflugers Arch. 465, 1159–1170 (2013).
Lazniewska, J., Rzhepetskyy, Y., Zhang, F. X., Zamponi, G. W. & Weiss, N. Cooperative roles of glucose and asparagine-linked glycosylation in T-type calcium channel expression. Pflugers Arch. 468, 1837–1851 (2016).
Liu, Y. et al. Asparagine-linked glycosylation modifies voltage-dependent gating properties of CaV3.1-T-type Ca2+ channel. J. Physiol. Sci. 69, 335–343 (2019).
Ondacova, K., Karmazinova, M., Lazniewska, J., Weiss, N. & Lacinova, L. Modulation of Cav3.2 T-type calcium channel permeability by asparagine-linked glycosylation. Channels 10, 175–184 (2016).
Weiss, N., Black, S. A. G., Bladen, C., Chen, L. & Zamponi, G. W. Surface expression and function of Cav3.2 T-type calcium channels are controlled by asparagine-linked glycosylation. Pflugers Arch. Eur. J. Physiol. 465, 1159–1170 (2013).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Tao, X., Lee, A., Limapichat, W., Dougherty, D. A. & MacKinnon, R. A gating charge transfer center in voltage sensors. Science 328, 67–73 (2010).
Coutelier, M. et al. A recurrent mutation in CACNA1G alters Cav3.1 T-type calcium-channel conduction and causes autosomal-dominant cerebellar ataxia. Am. J. Hum. Genet. 97, 726–737 (2015).
Morino, H. et al. A mutation in the low voltage-gated calcium channel CACNA1G alters the physiological properties of the channel, causing spinocerebellar ataxia. Mol. Brain 8, 89 (2015).
Chemin, J. et al. De novo mutation screening in childhood-onset cerebellar atrophy identifies gain-of-function mutations in the CACNA1G calcium channel gene. Brain 141, 1998–2013 (2018).
Splawski, I. et al. CACNA1H mutations in autism spectrum disorders. J. Biol. Chem. 281, 22085–22091 (2006).
Heron, S. E. et al. Extended spectrum of idiopathic generalized epilepsies associated with CACNA1H functional variants. Ann. Neurol. 62, 560–568 (2007).
Chen, Y. et al. Association between genetic variation of CACNA1H and childhood absence epilepsy. Ann. Neurol. 54, 239–243 (2003).
Heron, S. E. et al. Genetic variation of CACNA1H in idiopathic generalized epilepsy. Ann. Neurol. 55, 595–596 (2004).
Meyer, K. et al. Mutations in disordered regions can cause disease by creating dileucine motifs. Cell 175, 239–253 (2018).
Daniil, G. et al. CACNA1H mutations are associated with different forms of primary aldosteronism. EBioMedicine 13, 225–236 (2016).
Scholl, U. I. et al. Recurrent gain of function mutation in calcium channel CACNA1H causes early-onset hypertension with primary aldosteronism. eLife 4, e06315 (2015).
Acknowledgements
We thank X. Li for technical support during EM image acquisition. We thank J. Han for sharing the cDNA for human Cav3.1 (Uniprot O43497-9). This work was funded by the National Natural Science Foundation of China (projects 81920108015, 31800628 and 31621092), and the National Key R&D Program (2016YFA0500402 to X.P. and 2016YFA0501100 to J.L.) from Ministry of Science and Technology of China. We thank the Tsinghua University Branch of China National Center for Protein Sciences (Beijing) for providing the cryo-EM facility support. We thank the computational facility support on the cluster of Bio-Computing Platform (Tsinghua University Branch of China National Center for Protein Sciences Beijing) and the ‘Explorer 100’ cluster system of Tsinghua National Laboratory for Information Science and Technology. N.Y. is supported by the Shirley M. Tilghman endowed professorship from Princeton University.
Author information
Authors and Affiliations
Contributions
N.Y. conceived the project. Y.Z. and Q.W. conducted molecular cloning and protein purification. Y.Z., G.H., Q.W. and J.L. performed experiments for structural determination. K.W., X.P. and R.L. performed and analysed electrophysiological measurements. All authors contributed to data analysis. N.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Jörg Striessnig, Gerald W. Zamponi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Brief introduction to Cav channels.
a, A brief overview of the classification, physiology and pharmacology of mammalian Cav channels. The evolutionary distance is calculated by Clustal W55. The table is summarized from several reviews4,20,56,57. HVA, high-voltage-activated; LVA, low-voltage-activated; E-C coupling, excitation–contraction coupling; E-T coupling, excitation–transcription coupling. b, Pairwise comparison of sequence similarity and identity of full-length human Cav channels. The sequence alignment is provided as Supplementary Fig. 1. c, Topological structure of the Cav channels. The panel is adapted from our previous publication with some modifications23. For Cav3.1-Δ8b, residues 509–642, shown as dashed lines on the I–II linker, were deleted. No human splice variant corresponding to the mouse Cav3.1-Δ8b has been identified. In fact, the exon–intron boundaries do not support existence of such a variant in humans. Nevertheless, we name this construct Cav3.1-Δ8b to acknowledge the source where this construct was generated. Five glycosylation sites are observed on the extracellular loops, including Asn246/322/1428/1425/1675 (Fig. 1a). Glycosylation of the counterparts of Asn246 and Asn1425 has been reported in Cav3.258,59, and the glycosylation might modulate channel expression and activity60,61,62. d, The typical domain-swapped architecture of most voltage-gated ion channels63. Shown here is an extracellular view in which the voltage-sensing domains are shown as round rectangles.
Extended Data Fig. 2 Cryo-EM analysis of the human Cav3.1-Δ8b alone and in complex with Z944.
a, Whole-cell patch clamp measurements of the full-length human Cav3.1 and Cav3.1-Δ8b. n values indicate the number of independent cells; mean ± s.e.m. b, Last-step purification of human Cav3.1-Δ8b. Shown here is a representative size-exclusion chromatogram for proteins obtained from 30 l of HEK293F cells transfected with plasmids. The indicated peak fractions on the Coomassie-blue-stained SDS–PAGE (Supplementary Fig. 2) were pooled and concentrated for cryo-EM sample preparation. c, Representative electron micrograph and 2D class averages. The green circles indicate representative particles in distinct orientations. The black and white scale bars in the top and bottom panels represent 100 nm and 10 nm, respectively. d, Flowchart for EM data processing. Details can be found in Methods. e, The gold-standard Fourier shell correlation (FSC) curves for the 3D reconstructions. The middle and right panels show FSC curves for phase-randomized half maps and unmasked half maps for the apo (middle) and complex (right) datasets. f, FSC curves of the refined model versus the overall map that it was refined against (black); of the model refined in the first of the two independent maps used for the gold-standard FSC versus that same map (red); and of the model refined in the first of the two independent maps versus the second independent map (green). The small difference between the red and green curves indicates that the refinement of the atomic coordinates did not suffer from overfitting. Before calculation of FSC against model-generated map, both half maps and the merged map were multiplied by a solvent mask that only includes the protein region. The merged map was brought to a threshold at which the micelle is invisible and all transmembrane helices are visible. Dust points were manually removed using the hide dust function in Chimera. Caution was taken not to mask out the densities for the bound ligand and lipids. The map was then extended by 2 pixels and supplied with a soft edge width of 12 pixels using relion_mask_create. g, Local-resolution map for the 3D EM reconstruction of Cav3.1-Δ8b in the presence of Z944. The map, calculated in RELION-3.1, was generated in Chimera49.
Extended Data Fig. 3 EM maps for the transmembrane segments and lipids in Cav3.1-Δ8b.
a, EM maps for the S1–S6 segments in each repeat, shown as blue mesh, are contoured at 4–5σ. The maps were prepared in PyMol. b, Densities reminiscent of lipids and cholesteryl hemisuccinate (CHS) surrounding the pore domain. The densities are contoured at 7σ. For visual clarity, the ECLI and ECLIII are omitted in the extracellular view. PE, phosphatidylethanolamine. c, EM densities for the two calcium ions and surrounding residues from two half maps. The densities are contoured at 4.5σ.
Extended Data Fig. 4 EM maps for the transmembrane segments of the Z complex.
EM maps for the S1–S6 segments in each repeat, shown as magenta mesh, are contoured at 4–5σ.
Extended Data Fig. 5 Depolarized (‘up’) conformations of the four VSDs.
Structures of the four VSDs are presented in similar views. After purification in the absence of electric field with a lengthy duration, Nav and Cav channels are expected to be trapped in the inactivated states that are featured with depolarized or ‘up’ VSDs and closed intracellular gate. The S4 segments are in the 310 helix form. The gating charge residues and the conserved charge transfer centre64 are shown as ball and sticks. Other polar residues that form potential hydrogen bonds are represented by red dashed lines, with the gating charge residues shown as sticks. The two conserved polar or acidic residues on S2 that facilitate charge transfer, designated An1 and An2, are also labelled.
Extended Data Fig. 6 Local structural shifts of Cav3.1-Δ8b upon Z944 binding.
a, Lys1462, which is conserved in T-type channels only, is important for Z944 inhibition. State-dependent blockade by Z944 at indicated concentrations in cells expressing Cav3.1-Δ8b (left), Cav3.1-Δ8b (K1462F) (middle) and Cav3.1-Δ8b (K1462G) (right) are tested. n values indicate the number of independent cells; mean ± s.e.m. The sample sizes (n) tested from low to high concentrations are: n = 4, 5, 5, 3, 8, 6, 3 for Cav3.1-∆8b; n = 3, 4, 8, 8, 8, 8, 3 for Cav3.1-∆8b (K1462F); and n = 8, 8, 10, 10, 8, 6, 3 for Cav3.1-∆8b (K1462G). b, Several lipid and CHS molecules are resolved in the structure of Cav3.1-Δ8b. Shown here is an extracellular view. The lipids, the precise identities of which remain unclear, are shown as sticks. Phosphatidylethanolamine (PE) molecules were tentatively modelled into these densities. Three densities are reminiscent of cholesteryl hemisuccinate (CHS1–CHS3), although they may also belong to the detergent glyco-diosgenin (GDN). c, Structures of Cav3.1-Δ8b alone and in complex with Z944 can be superimposed, with a r.m.s.d. of 0.45 Å over 851 Cα atoms. Two perpendicular views of the superimposed structures are shown. Cav3.1-Δ8b alone is coloured by domain and the complex is coloured light blue. d, Change of lipid distribution in the pore domain in the presence of Z944. Left: an extracellular view of the superimposed pore domain of Cav3.1-Δ8b with or without Z944. The bound lipids, shown as thin sticks, are coloured dark grey for the apo structure and light blue for the complex. Z944 is shown as silver sticks. An extra lipid molecule, highlighted with a red rectangle, was resolved in the pore domain of the complex. Right: the densities for Z944 and the nearby transverse lipid are contoured at 4.5σ. It is noted that the densities that were tentatively assigned with two Ca2+ ions are contiguous with that for the transverse lipid. Although we cannot entirely exclude the possibility that the densities in the selectivity filter (SF) may belong to a lipid, they are more likely to be bound ions because: (1) If the density belongs to the head group of a lipid, the SF is too narrow to accommodate any known positively charged linear head group with the length corresponding to the density; if the density belongs to a tail, then the hydrophobic property is incompatible with the polar environment within the SF. (2) Lipid-like densities have been observed traversing the pore domain in nearly all structures of Nav and Cav channels with fenestrations. In these channels, a highly conserved inner site constituted by backbone C=O groups has been demonstrated to coordinate Na+ or Ca2+ by X-ray crystallographic and molecular dynamics simulation analyses. Taken together, two Ca2+ ions, instead of a lipid moiety, were tentatively assigned to the density in the SF. e, Half-map densities for the SF from two diagonal repeats, contoured at 4.5σ.
Extended Data Fig. 7 Structural mapping of disease mutations identified in Cav3.1 and Cav3.2.
Please refer to Extended Data Table 1 for details. Side views of the diagonal repeats are shown. SCA42, spinocerebellar ataxia 42; SCA42ND, spinocerebellar ataxia 42, early-onset, severe, with neurodevelopmental deficits; HALD4, hyperaldosteronism, familial, 4.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Zhao, Y., Huang, G., Wu, Q. et al. Cryo-EM structures of apo and antagonist-bound human Cav3.1. Nature 576, 492–497 (2019). https://doi.org/10.1038/s41586-019-1801-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1801-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.