Abstract
Melanoma is notable for its metastatic propensity, lethality in the advanced setting and association with ultraviolet exposure early in life1. To obtain a comprehensive genomic view of melanoma in humans, we sequenced the genomes of 25 metastatic melanomas and matched germline DNA. A wide range of point mutation rates was observed: lowest in melanomas whose primaries arose on non-ultraviolet-exposed hairless skin of the extremities (3 and 14 per megabase (Mb) of genome), intermediate in those originating from hair-bearing skin of the trunk (5–55 per Mb), and highest in a patient with a documented history of chronic sun exposure (111 per Mb). Analysis of whole-genome sequence data identified PREX2 (phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2)—a PTEN-interacting protein and negative regulator of PTEN in breast cancer2—as a significantly mutated gene with a mutation frequency of approximately 14% in an independent extension cohort of 107 human melanomas. PREX2 mutations are biologically relevant, as ectopic expression of mutant PREX2 accelerated tumour formation of immortalized human melanocytes in vivo. Thus, whole-genome sequencing of human melanoma tumours revealed genomic evidence of ultraviolet pathogenesis and discovered a new recurrently mutated gene in melanoma.
Similar content being viewed by others
Main
To gain a comprehensive view of the genomic landscape in human melanoma tumours, we sequenced the genomes of 25 metastatic melanomas and peripheral blood obtained from the same patients (Supplementary Table 1). Two tumours (ME015 and ME032) were metastases from cutaneous melanomas arising on glabrous (that is, hairless) skin of the extremities, representing the acral subtype. The other tumours were primarily metastases from melanomas originating on hair-bearing skin of the trunk (the most common clinical subtype). Further, ME009 represented a metastasis from a primary melanoma of a patient with a clinical history of chronic ultraviolet exposure.
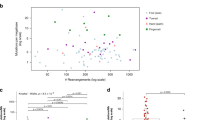
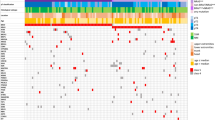
We obtained 59-fold mean haploid genome coverage for tumour DNA and 32-fold for normal DNA (Supplementary Table 2). On average, 78,775 somatic base substitutions per tumour were identified, consistent with prior reports3,4 (Supplementary Table 3). This corresponded to an average mutation rate of 30 per Mb. However, the mutation rate varied by nearly two orders of magnitude across the 25 tumours (Fig. 1). The acral melanomas showed mutation rates comparable to other solid tumour types (3 and 14 mutations per Mb)5,6, whereas melanomas from the trunk harboured substantially more mutations, in agreement with previous studies3,7,8. In particular, sample ME009 exhibited a striking rate of 111 somatic mutations per Mb, consistent with a history of chronic sun exposure.
Top bar plot shows somatic mutation rate of 25 sequenced melanoma genomes, in decreasing order. Middle matrix indicates BRAF and NRAS somatic mutation status, with left-adjacent bar plot indicating total number of mutations in each oncogene as well as percent frequency. Bottom plot displays each tumour’s somatic mutation spectrum as a percentage of all mutations (right axis). Tumour sample names are indicated at the bottom of the figure, with acral melanomas in red.
In tumours with elevated mutation rates, most nucleotide substitutions were C → T or G → A transitions consistent with ultraviolet irradiation9. The variations in mutation rate correlated with differences in the ultraviolet mutational signature. For example, 93% of substitutions in ME009 but only 36% in acral melanoma ME015 were C → T transitions (Fig. 1); these tumours contained the highest and lowest base mutation rates, respectively (111 and 3 mutations per Mb). Interestingly, the acral tumour ME032 also showed a discernible enrichment of ultraviolet-associated mutations (Fig. 1). Thus, genome sequencing readily confirmed the contribution of sun exposure in melanoma aetiology.
In agreement with prior studies7,9, we detected an overall enrichment for dipyrimidines at C → T transitions. Analysis of intragenic C → T mutations yielded a significant bias against such mutations on the transcribed strand for most melanomas, consistent with transcription-coupled repair (Supplementary Fig. 1)3,7,10. Most commonly, C → T mutations occurred at the 3′ base of a pyrimidine dinucleotide (CpC or TpC; Supplementary Fig. 2). In contrast, the C → T mutations in sample ME009 (with hypermutation and chronic sun exposure history) more often occurred at the 5′ base of a pyrimidine dinucleotide. As expected, the acral tumour ME015 exhibited mutation patterns observed in non-ultraviolet-associated tumour types11, such as an increased mutation rate at CpG dinucleotides relative to their overall genome-wide frequency (Supplementary Fig. 2). These different mutational signatures suggest a complex mechanism of ultraviolet mutagenesis across the clinical spectrum of melanoma, probably reflecting distinct histories of environmental exposures and cutaneous biology.
We detected 9,653 missense, nonsense or splice site mutations in 5,712 genes (out of a total of 14,680 coding mutations; Supplementary Tables 4 and 5), with an estimated specificity of 95% (Supplementary Methods). A mutation of BRAF, BRAFV600E , was present in 16 of 25 tumours (64%), including the acral melanoma ME015. NRAS was mutated in 9 of 25 tumours (36%) in a mutually exclusive fashion with BRAF, with the exception of one non-canonical substitution (NRAST50I ) in the hypermutated sample ME009. We also identified 6 insertions and 34 deletions in protein coding exons (Supplementary Table 6), including a 21-base-pair (bp) in-frame deletion involving exon 11 of the KIT oncogene in the acral tumour ME032 (Supplementary Fig. 3). KIT mutations occur in 15% of acral and mucosal melanomas12, and melanoma patients with activating KIT mutations in exon 11 have demonstrated marked responses to imatinib treatment13.
We identified an average of 97 structural rearrangements per melanoma genome (range: 6–420) (Supplementary Table 7). In addition to displaying a wide range of rearrangement frequencies, the proportion of intrachromosomal and interchromosomal rearrangements varied widely across genomes. ME029, which harboured the largest number of rearrangements (420), contained only 8 interchromosomal events (Fig. 2a). In contrast, ME020 and ME035 contained 95 and 90 interchromosomal rearrangements, respectively (Fig. 2a). In both cases, the vast majority of interchromosomal rearrangements were restricted to two chromosomes. This pattern is reminiscent of chromothripsis14, a process involving catastrophic chromosome breakage that has been observed in several tumour types15,16.
a, Circos plots representing four melanoma genomes with notable structural alterations. Interchromosomal and intrachromosomal rearrangements are shown in purple and green, respectively. b, Location of breakpoints associated with ETV1 in melanoma ME032. c, Location of breakpoints associated with PREX2 in melanoma ME032. The red arrow indicates a premature stop codon (E824*). All rearrangements in ETV1 and PREX2 were validated by high-throughput PCR and deep sequencing. d, Confirmation of high-level amplification and rearrangement of PREX2 in ME032 by dual-colour break-apart FISH. The assignment of red and green FISH probes to the PREX2 gene region is delineated as bars. Lack of co-localization of red and green probes is indicative of break-apart.
106 genes harboured chromosomal rearrangements in two or more samples (Supplementary Table 8). Many recurrently rearranged loci contain large genes or reside at known or suspected fragile sites17; examples include FHIT (six tumours), MACROD2 (five tumours) and CSMD1 (four tumours). On the other hand, several known cancer genes were also recurrently rearranged, including the PTEN tumour suppressor (four tumours) and MAGI2 (three tumours), which encodes a protein known to bind and stabilize PTEN. MAGI2 was also found disrupted in recent whole-genome studies of prostate cancer18 and a melanoma cell line7. Rearrangements involving the 5′ untranslated region of the ataxin 2-binding protein 1 gene (A2BP1) were observed in 4 tumours. A2BP1 encodes an RNA binding protein whose genetic disruption has been linked to spinocerebellar ataxia and other neurodegenerative diseases. A2BP1 undergoes complex splicing regulation in the central nervous system and other tissues19; in melanoma, these rearrangements may disrupt a known A2BP1 splice isoform or enable a de novo splicing product. Together, these results suggest that chromosomal rearrangements may contribute importantly to melanoma genesis or progression.
An acral melanoma (ME032) harboured the second-largest number of total rearrangements (314; Fig. 2a). We employed high-throughput PCR followed by massively parallel sequencing to successfully validate 177 of 182 events tested in this sample, confirming its high rate of rearrangement. The elevated frequency of genomic rearrangements in acral melanomas has been reported previously20. In comparison, ME032 exhibited one of the lowest base-pair mutation rates of the melanomas examined (22nd out of 25 samples), suggesting that different tumours might preferentially enact alternative mechanisms of genomic alteration to drive tumorigenesis.
As noted above, many rearrangements in ME032 involved multiple breakpoints within a narrow genomic interval. One such event disrupted the ETV1 locus. We previously demonstrated an oncogenic role for ETV1 in melanoma, whose dysregulated expression was associated with upregulation of microphthalmia-associated transcription factor (MITF)21, the master melanocyte transcriptional regulator and a melanoma lineage survival oncogene22. We validated six distinct rearrangements (four interchromosomal translocations) in ME032 involving breakpoints within ETV1 introns (Fig. 2b). These events join regions of ETV1 to distal loci on chromosomes 8, 9, 11 and 15. In support of their possible functional relevance, these rearrangements were associated with high-level ETV1 amplification in this tumour.
A second complex rearrangement involved the PREX2 locus. PREX2 encodes a phosphatidylinositol 3,4,5-trisphosphate RAC exchange factor recently shown to interact with the PTEN tumour suppressor and modulate its function2. We validated nine somatic rearrangements in the vicinity of PREX2 (six interchromosomal translocations), including five with intronic breakpoints (Fig. 2c, Supplementary Fig. 4). One event joined specific intronic regions of PREX2 and ETV1. Like ETV1, PREX2 is highly amplified in this tumour, as verified by FISH (fluorescence in situ hybridization) analysis (Fig. 2d, Supplementary Fig. 5). The presence of these complex structural rearrangements in addition to amplification may indicate multiple mechanisms of PREX2 dysregulation in melanoma. More generally, these findings raised the possibility that sites of complex rearrangement might denote genes of functional importance in melanoma.
Next, we calculated the mutational significance of each gene based on the number of mutations detected, gene length and background mutation rates (Table 1, Supplementary Table 9) (see Methods). Eleven genes were found to be significantly mutated across the 25 samples (Q < 0.01, where Q is the false discovery rate adjusted P-value). As expected, the two most significant genes were BRAF and NRAS, mutated in 16 and 9 samples, respectively. Interestingly, PREX2 scored as one of the top significant genes (Table 1). Furthermore, four samples harboured nonsense truncation mutations in PREX2, more than any of the other genes identified as statistically significant in this analysis. PREX2 mutations have occasionally been reported in colon, lung and pancreatic cancer23, albeit at low frequencies. Here, we detected 13 non-synonymous point mutations in PREX2—including 4 nonsense mutations—and 1 synonymous mutation, with 11 of 25 melanomas harbouring at least 1 non-synonymous mutation. The mutations were distributed throughout the entire length of PREX2 (Fig. 3a, green circles), and 13 of 14 mutations were non-synonymous, suggestive of positive selection. An analysis of the mutant allele frequencies and estimated tumour purities indicates that at least two mutations are homozygous. One melanoma, ME018, harbours three missense mutations, two of which (I534M and G1581R) appear to co-occur on a single allele based on their observed mutation frequencies. Notably, a PREX2 nonsense mutation was detected in ME032, in addition to the rearrangements and amplification of this locus present in this tumour (Fig. 2c). This PREX2 mutation was truncating (E824*), removing the carboxy-terminal region with homology to an inositol phosphatase domain. Based on the allele frequency of this mutation, we infer that it occurs on the non-amplified allele. Taken together, whole-genome sequencing of this 25-sample discovery cohort identified PREX2 as a candidate melanoma gene whose amplifications, rearrangements or mutations appeared to undergo positive selection in human melanoma genesis.
a, Non-synonymous sequence mutations detected from Illumina sequencing of 25 melanomas (green) or from capillary sequencing of a validation cohort of 107 additional melanomas (purple). Mutations are dispersed throughout all annotated structural domains of PREX2. (DH, DBL homology domain; PH, plekstrin homology domain). The C-terminal half of PREX2 exhibits sequence homology to an inositol phosphatase domain. Engineered PREX2 mutants are labelled with red arrowheads. b, Kaplan-Meier curve showing tumour-free survival of NUDE mice (n = 10) injected with PMEL-NRAS* cells expressing GFP, wild-type (left plot), truncated (middle plot) and mutated (right plot) PREX2 subcutaneously.
To determine the prevalence of PREX2 mutations in melanoma, we performed bidirectional capillary sequencing in an extension cohort of 107 tumour/normal pairs, comprising 45 tumours and 62 short-term cultures collected from multiple institutions and geographic regions (Supplementary Table 10). We identified 23 somatic base pair mutations and one frame-shift insertion in PREX2 in this cohort (Fig. 3a; Supplementary Table 11), 15 of which represented non-synonymous changes. We therefore inferred a 14% frequency of non-synonymous PREX2 mutations in this melanoma cohort.
Discrepant non-synonymous:synonymous ratios were observed between the tumour samples and short-term cultures in the extension cohort. In line with results from the discovery cohort, 100% of PREX2 mutations detected across 45 tumour samples were non-synonymous in nature (n = 4), consistent with positive selection. In contrast, only 55% of the sequence mutations found in the 64 short-term cultures were non-synonymous (a ratio of 11:9). Conceivably, these findings may indicate that subsets of melanoma cells capable of robust growth in vitro may have experienced reduced selective pressure for PREX2 mutations. Alternatively (or in addition), the PREX2 locus may exhibit an enhanced ‘local’ mutation rate, a by-product of which is the production of variants that undergo positive selection in vivo.
To demonstrate the functional relevance of PREX2 mutations in melanoma tumorigenesis, we ectopically expressed six representative mutations (three truncation variants and three non-synonymous point mutations predicted to carry functional impact24) in TERT-immortalized human melanocytes engineered to express NRAS(G12D) (PMEL-NRAS*)21. These melanocytic lines were transplanted into immunodeficient mice alongside control melanocytes expressing either wild-type PREX2 or GFP (green fluorescent protein). Overexpression of all three truncated variants as well as a point mutant (G844D) of PREX2 significantly accelerated in vivo tumorigenesis when compared to GFP control or wild-type PREX2-expressing melanocytes (Fig. 3b, Supplementary Fig. 6). These results therefore affirmed the aforementioned genomic data suggesting that PREX2 mutations may undergo positive selection in vivo. Although the spectrum of PREX2 mutations in human melanoma (Fig. 3a) is reminiscent of inactivating mutations, our findings suggest that PREX2 somatic mutations generate truncated or variant proteins that gain oncogenic activity in melanoma cells.
In summary, following recent efforts to characterize whole genomes from several haematologic and solid tumours, we provide the first (to our knowledge) high-resolution view of the genomic landscape across a spectrum of metastatic melanoma tumours. The analysis reveals global genomic evidence for the role of ultraviolet mutagenesis in melanoma, and identifies several recurrently mutated and rearranged genes not previously implicated in this malignancy. In particular, we discovered that PREX2 mutations are both recurrent and functionally consequential in melanoma biology. Although its precise mechanism(s) of action remains to be elucidated in melanoma, PREX2 appears to acquire oncogenic activity through mutations that perturb or inactivate one or more of its cellular functions. This pattern of mutations may exemplify a category of cancer genes that is distinct from ‘classic’ oncogenes (often characterized by highly recurrent gain-of-function mutations) and tumour suppressors (inactivated by simple loss-of-function alterations). Instead, (over)expression of certain cancer genes with distributed mutation patterns may promote tumorigenicity either through dominant negative effects or more subtle dysregulation of normal protein functions.
Cancer genomics has enabled the discovery and rational application of the first truly effective targeted therapy for metastatic melanoma: BRAF mutations predict sensitivity to selective RAF inhibitors25,26,27. However, the emergence of acquired resistance is rapid and often driven by other genomic events28. Our genomic exploration of the melanoma genomes revealed a large number of complex alterations that probably affect many other genes in addition to PREX2. Understanding how this spectrum of genomic aberrations contributes to melanoma genesis and progression should provide new insights into tumour biology, therapeutic resistance and development of treatment regimens aimed at durable control of this malignancy.
Methods Summary
The complete genomes of 25 metastatic melanomas and patient-matched germline samples were sequenced to approximately 30× and 30× haploid coverage, respectively, on an Illumina GAIIx sequencer (5 cases), and approximately 65× and 32× haploid coverage, respectively, on an Illumina HiSeq 2000 sequencer (20 cases) as paired-end 101-nucleotide reads. Read pairs were aligned to the reference human genome (hg19) using BWA29. Somatic alterations (single base substitutions, small insertions and deletions, and structural rearrangements) were identified according to their presence in the tumour genome and absence from the corresponding normal genome. A subset of rearrangements was validated by PCR and an independent sequencing technology in order to assess the specificity of the detection algorithm. Fluorescence in situ hybridization (FISH) was performed to confirm the high level amplification and rearrangement of PREX2. Significantly mutated genes were identified by comparing the observed mutations to the background mutation rates calculated for different sequence context categories per tumour sample. 40 exons of PREX2 were sequenced by PCR and bidirectional capillary sequencing in a validation panel of 107 additional melanoma tumours and short term cultures; mutations were confirmed as somatic by sequencing matched normal DNA. For gain of function studies, PREX2 mutation constructs were engineered and introduced to PMEL cell lines by lentiviral transduction. To assess the oncogenic roles of PREX2 mutants, PMEL-NRAS* cells were injected subcutaneously into NUDE mice, and tumour growth was measured over time. A complete description of the materials and methods is provided in Supplementary Information.
Accession codes
Data deposits
All Illumina sequence data are publicly available in dbGaP (accession number phs000452.v1.p1).
References
Chin, L. The genetics of malignant melanoma: lessons from mouse and man. Nature Rev. Cancer 3, 559–570 (2003)
Fine, B. et al. Activation of the PI3K pathway in cancer through inhibition of PTEN by exchange factor P-REX2a. Science 325, 1261–1265 (2009)
Berger, M. F. et al. Integrative analysis of the melanoma transcriptome. Genome Res. 20, 413–427 (2010)
Chapman, M. A. et al. Initial genome sequencing and analysis of multiple myeloma. Nature 471, 467–472 (2011)
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007)
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010)
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010)
Wei, X. et al. Exome sequencing identifies GRIN2A as frequently mutated in melanoma. Nature Genet. 43, 442–446 (2011)
Drobetsky, E. A., Grosovsky, A. J. & Glickman, B. W. The specificity of UV-induced mutations at an endogenous locus in mammalian cells. Proc. Natl Acad. Sci. USA 84, 9103–9107 (1987)
Vrieling, H. et al. Strand specificity for UV-induced DNA repair and mutations in the Chinese hamster HPRT gene. Nucleic Acids Res. 19, 2411–2415 (1991)
Rubin, A. F. & Green, P. Mutation patterns in cancer genomes. Proc. Natl Acad. Sci. USA 106, 21766–21770 (2009)
Curtin, J. A., Busam, K., Pinkel, D. & Bastian, B. C. Somatic activation of KIT in distinct subtypes of melanoma. J. Clin. Oncol. 24, 4340–4346 (2006)
Hodi, F. S. et al. Major response to imatinib mesylate in KIT-mutated melanoma. J. Clin. Oncol. 26, 2046–2051 (2008)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012)
Kloosterman, W. P. et al. Chromothripsis is a common mechanism driving genomic rearrangements in primary and metastatic colorectal cancer. Genome Biol. 12, R103 (2011)
Bignell, G. R. et al. Signatures of mutation and selection in the cancer genome. Nature 463, 893–898 (2010)
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011)
Nakahata, S. & Kawamoto, S. Tissue-dependent isoforms of mammalian Fox-1 homologs are associated with tissue-specific splicing activities. Nucleic Acids Res. 33, 2078–2089 (2005)
Curtin, J. A. et al. Distinct sets of genetic alterations in melanoma. N. Engl. J. Med. 353, 2135–2147 (2005)
Jané-Valbuena, J. et al. An oncogenic role for ETV1 in melanoma. Cancer Res. 70, 2075–2084 (2010)
Garraway, L. A. et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436, 117–122 (2005)
Forbes, S. A. et al. COSMIC (the Catalogue of Somatic Mutations in Cancer): a resource to investigate acquired mutations in human cancer. Nucleic Acids Res. 38, D652–D657 (2010)
Reva, B., Antipin, Y. & Sander, C. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res. 39, e118 (2011)
Flaherty, K. T., Hodi, F. S. & Bastian, B. C. Mutation-driven drug development in melanoma. Curr. Opin. Oncol. 22, 178–183 (2010)
Flaherty, K. T. et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N. Engl. J. Med. 363, 809–819 (2010)
Chapman, P. B. et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N. Engl. J. Med. 364, 2507–2516 (2011)
Solit, D. B. & Rosen, N. Resistance to BRAF inhibition in melanomas. N. Engl. J. Med. 364, 772–774 (2011)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Acknowledgements
Illumina sequencing was performed at the Broad Institute and array-based genomic characterization and functional studies were performed at the Belfer Institute of DFCI. We are grateful to the Broad Institute Genome Sequencing Platform, Genome Analysis Platform and Biological Samples Platform. This work was supported by the National Human Genome Research Institute (S.B.G., E.S.L.), National Cancer Institute (M.M., L.C.), FWF-Austrian Science Fund (S.N.W.), NIH Director’s New Innovator Award (L.A.G.), Melanoma Research Alliance (L.A.G., L.C.), Starr Cancer Consortium (L.A.G.) and the Burroughs-Wellcome Fund (L.A.G.).
Author information
Authors and Affiliations
Contributions
M.F.B., E.H., T.P.H. and Y.L.D. are lead authors; E.S.L., G.G., L.C. and L.A.G. are senior authors. M.F.B. and E.H. performed the genomic analysis of the whole-genome sequencing data. T.P.H. and Y.L.D. performed the molecular biology and mouse experiments to interrogate the function of PREX2. E.I., P.G., R.Z. and M.A.S. participated in the functional experiments. M.S.L. performed genomic analysis of mutations and rearrangements. A.P. and X.R. performed FISH studies of PREX2. H.Z, K.C, A.Y.S., T.F., S.L.C., Y.D., P.S., D.V., R.J., G.S., A.H.R., T.J.P. and N.S. provided additional computational analyses. A.S. and D.S. contributed melanoma short-term cultures for the extension cohort. I.R.W. contributed clinical information. C.S., R.O., W.W., S.M., D.A. and K.A. participated in DNA sample processing and quality control. E.N. played a project management role. M.F.B., M.S.L., R.O., M.P., L.A., W.W., G.G. and L.A.G. validated candidate rearrangements. J.B. and S.B.G. oversaw the generation of DNA sequence data. J.B., M.M., S.B.G. and T.R.G contributed to the study design and interpretation of data. S.N.W. contributed most melanoma tumours for the discovery and extension cohort and participated in the study design. J.W. and N.W. also contributed tumour material for the discovery cohort. E.S.L., G.G., L.C. and L.A.G. conceived of and designed the study and participated in the data analysis and interpretation. M.F.B., T.P.H., L.C. and L.A.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains a list of Supplementary Figures and Tables, Supplementary Methods, Supplementary Figures 1-6 and Supplementary References. (PDF 2143 kb)
Supplementary Tables
This file contains Supplementary Tables 1-12. (XLS 13173 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Berger, M., Hodis, E., Heffernan, T. et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 485, 502–506 (2012). https://doi.org/10.1038/nature11071
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11071
This article is cited by
-
Cooperative function of oncogenic MAPK signaling and the loss of Pten for melanoma migration through the formation of lamellipodia
Scientific Reports (2024)
-
Molecular profiling and prognostic biomarkers in chinese non-small cell lung cancer cohort
Diagnostic Pathology (2023)
-
Integrated DNA and RNA sequencing reveals early drivers involved in metastasis of gastric cancer
Cell Death & Disease (2022)
-
Evolving Treatment Approaches to Mucosal Melanoma
Current Oncology Reports (2022)
-
Targeted Therapy for Melanomas Without BRAF V600 Mutation
Current Oncology Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.