Abstract
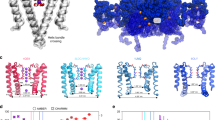
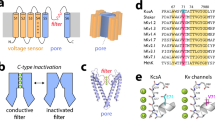
The coupled interplay between activation and inactivation gating is a functional hallmark of K+ channels1,2. This coupling has been experimentally demonstrated through ion interaction effects3,4 and cysteine accessibility1, and is associated with a well defined boundary of energetically coupled residues2. The structure of the K+ channel KcsA in its fully open conformation, in addition to four other partial channel openings, richly illustrates the structural basis of activation–inactivation gating5. Here, we identify the mechanistic principles by which movements on the inner bundle gate trigger conformational changes at the selectivity filter, leading to the non-conductive C-type inactivated state. Analysis of a series of KcsA open structures suggests that, as a consequence of the hinge-bending and rotation of the TM2 helix, the aromatic ring of Phe 103 tilts towards residues Thr 74 and Thr 75 in the pore-helix and towards Ile 100 in the neighbouring subunit. This allows the network of hydrogen bonds among residues Trp 67, Glu 71 and Asp 80 to destabilize the selectivity filter6,7, allowing entry to its non-conductive conformation. Mutations at position 103 have a size-dependent effect on gating kinetics: small side-chain substitutions F103A and F103C severely impair inactivation kinetics, whereas larger side chains such as F103W have more subtle effects. This suggests that the allosteric coupling between the inner helical bundle and the selectivity filter might rely on straightforward mechanical deformation propagated through a network of steric contacts. Average interactions calculated from molecular dynamics simulations show favourable open-state interaction-energies between Phe 103 and the surrounding residues. We probed similar interactions in the Shaker K+ channel where inactivation was impaired in the mutant I470A. We propose that side-chain rearrangements at position 103 mechanically couple activation and inactivation in KcsA and a variety of other K+ channels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Panyi, G. & Deutsch, C. Cross talk between activation and slow inactivation gates of Shaker potassium channels. J. Gen. Physiol. 128, 547–559 (2006)
Sadovsky, E. & Yifrach, O. Principles underlying energetic coupling along an allosteric communication trajectory of a voltage-activated K+ channel. Proc. Natl Acad. Sci. USA 104, 19813–19818 (2007)
Baukrowitz, T. & Yellen, G. Use-dependent blockers and exit rate of the last ion from the multi-ion pore of a K+ channel. Science 271, 653–656 (1996)
Lopez-Barneo, J., Hoshi, T., Heinemann, S. H. & Aldrich, R. W. Effects of external cations and mutations in the pore region on C-type inactivation of Shaker potassium channels. Receptors Channels 1, 61–71 (1993)
Cuello, L., Jogini, V. & Perozo, E. Structural mechanism of C-type inactivation in K+ channnels. Nature 10.1038/nature09153 (this issue) (2010)
Cordero-Morales, J. F. et al. Molecular determinants of gating at the potassium-channel selectivity filter. Nature Struct. Mol. Biol. 13, 311–318 (2006)
Cordero-Morales, J. F. et al. Molecular driving forces determining potassium channel slow inactivation. Nature Struct. Mol. Biol. 14, 1062–1069 (2007)
Armstrong, C. M. Voltage-gated K channels. Sci. STKE 2003, re10 (2003)
Yellen, G. The voltage-gated potassium channels and their relatives. Nature 419, 35–42 (2002)
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002)
Liu, Y., Holmgren, M., Jurman, M. E. & Yellen, G. Gated access to the pore of a voltage-dependent K+ channel. Neuron 19, 175–184 (1997)
Perozo, E., Cortes, D. M. & Cuello, L. G. Structural rearrangements underlying K+-channel activation gating. Science 285, 73–78 (1999)
Choi, K. L., Aldrich, R. W. & Yellen, G. Tetraethylammonium blockade distinguishes two inactivation mechanisms in voltage-activated K+ channels. Proc. Natl Acad. Sci. USA 88, 5092–5095 (1991)
Hoshi, T., Zagotta, W. N. & Aldrich, R. W. Biophysical and molecular mechanisms of Shaker potassium channel inactivation. Science 250, 533–538 (1990)
Kiss, L., LoTurco, J. & Korn, S. J. Contribution of the selectivity filter to inactivation in potassium channels. Biophys. J. 76, 253–263 (1999)
Demo, S. D. & Yellen, G. Ion effects on gating of the Ca2+-activated K+ channel correlate with occupancy of the pore. Biophys. J. 61, 639–648 (1992)
Swenson, R. P. Jr & Armstrong, C. M. K+ channels close more slowly in the presence of external K+ and Rb+. Nature 291, 427–429 (1981)
Alagem, N., Yesylevskyy, S. & Reuveny, E. The pore helix is involved in stabilizing the open state of inwardly rectifying K+ channels. Biophys. J. 85, 300–312 (2003)
Chapman, M. L., Blanke, M. L., Krovetz, H. S. & Vandongen, A. M. Allosteric effects of external K+ ions mediated by the aspartate of the GYGD signature sequence in the Kv2.1 K+ channel. Pflügers Arch. 451, 776–792 (2006)
Proks, P., Capener, C. E., Jones, P. & Ashcroft, F. M. Mutations within the P-loop of Kir6.2 modulate the intraburst kinetics of the ATP-sensitive potassium channel. J. Gen. Physiol. 118, 341–353 (2001)
Schulte, U., Weidemann, S., Ludwig, J., Ruppersberg, J. & Fakler, B. K+-dependent gating of Kir1.1 channels is linked to pH gating through a conformational change in the pore. J. Physiol. (Lond.) 534, 49–58 (2001)
Ben-Abu, Y., Zhou, Y., Zilberberg, N. & Yifrach, O. Inverse coupling in leak and voltage-activated K+ channel gates underlies distinct roles in electrical signaling. Nature Struct. Mol. Biol. 16, 71–79 (2009)
Monod, J., Wyman, J. & Changeux, J. P. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965)
Chakrapani, S., Cordero-Morales, J. F. & Perozo, E. A quantitative description of KcsA gating I: macroscopic currents. J. Gen. Physiol. 130, 465–478 (2007)
Liu, Y., Jurman, M. E. & Yellen, G. Dynamic rearrangement of the outer mouth of a K+ channel during gating. Neuron 16, 859–867 (1996)
Ader, C. et al. Coupling of activation and inactivation gate in a K+-channel: potassium and ligand sensitivity. EMBO J. 28, 2825–2834 (2009)
Rotem, D., Mason, A. & Bayley, H. Inactivation of the KcsA potassium channel explored with heterotetramers. J. Gen. Physiol. 135, 29–42 (2010)
Olcese, R., Sigg, D., Latorre, R., Bezanilla, F. & Stefani, E. A conducting state with properties of a slow inactivated state in a Shaker K+ channel mutant. J. Gen. Physiol. 117, 149–163 (2001)
Chen, J., Seebohm, G. & Sanguinetti, M. C. Position of aromatic residues in the S6 domain, not inactivation, dictates cisapride sensitivity of HERG and eag potassium channels. Proc. Natl Acad. Sci. USA 99, 12461–12466 (2002)
Klement, G., Nilsson, J., Arhem, P. & Elinder, F. A tyrosine substitution in the cavity wall of a K channel induces an inverted inactivation. Biophys. J. 94, 3014–3022 (2008)
Cuello, L. G., Romero, J. G., Cortes, D. M. & Perozo, E. pH-dependent gating in the Streptomyces lividans K+ channel. Biochemistry 37, 3229–3236 (1998)
Cortes, D. M. & Perozo, E. Structural dynamics of the Streptomyces lividans K+ channel (SKC1): oligomeric stoichiometry and stability. Biochemistry 36, 10343–10352 (1997)
Liu, Y. S., Sompornpisut, P. & Perozo, E. Structure of the KcsA channel intracellular gate in the open state. Nature Struct. Biol. 8, 883–887 (2001)
Cuello, L. G., Cortes, D. M. & Perozo, E. Molecular architecture of the KvAP voltage-dependent K+ channel in a lipid bilayer. Science 306, 491–495 (2004)
Delcour, A. H., Martinac, B., Adler, J. & Kung, C. Modified reconstitution method used in patch-clamp studies of Escherichia coli ion channels. Biophys. J. 56, 631–636 (1989)
Cortes, D. M., Cuello, L. G. & Perozo, E. Molecular architecture of full-length KcsA: role of cytoplasmic domains in ion permeation and activation gating. J. Gen. Physiol. 117, 165–180 (2001)
Zhou, Y., Morais-Cabral, J. H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 Å resolution. Nature 414, 43–48 (2001)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–332 (1997)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Morais-Cabral, J. H., Zhou, Y. & MacKinnon, R. Energetic optimization of ion conduction rate by the K+ selectivity filter. Nature 414, 37–42 (2001)
Brooks, B. R. et al. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983)
Bernèche, S. & Roux, B. Molecular dynamics of the KcsA K+ channel in a bilayer membrane. Biophys. J. 78, 2900–2917 (2000)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
Jogini, V. & Roux, B. Dynamics of the Kv1.2 voltage-gated K+ channel in a membrane environment. Biophys. J. 93, 3070–3082 (2007)
Acknowledgements
We thank F. Bezanilla, H Mchaourab and R. Nakamoto for discussions and comments on the manuscript. We also thank K. Locher for comments on the manuscript. R. Mackinnon provided the Fab-expressing hybridoma cells. We thank the members of the Perozo laboratory for comments on the manuscript. We thank K. R. Rajashankar, R. Sanishvili and the staff at the NE-CAT 24ID and GM-CA 23ID beamlines at the Advanced Photon Source, Argonne National Laboratory for assistance during data collection. This work was supported in part by NIH grant R01-GM57846 and a gift from the Palmer family to E.P., by grant R01-GM62342 to B.R. and by an NRSA postdoctoral fellowship to A.C.P.
Author information
Authors and Affiliations
Contributions
E.P. and L.G.C. conceived the project. L.G.C. and D.M.C. generated constructs, performed biochemical analysis, and expressed, purified and crystallized the proteins. L.G.C. performed EPR experiments. L.G.C., V.J. and E.P. collected X-ray diffraction data. L.G.C. and V.J. determined and analysed the structures. A.C.P. and B.R. performed computation analysis. O.D. and L.G.C. conducted FRET measurements. S.C. measured inactivation in truncated KcsA and in Rb+ ions. J.F.C.-M. measured E71H single-channel activity and made the F103A mutation. L.G.C. measured F103X mutant series electrophysiology. D.G.G. made electrophysiology measurements in Shaker channels. E.P., L.G.C. and V.J. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and Supplementary Figures S1-S10 with legends. (PDF 1962 kb)
Supplementary Movie 1
This movie shows coupling between F103-I100-T74/T75 during C-type inactivation. (MPG 3599 kb)
Rights and permissions
About this article
Cite this article
Cuello, L., Jogini, V., Cortes, D. et al. Structural basis for the coupling between activation and inactivation gates in K+ channels. Nature 466, 272–275 (2010). https://doi.org/10.1038/nature09136
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09136
This article is cited by
-
Calcium-gated potassium channel blockade via membrane-facing fenestrations
Nature Chemical Biology (2024)
-
The structural basis of divalent cation block in a tetrameric prokaryotic sodium channel
Nature Communications (2023)
-
Structures of the T cell potassium channel Kv1.3 with immunoglobulin modulators
Nature Communications (2022)
-
Ion-dependent structure, dynamics, and allosteric coupling in a non-selective cation channel
Nature Communications (2021)
-
Correlated conformational dynamics of the human GluN1-GluN2A type N-methyl-D-aspartate (NMDA) receptor
Journal of Molecular Modeling (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.