Abstract
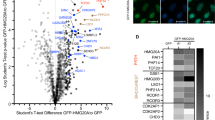
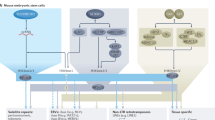
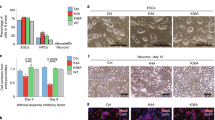
Diverse histone modifications are catalysed and recognized by various specific proteins, establishing unique modification patterns that act as transcription signals1,2. In particular, histone H3 trimethylation at lysine 36 (H3K36me3) is associated with actively transcribed regions and has been proposed to provide landmarks for continuing transcription3,4; however, the control mechanisms and functions of H3K36me3 in higher eukaryotes are unknown. Here we show that the H3K36me3-specific histone methyltransferase (HMTase) Wolf–Hirschhorn syndrome candidate 1 (WHSC1, also known as NSD2 or MMSET) functions in transcriptional regulation together with developmental transcription factors whose defects overlap with the human disease Wolf–Hirschhorn syndrome (WHS)5,6. We found that mouse Whsc1, one of five putative Set2 homologues2,7,8, governed H3K36me3 along euchromatin by associating with the cell-type-specific transcription factors Sall1, Sall4 and Nanog in embryonic stem cells, and Nkx2-5 in embryonic hearts, regulating the expression of their target genes. Whsc1-deficient mice showed growth retardation and various WHS-like midline defects, including congenital cardiovascular anomalies. The effects of Whsc1 haploinsufficiency were increased in Nkx2-5 heterozygous mutant hearts, indicating their functional link. We propose that WHSC1 functions together with developmental transcription factors to prevent the inappropriate transcription that can lead to various pathophysiologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sims, R. J. & Reinberg, D. Histone H3 Lys 4 methylation: caught in a bind? Genes Dev. 20, 2779–2786 (2006)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007)
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007)
Hirschhorn, K. & Cooper, H. L. Chromosomal aberrations in human disease. A review of the status of cytogenetics in medicine. Am. J. Med. 31, 442–470 (1961)
Bergemann, A. D., Cole, F. & Hirschhorn, K. The etiology of Wolf–Hirschhorn syndrome. Trends Genet. 21, 188–195 (2005)
Strahl, B. D. et al. Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol. Cell. Biol. 22, 1298–1306 (2002)
Sun, X. J. et al. Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J. Biol. Chem. 280, 35261–35271 (2005)
Stec, I. et al. WHSC1, a 90 kb SET domain-containing gene, expressed in early development and homologous to a Drosophila dysmorphy gene maps in the Wolf-Hirschhorn syndrome critical region and is fused to IgH in t(4;14) multiple myeloma. Hum. Mol. Genet. 7, 1071–1082 (1998)
Marango, J. et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood 111, 3145–3154 (2008)
Kohlhase, J., Wischermann, A., Reichenbach, H., Froster, U. & Engel, W. Mutations in the SALL1 putative transcription factor gene cause Townes-Brocks syndrome. Nature Genet. 18, 81–83 (1998)
Shafi, R. et al. The O-GlcNAc transferase gene resides on the X chromosome and is essential for embryonic stem cell viability and mouse ontogeny. Proc. Natl Acad. Sci. USA 97, 5735–5739 (2000)
Sakaki-Yumoto, M. et al. The murine homolog of SALL4, a causative gene in Okihiro syndrome, is essential for embryonic stem cell proliferation, and cooperates with Sall1 in anorectal, heart, brain and kidney development. Development 133, 3005–3013 (2006)
Wu, Q. et al. Sall4 interacts with Nanog and co-occupies Nanog genomic sites in embryonic stem cells. J. Biol. Chem. 281, 24090–24094 (2006)
Liang, J. et al. Nanog and Oct4 associate with unique transcriptional repression complexes in embryonic stem cells. Nature Cell Biol. 10, 731–739 (2008)
Carrozza, M. J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005)
Li, B. et al. Combined action of PHD and chromo domains directs the Rpd3S HDAC to transcribed chromatin. Science 316, 1050–1054 (2007)
Li, M. et al. Solution structure of the Set2-Rpb1 interacting domain of human Set2 and its interaction with the hyperphosphorylated C-terminal domain of Rpb1. Proc. Natl Acad. Sci. USA 102, 17636–17641 (2005)
Liu, K. J., Arron, J. R., Stankunas, K., Crabtree, G. R. & Longaker, M. T. Chemical rescue of cleft palate and midline defects in conditional GSK-3β mice. Nature 446, 79–82 (2007)
Bruneau, B. G. The developmental genetics of congenital heart disease. Nature 451, 943–948 (2008)
Prall, O. W. et al. An Nkx2-5/Bmp2/Smad1 negative feedback loop controls heart progenitor specification and proliferation. Cell 128, 947–959 (2007)
Hiroi, Y. et al. Tbx5 associates with Nkx2-5 and synergistically promotes cardiomyocyte differentiation. Nature Genet. 28, 276–280 (2001)
Moses, K. A., DeMayo, F., Braun, R. M., Reecy, J. L. & Schwartz, R. J. Embryonic expression of an Nkx2-5/Cre gene using ROSA26 reporter mice. Genesis 31, 176–180 (2001)
Biben, C. et al. Cardiac septal and valvular dysmorphogenesis in mice heterozygous for mutations in the homeobox gene Nkx2-5. Circ. Res. 87, 888–895 (2000)
Koshiba-Takeuchi, K. et al. Cooperative and antagonistic interactions between Sall4 and Tbx5 pattern the mouse limb and heart. Nature Genet. 38, 175–183 (2006)
Gromak, N. et al. The PTB interacting protein raver1 regulates α-tropomyosin alternative splicing. EMBO J. 22, 6356–6364 (2003)
Nimura, K. et al. Dnmt3a2 targets endogenous Dnmt3L to ES cell chromatin and induces regional DNA methylation. Genes Cells 11, 1225–1237 (2006)
Hayashi, M. et al. Comparative roles of Twist-1 and Id1 in transcriptional regulation by BMP signaling. J. Cell Sci. 120, 1350–1357 (2007)
Ura, K. & Kaneda, Y. Reconstitution of chromatin in vitro. Methods Mol. Biol. 181, 309–325 (2001)
Tachiwana, H., Osakabe, A., Kimura, H. & Kurumizaka, H. Nucleosome formation with the testis-specific histone H3 variant, H3t, by human nucleosome assembly proteins in vitro. Nucleic Acids Res. 36, 2208–2218 (2008)
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998)
Dorigo, B., Schalch, T., Bystricky, K. & Richmond, T. J. Chromatin fiber folding: requirement for the histone H4 N-terminal tail. J. Mol. Biol. 327, 85–96 (2003)
Saeki, H. et al. Linker histone variants control chromatin dynamics during early embryogenesis. Proc. Natl Acad. Sci. USA 102, 5697–5702 (2005)
Nishioka, K. & Reinberg, D. Methods and tips for the purification of human histone methyltransferases. Methods 31, 49–58 (2003)
Seo, H. et al. Rapid generation of specific antibodies by enhanced homologous recombination. Nature Biotechnol. 23, 731–735 (2005)
Wade, P. A. et al. Mi-2 complex couples DNA methylation to chromatin remodelling and histone deacetylation. Nature Genet. 23, 62–66 (1999)
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotechnol. 17, 1030–1032 (1999)
Bruneau, B. G. et al. A murine model of Holt–Oram syndrome defines roles of the T-box transcription factor Tbx5 in cardiogenesis and disease. Cell 106, 709–721 (2001)
Acknowledgements
We thank T. Richmond, R. Nishinakamura, H. Kurumizaka and M. Shirai for providing reagents and mice; S. Khochbin, S. Hirose, J. Godde and J. Takeuchi for their critical reading of the manuscript; and H. Niwa, H. Hamada, M. Yamamoto, S. Ohishi, H. Nakagami and members of the GTS laboratory for discussion and support. This work was supported by grants from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (MEXT) and the Naito Foundation, and by funds from Osaka University for female researchers.
Author Contributions Y.K. provided support and general guidance for this work. K.U. planned and organized the project. K.N. designed and performed experiments. H.S. performed histological analysis of hearts. R.J.S., M.I. and M.O. contributed mouse resources. K.U. wrote the paper with K.N. and Y.K.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S16 with Legends, Supplementary Table 1 and Supplementary References. (PDF 1951 kb)
Rights and permissions
About this article
Cite this article
Nimura, K., Ura, K., Shiratori, H. et al. A histone H3 lysine 36 trimethyltransferase links Nkx2-5 to Wolf–Hirschhorn syndrome. Nature 460, 287–291 (2009). https://doi.org/10.1038/nature08086
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08086
This article is cited by
-
Diverse and dynamic forms of gene regulation by the S. cerevisiae histone methyltransferase Set1
Current Genetics (2023)
-
Dual function of SF3B2 on chromatin and RNA to regulate transcription in head and neck squamous cell carcinoma
Cell & Bioscience (2022)
-
Interplay between chromatin marks in development and disease
Nature Reviews Genetics (2022)
-
Multiple myeloma with high-risk cytogenetics and its treatment approach
International Journal of Hematology (2022)
-
The role of NSD1, NSD2, and NSD3 histone methyltransferases in solid tumors
Cellular and Molecular Life Sciences (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.