Abstract
The neuronal repressor REST (RE1-silencing transcription factor; also called NRSF) is expressed at high levels in mouse embryonic stem (ES) cells1, but its role in these cells is unclear. Here we show that REST maintains self-renewal and pluripotency in mouse ES cells through suppression of the microRNA miR-21. We found that, as with known self-renewal markers, the level of REST expression is much higher in self-renewing mouse ES cells than in differentiating mouse ES (embryoid body, EB) cells. Heterozygous deletion of Rest (Rest+/-) and its short-interfering-RNA-mediated knockdown in mouse ES cells cause a loss of self-renewal—even when these cells are grown under self-renewal conditions—and lead to the expression of markers specific for multiple lineages. Conversely, exogenously added REST maintains self-renewal in mouse EB cells. Furthermore, Rest+/- mouse ES cells cultured under self-renewal conditions express substantially reduced levels of several self-renewal regulators, including Oct4 (also called Pou5f1), Nanog, Sox2 and c-Myc, and exogenously added REST in mouse EB cells maintains the self-renewal phenotypes and expression of these self-renewal regulators. We also show that in mouse ES cells, REST is bound to the gene chromatin of a set of miRNAs that potentially target self-renewal genes. Whereas mouse ES cells and mouse EB cells containing exogenously added REST express lower levels of these miRNAs, EB cells, Rest+/- ES cells and ES cells treated with short interfering RNA targeting Rest express higher levels of these miRNAs. At least one of these REST-regulated miRNAs, miR-21, specifically suppresses the self-renewal of mouse ES cells, corresponding to the decreased expression of Oct4, Nanog, Sox2 and c-Myc. Thus, REST is a newly discovered element of the interconnected regulatory network that maintains the self-renewal and pluripotency of mouse ES cells.
Similar content being viewed by others
Main
REST is believed to be a major transcriptional repressor of neurogenesis2,3,4,5, and activation of REST target genes was found to be sufficient to convert neural stem/progenitor cells to neuronal phenotypes6,7. However, REST activity seems to depend on the cellular context; for example, REST can show both an oncogenic8,9,10 and tumour-suppressor function5 as well as involvement in haematopoietic and cardiac differentiation3,4,5. Embryonic stem (ES) cells are pluripotent cells that have the potential for both indefinite self-renewal and differentiation into all three germ layers of the body11. Here we provide evidence that REST has a unique role as a protector of self-renewal and pluripotency in mouse ES cells, corresponding to the expression of critical regulators such as Oct4, Nanog, Sox2 and c-Myc.
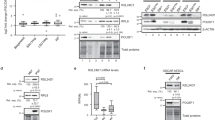
We began by assessing the levels of REST protein in mouse ES cells growing under self-renewal conditions and differentiation conditions (Fig. 1a; ES and EB, respectively). As expected, western blotting showed that the ES cells had higher levels of REST expression and of the representative markers of self-renewal (proteins Oct4, Sox2 and c-Myc) than did the EB cells, indicating that REST expression is associated with self-renewal. To determine whether REST regulates the self-renewal of mouse ES cells, we took two approaches using two cell lines—YHC334 (YHC) and RRC160 (RRC) (see Supplementary Information). In the first approach, we performed polymerase chain reaction with reverse transcription (RT–PCR; Fig. 1b), quantitative RT–PCR (qRT–PCR; Fig. 1c) using different primer sets than those used for regular RT–PCR, and western blotting (Fig. 1d), all of which showed that both of the Rest+/- mouse ES cell lines (YHC and RRC) had substantially lower levels of Rest transcript and protein than the parental mouse ES cells. Alkaline phosphatase assays12, performed to measure self-renewal within these cells, revealed that the levels of self-renewal in both of the Rest+/- mouse ES cell lines were less than half that in the parental mouse ES cells (Fig. 1e, f).
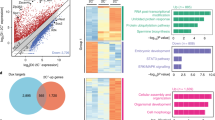
a, EB cells (ES cells grown in the absence of LIF under non-adherent conditions for 4 days) showed reduced levels of self-renewal markers and REST compared with ES cells (grown in the presence of LIF under adherent conditions). Western blot analysis of whole-cell extracts prepared from ES cells and EB cells showed an association between expression of REST and self-renewal markers. Actin was used as a loading control. b–f, Mouse ES cell lines with heterozygous deletion of Rest (RRC and YHC) show loss of self-renewal. b, c, RT–PCR (b) and qRT–PCR (c) analyses with different primer sets of total RNA isolated from ES, RRC and YHC cells show reduced Rest transcripts in Rest+/- cells. Gapdh was used as a loading control. *, P < 0.0001. The values are represented as mean ± s.d. (n = 3). d, Western blot analysis of whole-cell extracts from ES, RRC and YHC cells shows reduced levels of REST protein in Rest+/- cells. α-Tubulin was used as a loading control. e, Alkaline phosphatase staining of ES colonies shows loss of self-renewal in Rest+/- cells as compared with wild-type ES cells. f, Percentages of self-renewing colonies of ES, RRC and YHC cells calculated after alkaline phosphatase assays when cultured under self-renewing conditions show significant reductions in the self-renewal capacity of both Rest+/- cell lines compared with ES cells. *, P < 0.0001. The error bars correspond to three replicates (n = 3) and show mean ± s.d. g–i, siRNA-mediated knockdown of REST causes loss of self-renewal in mouse ES cells. g, Specific knockdown of targeted genes was achieved using siRNA. qRT–PCR of total RNA purified from mouse ES cells treated with siRest or siOct4 shows knockdown of specific genes. Analysis was performed 5 days after transfection. *, P < 0.0001. The values are represented as mean ± s.d. (n = 3). h, Western blotting shows reduced REST protein levels in siRest-treated cells compared with control (NT siRNA). i, siRest- and siOct4-treated cells show less self-renewal than NT-siRNA-treated cells. Mouse ES cell colonies were screened by alkaline phosphatase assays. *, P < 0.0001; **, P < 0.001. The error bars correspond to three replicates (n = 3) and show mean ± s.d. j, Exogenously added Rest, but not GFP, maintained self-renewal in mouse ES cells cultured under differentiation conditions. Mouse ES cells were transfected with plasmids encoding GFP or REST and grown in the absence of LIF. Percentages of self-renewing colonies from three independent experiments were averaged after alkaline phosphatase assay and are shown for each transfected gene. ***, P < 0.01. The error bars correspond to three replicates (n = 3) and show mean ± s.d.
In the second approach, done to rule out any adaptive response accumulated by Rest+/- mouse ES cell lines, we determined whether the decreased self-renewal seen in the Rest+/- mouse ES cells could also be seen in mouse ES cells treated with short interfering RNA (siRNA) against Rest messenger RNA (siRest). qRT–PCR assays, done to confirm that siRNA-treated cells actually had decreased Rest and Oct4 levels, showed a 50% knockdown of Rest and Oct4, respectively, in both siRest- and siOct4-treated cells (Fig. 1g). This was further confirmed by western blotting (Fig. 1h). Self-renewal assays of these cells (Fig. 1i) then showed that whereas mouse ES cells treated with non-targeting control (NT) siRNA showed almost no change in self-renewal (94.5%), mouse ES cells treated with siRest showed about a 50% decrease in self-renewal (see Supplementary Fig. 1 for the effect of four different siRest constructs on the self-renewal capacity of mouse ES cells). SiOct4-treated mouse ES cells also showed a decrease in self-renewal (47%), consistent with the role of Oct4 in mouse ES cell self-renewal13. Thus, siRest-mediated knockdown of Rest transcripts, and not nonspecific siRNA activity, caused decreased self-renewal in mouse ES cells, a result similar to that seen in the Rest+/- mouse ES cell lines.
Gain-of-function experiments (Fig. 1j) in which self-renewing mouse ES cells were transfected with plasmids encoding a neomycin resistance gene and either green fluorescent protein (GFP) or REST12 showed that cells transfected with GFP showed very little maintenance of self-renewal, whereas cells transfected with REST showed approximately 55% maintenance of self-renewal. Taken together, these results further confirmed the role of REST in maintaining self-renewal of mouse ES cells.
To ascertain the lineages into which Rest+/- haploinsufficient mouse ES cells differentiate on their exit from REST-mediated self-renewal, we performed RT–PCR (Fig. 2a) and qRT–PCR (Fig. 2b) assays of parental mouse ES cells, YHC and RRC cells growing under self-renewal conditions in the presence of leukaemia inhibitory factor (LIF). Rest+/- cells showed activation of markers specific for mesoderm, endoderm, ectoderm and trophectoderm. Expression of all of these differentiation markers was absent or detectable at only very low levels in self-renewing parental mouse ES cells. The results in RRC cells (data not shown) and YHC cells were very similar. Notably, although the neuronal marker gene Calbindin is a direct target of REST1, the other differentiation markers are not known to be direct REST targets, suggesting that REST can maintain the pluripotency of mouse ES cells partly by directly repressing the expression of some differentiation genes and partly by an indirect mechanism in which the non-REST-target differentiation genes are repressed as a consequence of the REST-mediated maintenance of self-renewal.
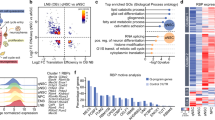
a, b, RT–PCR (a) and qRT–PCR (b) analyses of total RNA isolated from ES and YHC cells grown under self-renewing conditions were performed using primers specific for markers for ectoderm, mesoderm, endoderm and trophectoderm. Gapdh was used as an internal control. *, P = 0.001; **, P = 0.003; ***, P = 0.006; ****, P = 0.012. The values are represented as mean ± s.d. (n = 3). c, EB cells derived from YHC cells show lineage markers that are either absent or significantly reduced in EB cells derived from wild-type mouse ES cells. RT–PCR analysis of total RNA isolated from wild-type EB cells and YHC EB cells was performed using specific primers for various differentiated lineage genes (shown above each lane). Gapdh was used as a loading control. d, qRT–PCR analysis of total RNA isolated from mouse ES cells treated with either non-targeting siRNA (NT) or siRest was performed using primers specific for different lineage markers. Analysis was performed 5 days after transfection. *, P < 0.0001; **, P = 0.001; ***, P = 0.002; ****, P = 0.004; *****, P = 0.005; ******, P = 0.01. The values are represented as mean ± s.d. (n = 3). e, The inner cell mass of blastocysts showed co-expression of REST and self-renewal markers Oct4, Nanog and Sox2. Blastocysts were stained for nuclei (4,6-diamidino-2-phenylindole (DAPI), blue), REST (red) and self-renewal markers (green). Merged and differential interference contrast (DIC) images are also shown. Scale bar, 25 μm.
The differentiation of Rest+/- haploinsufficient cells into multiple lineages growing under self-renewal conditions prompted us to determine the differentiation characteristics of EB cells derived from wild-type as well as Rest+/- ES cells. RT–PCR of Rest+/- EB cells detected the expression of markers for all lineages, including ectoderm, mesoderm, endoderm and trophectoderm (Fig. 2c). Some of these markers were either absent or markedly reduced in wild-type ES-cell-derived EB cells. Thus, although both wild-type and Rest+/- cells started expressing multiple differentiation markers in EB cells, the Rest+/--derived cells expressed a greater number of lineage markers than did the EB cells derived from wild-type mouse ES cells, suggesting that REST deficiency predisposes mouse ES cells to differentiation. To determine whether siRest-treated mouse ES cells also differentiate into various lineages, we performed a qRT–PCR assay with these cells growing under self-renewal conditions (Fig. 2d). The results indicated that siRest indeed increased the expression of various lineage markers. Notably, the exact level of expression of each differentiation marker varied between siRest-treated and Rest+/- mouse ES cells (Fig. 2b), presumably because of the differences in the methods used to reduce the REST levels in these cells (that is, whereas Rest+/- cells may show additional adaptive responses as a result of being cultured for longer than the siRest-treated mouse ES cells, the latter show more immediate effects of REST knockdown). Nonetheless, the overall results indicated that on exit from REST-mediated self-renewal, both siRest-treated wild type and Rest+/- mouse ES cells were able to differentiate into various lineages.
The inner cell mass of the blastocyst has been shown to harbour self-renewing cells that express proteins such as Nanog, Oct4 and Sox2. To determine whether REST is expressed in blastocysts, we subjected mouse blastocysts to double-immunofluorescence analysis using anti-REST antibodies and antibodies against Nanog, Oct4, or Sox2. Consistent with its role in the self-renewal of mouse ES cells, REST was co-expressed with these self-renewal regulators in the blastocyst inner cell mass (Fig. 2e) (see Supplementary Information for a discussion on the role of REST in cells of the inner cell mass).
When we subjected the parental mouse ES cells and Rest+/- (YHC and RRC) haploinsufficient cells growing under self-renewal conditions to RT–PCR assays to examine the transcript levels of various self-renewal markers, we found that the decreased REST levels in Rest+/- cells corresponded with decreased transcript levels of several other self-renewal genes, such as Oct4, Nanog, Sox2, Tbx3 and c-myc (Fig. 3a), suggesting that REST protects the expression of these self-renewal genes. Western blot analysis of the same cells also indicated that Rest+/- cells have substantially reduced levels of c-Myc and Oct4 proteins compared with the parental mouse ES cells (Fig. 3b), indicating that the downregulation of REST negatively affects self-renewal signals. To confirm further the role of REST in maintaining the self-renewal signals, we performed the same experiment shown in Fig. 1j, in which exogenously added REST maintained the self-renewal of mouse ES cells growing under differentiation conditions, and then analysed the expression of c-Myc and Oct4 by western blotting. As shown in Fig. 3c, the exogenously added REST, but not GFP, maintained c-Myc and Oct4 protein levels in mouse ES cells growing under differentiation conditions. Taken together, these results indicate that REST protects the expression of several self-renewal genes.
a, b, Heterozygous deletion of Rest results in decreased expression of self-renewal genes. a, RT–PCR analysis of total RNA from ES, RRC and YHC cells. Genes are indicated on the left side of each panel. Gapdh was used as a loading control. b, Western blot analysis of whole-cell extracts from ES, RRC and YHC cells. Antibodies specific for REST, c-Myc and Oct4 proteins were used for the analysis. α-Tubulin was used as a loading control. c, Exogenously added Rest, but not GFP, maintained the expression of c-Myc and Oct4 in ES cells growing without LIF. Shown is a western blot analysis of whole-cell extracts from mouse ES cells transfected with plasmids encoding GFP or REST and then grown in the absence of LIF. Antibodies specific for REST, c-Myc and Oct4 proteins were used for the analysis. α-Tubulin was used as a loading control. d–h, REST suppresses the expression of miRNAs, which are associated with the absence of self-renewal regulators. d, e, ChIP (d) and quantitative ChIP (e) assays showed that REST binds to the gene chromatin of a set of miRNAs in ES cells (1, input; 2, IgG, negative control; 3, anti-H3 antibody, positive control; 4, anti-REST antibody; + and - represent predicted RE-1 binding sites occupied and unoccupied by REST, respectively). *, P < 0.0001; **, P = 0.001. The values of three replicates are represented as mean ± s.d. f, qRT–PCR analysis of ES, YHC, RRC, EB and EB plus REST showed that expression of the miRNAs (shown in d and e) was lower in ES than in YHC, RRC and EB cells. The higher expression of miRNAs in EB cells could be extinguished in a gain-of-function experiment in the presence of exogenously added REST. *, P < 0.0001. The values are represented as mean ± s.d. (n = 3). g, The expression of the miRNAs shown in d was upregulated in a loss-of-function experiment when mouse ES cells were treated with siRest but not when they are treated with non-targeting siRNA (NT). *, P < 0.0001. The values are represented as mean ± s.d. (n = 3). Analysis was performed 3 days after transfection. h, siRest-mediated knockdown of REST that produced increased expression of the miRNAs in g corresponded to the decreased expression of Rest and the known self-renewal genes Oct4, Nanog and Sox2. A qRT–PCR assay with mouse ES cells treated with siRest and NT siRNA is shown. Analysis was performed 5 days after transfection. *, P < 0.0001. The values are represented as mean ± s.d. (n = 3).
To determine how REST maintains the expression of multiple self-renewal genes, we examined the possible role of miRNAs in the self-renewal of REST-mediated mouse ES cells. We examined the miRNA profiles of wild-type and Rest+/- mouse ES cells growing under self-renewal conditions. The miRNAs from self-renewing wild-type mouse ES cells fell into two major groups, representing expressed and repressed miRNAs (Supplementary Fig. 1 and Supplementary Table 1). Rest+/- YHC and RRC cell lines had similar miRNA profiles, which were strikingly opposite to those of wild-type mouse ES cells. Furthermore, miR-124a, which was recently shown to be a target of REST14, and miR-106a and miR-106b, whose predicted targets include Rest, were upregulated in Rest+/- cells, suggesting that regulation involving double-negative feedback loops may exist to maintain homeostasis. Using miRNA databases (http://cbio.mskcc.org/mirnaviewer and http://pictar.bio.nyu.edu/), we found a set of miRNAs that can potentially target self-renewal genes such as Nanog, Sox2, Tbx3 and c-myc (Supplementary Table 2). These miRNAs were present only at low levels in self-renewing mouse ES cells and were present at higher levels in both the Rest+/- heterozygous cells and EB cells. To determine whether REST directly binds to the regulatory sequences (chromatin) of these miRNA-containing genes, we determined such potential REST binding sites (Supplementary Table 3) using the MatInspector module of the Genomatix database15. We found several potential REST-binding sites for each of these genes. Chromatin immunoprecipitation assays (Fig. 3d) and quantitative chromatin immunoprecipitation assays (Fig. 3e) in ES cells further revealed that REST was bound only to specific sites of the gene chromatin for each of these miRNAs. In a gain-of-function experiment similar to those shown in Figs 1j and 3c, in which exogenously added REST restored the self-renewal function of mouse ES cells growing under differentiation conditions, we analysed the expression of the miRNAs in these cells by qRT–PCR. As shown in Fig. 3f, levels of these miRNAs were compared with self-renewing mouse ES cells, expressed at higher levels in both Rest+/- heterozygous cells and EB cells, and expressed at lower levels in EB cells transfected with exogenous REST. To confirm further REST-mediated repression of the miRNAs, we performed a loss-of-function experiment in which we treated mouse ES cells with siRest and analysed the expression of the miRNAs by qRT–PCR. As shown, siRest-mediated knockdown of REST resulted in increased levels of miRNA expression when compared with non-targeting siRNA (Fig. 3g). To confirm that the siRest-mediated knockdown of REST, which increased the expression of the miRNAs, also decreased the expression of the self-renewal genes, we performed a qRT–PCR assay. As shown in Fig. 3h, siRest-treated mouse ES cells showed decreased levels of Rest, Oct4, Nanog and Sox2 expression. Thus, taken together, these results indicate that REST represses this set of miRNAs, and strongly suggest that at least some of the miRNAs, in turn, interfere with the expression of critical self-renewal regulators such as Oct4, Nanog and Sox2.
To determine whether the REST-regulated miRNAs are directly involved in maintaining self-renewal, we performed self-renewal assays in mouse ES cells after transfecting cells with individual precursors of these miRNAs. At least one of these precursor miRNAs, pre-miR-21, markedly decreased the self-renewing capacity of mouse ES cells by 60%, compared with that of an NT siRNA. In contrast, pre-miR-26a did not effectively alter the self-renewal efficiency when compared with the same NT siRNA (Fig. 4a). To confirm that the decreased self-renewal seen in mouse ES cells was actually due to the intracellular increase in the corresponding miRNAs, we measured levels of miR-21 and miR-26a by qRT–PCR. Indeed, the pre-miR-transfected cells showed more than tenfold and 300-fold increases in miR-21 and miR-26a expression, respectively (Fig. 4b).
a, Addition of exogenous precursor miR-21 (pre-miR-21), but not non-targeting siRNA (NT) or pre-miR-26a, caused a decrease of self-renewal in mouse ES cells. *, P < 0.0001. The error bars correspond to three replicates (n = 3) and show mean ± s.d. b, Addition of exogenous pre-miR-21 and exogenous pre-miR-26a, but not NT siRNA, to mouse ES cells resulted in increased levels of their corresponding miRNA expression. Results of qRT–PCR assays performed with NT-siRNA-, pre-miR-26a- and pre-miR-21-transfected cells 1 day after transfection are shown. *, P < 0.0001. The values are represented as mean ± s.d. (n = 3). c, The effect of miR-21 on loss of self-renewal was specific. Addition of exogenous pre-miR-21 along with anti-miR-21 rescued the loss of self-renewal by pre-miR-21 alone. *, P < 0.0001 (there was no significant difference between NT siRNA and pre-miR-21 plus anti-miR-21 values). The error bars correspond to three replicates (n = 3) and show mean ± s.d. d, Addition of exogenous pre-miR-21, but not NT siRNA, to mouse ES cells resulted in loss of self-renewal markers Oct4, Nanog, Sox2 and c-myc. qRT–PCR after transfection of mouse ES cells with pre-miR-21 and NT siRNA was performed 5 days after transfection to detect the expression levels of the self-renewal markers. *, P < 0.0001; **, P = 0.002; ***, P = 0.007; ****, P = 0.008. The values are represented as mean ± s.d. (n = 3). e, Proposed model of REST-mediated maintenance of self-renewal through miR-21. Rest maintains self-renewal and pluripotency of mouse ES cells by suppressing miR-21 expression.
To confirm that the effect of pre-miR-21 on self-renewal was specific, we transfected mouse ES cells with NT siRNA or a mixture of pre-miR-21 plus either NT siRNA or anti-miR-21 and performed self-renewal assays. As shown in Fig. 4c, pre-miR-21 plus NT siRNA lowered the self-renewal capacity by 30% when compared with NT siRNA alone. This lowering of self-renewal by 50 nM pre-miR-21 was half of what we had observed with 100 nM pre-miR-21 (Fig. 4a), indicating that under these conditions, 50–100 nM pre-miR-21 was in the linear range of its activity in suppressing self-renewal in mouse ES cells. Furthermore, the suppression of self-renewal by pre-miR-21 was rescued by anti-miR-21 but not by NT siRNA, indicating that the lowering of self-renewal was pre-miR-21-specific. We then performed qRT–PCR assays to determine the expression levels of the self-renewal markers Oct4, Nanog, Sox2 and c-myc in pre-mir-21-treated cells. The results showed a corresponding decrease in the expression levels of these markers in pre-mir-21-treated cells (Fig. 4d). This is the first evidence of a single miRNA as a regulator of self-renewal in mouse ES cells. Interestingly, an earlier report indicated that miR-21 was one of the miRs whose expression was higher in differentiated compared with undifferentiated ES cells16. Taken together, our results indicate that REST-repressed miR-21 regulates the self-renewal of mouse ES cells. Thus, we report that REST has a newly discovered role in maintaining self-renewal and pluripotency of mouse ES cells. Furthermore, although REST can perform this function partly by direct repression of some of its target differentiation genes, such as calbindin, we provide evidence that REST functions via repressing the transcription of a specific miRNA, miR-21, which suppresses self-renewal by means of a corresponding loss of expression of the critical self-renewing regulators Oct4, Nanog, Sox2 and c-Myc (Fig. 4e). Whether miR-21 directly suppresses the expression of these self-renewing regulators, thereby inhibiting self-renewal, or whether miR-21 inhibits self-renewal by a different mechanism, with consequent suppression of these self-renewal regulators, is unclear. Of note, bioinformatics-predicted miR-21-binding sites are present in the mRNAs of Sox2 and Nanog but not those of Oct4 and c-myc (Supplementary Table 2). If these sites truly function as miR-21 target sites, and if miR-21 target sites are indeed not present in Oct4 and c-myc mRNAs, then these observations indicate that the miR-21-mediated regulation of mouse ES cell self-renewal involves a regulatory cascade initiated by the loss of Sox2 and Nanog expression that leads to the loss of expression of Oct4 and c-Myc. It is noteworthy that Oct4, Nanog and Sox2 were found to co-occupy Rest promoter/enhancer sequences, apparently as an activator complex17. Furthermore, REST formed part of the Nanog–Oct4 complex18 and was predicted to be a major component of an Oct4–Sox2–Nanog network in ES cells19. Taken together with our data presented here, these observations indicate that REST is a new element of the interconnected regulatory network required to maintain the self-renewal and pluripotency of cultured mouse ES cells.
Methods Summary
Embryonic stem cells (E14Tg2a) were cultured in the presence of LIF on gelatin-coated dishes. Embryoid bodies were grown on bacterial dishes in the absence of LIF and β-mercaptoethanol. Alkaline phosphatase staining after loss- and gain of function was carried out using a detection kit from Chemicon. Chromatin immunoprecipitation (ChIP) assays were performed as described in the Methods. For the miRNA microarrays, total RNA was prepared using TRIzol, and RNA samples were analysed by LC Sciences on their microarray platform with a probe set based on Sanger version 9.0. Blastocysts were obtained from C57BL/6 NCr mice, fixed in 4% paraformaldehyde, permeabilized with 0.25% Triton X-100, and were then double immunostained with antibodies against REST, Sox2, Nanog and Oct4. Primers used for RT–PCR and ChIP are available in the Methods.
Online Methods
ES cell lines and culture conditions
Mouse ES cells (E14Tg2A) obtained from Bay Genomics were cultured under self-renewal conditions (in the presence of LIF, cultured on adherent tissue culture dishes) without feeder cells in 1,000 units ml-1 LIF (ESGRO) on gelatin-coated tissue culture dishes20. Embryoid bodies in the absence of LIF and β-mercaptoethanol on non-adherent tissue culture dishes were grown for 3 days in bacterial dishes as described1. The Rest+/- cell lines YHC334 (YHC) and RRC160 (RRC) that have gene traps at Rest exons 1 and 2, respectively, were obtained from Bay Genomics. Both of these lines were derived from parental E14Tg2A mouse ES cells, the same cells that we used for all of our wild-type mouse ES cell experiments.
ChIP assays
ChIP assays of cultured mouse ES cells and EBs were performed as described21. We used antibodies against mouse IgG (2 μl; catalogue number I 5381, Sigma), REST (2 μl; 07-579, Upstate) and histone H3 (2 μl; ab1791, Abcam). One microgram of each antibody and approximately 15 micrograms of chromatin were added to each reaction mixture and incubated overnight at 4 °C. The mixtures were then incubated with 33 μl of protein A-agarose beads (Amersham Pharmacia Biotech AB; pre-equilibrated with 1 mg ml-1 bovine serum albumin and 0.3 mg ml-1 salmon sperm DNA).
The PCR primer sequences used are as follows: miR-93/miR-106b forward 5′-CCCCCAAAACCAGTATCCTT-3′, reverse 5′-TCGTACTTCCCGGATCACTC-3′; miR-27a forward 5′-AAATCCCCAGAAGCTGGAGT-3′, reverse 5′-AGCACTTGAAAGGCAAAGGA-3′; miR-21 forward 5′-AGGGCAGGAAGATGACACAC-3′, reverse 5′-GGGCTGATGAGCACTAAGGA-3′; miR-26a forward 5′-GCCTAACCCAAGAAGGGAAA-3′, reverse 5′-TCCCTCTCATCTGGACAACC-3′; miR-152 forward 5′-TGGTCTCTGTCCAGCACAAC-3′, reverse 5′-CAGGTCACAGCTGCACTCAT-3′; miR-124a forward 5′-CTCTGCGTGTTCACAGCGG-3′, reverse 5′-CTCTTGGCATTCACCGCGTG-3′.
Quantitative ChIP PCR was performed using the same extracts used above. Briefly, qPCR was performed and analysed using real-time PCR (ABI prism 7500). Primers were designed by Primer Express 2.0 (Applied Biosystems) to amplify 60- to 150-bp amplicons. Amplicons were measured by SYBR Green fluorescence (SYBR Green Master mix, Applied Biosystems) in 10-μl reactions. Reactions were performed in triplicates. The amount of product was determined relative to a standard curve of input chromatin. Melting curves showed that PCRs yielded single product. Ct values for each sample were normalized against that of input DNA and percentage recovery is plotted. The primers used for qPCR will be available on request.
Western blotting, RT–PCR and miRNA microarray
Whole-cell extracts were prepared, and approximately 10 μg of proteins were resolved on SDS–PAGE using antibodies against REST (07-579, Upstate), c-Myc (sc-764, Santa Cruz Biotechnology), Oct4 (ab19857, Abcam), Sox2 (ab15830, Abcam), actin (sc-1616, Santa Cruz) and α-tubulin (MMS-407R, Covance), following the manufacturers’ recommendations. Total RNA was extracted using TRIzol (Invitrogen), and approximately 50 ng of total RNA was used as a template. RT–PCR was performed using a OneStep RT–PCR kit (Qiagen).
The following gene-specific primers were designed: Rest forward 5′-AGCGAGTACCACTGGAGGAA-3′, reverse 5′-CTGAATGAGTCCGCATGTGT-3′; calbindin forward 5′-GCTTCTATCTGGCGGAAGG-3′, reverse 5′-TGTCATCTGGCTACCTTCCC-3′; Pitx2 forward 5′-CGGCAGAGGACTCATTTCAC-3′, reverse 5′-GTACGAATAGCCGGGGTACA-3′; Snai1 forward 5′-CTTGTGTCTGCACGACCTGT-3′, reverse 5′-CTTCACATCCGAGTGGGTTT-3′; Sox18 forward 5′-AACAAAATCCGGATCTGCAC-3′, reverse 5′-CGAGGCCGGTACTTGTAGTT-3′; Cebpa forward 5′-CCGACTTCTACGAGGTGGAG-3′, reverse 5′-TGGCCTTCTCCTGCTGTC-3′. The following primers were based on published sequences: Gapdh, Oct4, Nanog, Gata6, Fgf5, brachyury, Flk1 and Pl1 (ref. 22); Mash1, Mash2, β-globin, Cd34, Cxcl12, Ehox, Eomes, Gata4, Isl1, Meox1, Mixl1, Ngn2, Shh and Tbx3 (ref. 23).
For quantitative RT–PCR, first-strand synthesis was performed using Applied Biosystems’ high-capacity cDNA reverse transcription kit (4368814). Real-time PCR was carried out using cDNAs with SYBR Green PCR master mix from Applied Biosystems (4309155). Reactions were carried out in triplicates using ABI prism 7500 and the analysis was performed similar to that described for quantitative ChIP PCR above, except that ΔCt was obtained after normalization with Ct of Gapdh for mouse-specific primers and that of actin for human-specific primers. Mouse-specific primers will be provided on request; human-specific primers are as published24.
For the miRNA microarrays, total RNA was prepared using TRIzol, and RNA samples were analysed by LC Sciences on their microarray with a probe set based on Sanger version 9.0. Multiple sample analysis involves normalization, data adjustment, t-test/analysis of variance (ANOVA) analysis, and clustering. Normalization is carried out using a cyclic Lowess (locally weighted regression) method. The normalization is to remove system-related variations, such as sample amount variations, different labelling dyes, and signal gain differences of scanners so that biological variations can be faithfully revealed. Data adjustment includes data filtering, log2 transformation, and gene centring and normalization. The data filtering removes genes (or miRNAs) with (normalized) intensity values below a threshold value of 32 across all samples. The log2 transformation converts intensity values into the log2 scale. Gene centring and normalization transform the log2 values using the mean and the standard deviation of individual genes across all samples using the following formula:

In certain cases, expression ratios of dual-sample arrays are used in clustering analysis. Gene normalization would then be performed using the following formula:

The t-test is performed between ‘control’ and ‘test’ sample groups, with each group containing at least two samples. t-values are calculated for each miRNA, and P-values are computed from the theoretical t-distribution. MicroRNAs with P-values below a critical P-value (typically 0.01) are selected for cluster analysis. The clustering is done using a hierarchical method and is performed with average linkage and euclidean distance metric.
ANOVA is an extension of the t-test to more than two experimental conditions. It picks out miRNAs that have significant differences in means across three or more groups of samples, with each group containing at least two samples. In this analysis, P-values are computed from the F-distribution. In certain cases, such as time point experiments, in which only one sample is collected and assayed for each condition or time point, repeating sets of reporters (or probes) on each array may be used as a ‘group’ in an ANOVA analysis.
All data processes, except clustering plot, were carried out using in-house developed computer programs. The clustering plot is generated using TIGR MeV (multiple experimental viewer) software from The Institute for Genomic Research.
miRNA validation was carried out using Ambion’s mirVana qRT–PCR miRNA detection kit (AM1558). mirVana qRT–PCR primer sets for the miRNA of interest were also obtained from Ambion (30000–30999). Reactions were performed as per the manufacturer’s recommendations. The values obtained were normalized with primer set corresponding to 5S rRNA (AM 30302).
RNAi-mediated knockdown, exogenous expression of miR-21, anti-miR-21 and gain-of-function experiments
RNAi-mediated knockdown of REST and Oct4 were carried out per the manufacturer’s protocol (Dharmacon). siRest (60 pmol), siOct4 (60 pmol) and non-targeting siRNA (60 pmol) were transfected using the Amaxa nucleofection method. Colonies were assayed for alkaline phosphatase activity 3 days after transfection.
Amaxa nucleofection was also used to deliver precursor miR-21 and anti-miR-21 molecules into mouse ES cells. Transfection was carried out using 100 pmol of precursor or 50 pmol of miR-21 and anti-miR-21 each using 4 × 106 cells. Non-targeting siRNA was used as a control. Delivery and mature miR-21 product was inspected in cells by qRT–PCR 1 day after transfection, as described above.
For the gain-of-function experiments, mouse ES cells were transfected with 2 μg of plasmid DNA using Lipofectamine 2000. The Flag–REST construct was a gift from G. Mandel. The transfected cells were selected with G418 (125 μg ml-1) under self-renewing conditions in the presence of LIF for 3 days. After selection, we grew the cells under differentiation conditions in the absence of LIF for 4 days, and subjected them to self-renewal assays. An alkaline phosphatase assay was then performed using an alkaline phosphatase detection kit (Chemicon) after 4 days. Percentages of self-renewing colonies from three independent experiments were plotted. Standard deviation was calculated and is depicted by error bars in the figures.
Immunofluorescence microscopy of blastocysts
Blastocysts from C57BL/6 NCr mice were fixed in 4% paraformaldehyde, permeabilized with 0.25% Triton X-100, and blocked with 1% goat serum in PBS25. The blastocysts were then double immunostained with mouse anti-REST antibody (raised against the amino-terminal portion of human REST, A372) in combination with antibodies against Sox2 (Abcam, AB15830), Oct4 (Abcam, AB19857), Nanog (Abcam, AB21603) followed by Alexa-Fluor-488-conjugated secondary antibody detecting mouse IgG and Fluor-555-conjugated secondary antibody detecting rabbit IgG (Molecular Probes). Images were captured with a Nikon microscope (Nikon Eclipse TE 2000U).
References
Ballas, N., Grunseich, C., Lu, D. D., Speh, J. C. & Mandel, G. REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell 121, 645–657 (2005)
Ballas, N. & Mandel, G. The many faces of REST oversee epigenetic programming of neuronal genes. Curr. Opin. Neurobiol. 15, 500–506 (2005)
Ooi, L. & Wood, I. C. Chromatin crosstalk in development and disease: lessons from REST. Nature Rev. Genet. 8, 544–554 (2007)
Coulson, J. M. Transcriptional regulation: cancer, neurons and the REST. Curr. Biol. 15, R665–R668 (2005)
Majumder, S. REST in good times and bad: roles in tumor suppressor and oncogenic activities. Cell Cycle 5, 1929–1935 (2006)
Su, X., Kameoka, S., Lentz, S. & Majumder, S. Activation of REST/NRSF target genes in neural stem cells is sufficient to cause neuronal differentiation. Mol. Cell. Biol. 24, 8018–8025 (2004)
Watanabe, Y. et al. Conversion of myoblasts to physiologically active neuronal phenotype. Genes Dev. 18, 889–900 (2004)
Lawinger, P. et al. The neuronal repressor REST/NRSF is an essential regulator in medulloblastoma cells. Nature Med. 6, 826–831 (2000)
Fuller, G. N. et al. Many human medulloblastoma tumors overexpress repressor element-1 silencing transcription (REST)/neuron-restrictive silencer factor, which can be functionally countered by REST-VP16. Mol. Cancer Ther. 4, 343–349 (2005)
Su, X. et al. Abnormal expression of REST/NRSF and Myc in neural stem/progenitor cells causes cerebellar tumors by blocking neuronal differentiation. Mol. Cell. Biol. 26, 1666–1678 (2006)
Pan, G. & Thomson, J. A. Nanog and transcriptional networks in embryonic stem cell pluripotency. Cell Res. 17, 42–49 (2007)
Pritsker, M., Ford, N. R., Jenq, H. T. & Lemischka, I. R. Genomewide gain-of-function genetic screen identifies functionally active genes in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 6946–6951 (2006)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Conaco, C., Otto, S., Han, J. J. & Mandel, G. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc. Natl Acad. Sci. USA 103, 2422–2427 (2006)
Cartharius, K. et al. MatInspector and beyond: promoter analysis based on transcription factor binding sites. Bioinformatics 21, 2933–2942 (2005)
Houbaviy, H. B., Murray, M. F. & Sharp, P. A. Embryonic stem cell-specific microRNAs. Dev. Cell 5, 351–358 (2003)
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005)
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006)
Zhou, Q., Chipperfield, H., Melton, D. A. & Wong, W. H. A gene regulatory network in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 104, 16438–16443 (2007)
Nichols, J., Evans, E. P. & Smith, A. G. Establishment of germ-line-competent embryonic stem (ES) cells using differentiation inhibiting activity. Development 110, 1341–1348 (1990)
Cui, R. et al. Family members p53 and p73 act together in chromatin modification and direct repression of α-fetoprotein transcription. J. Biol. Chem. 280, 39152–39160 (2005)
Kaji, K. et al. The NuRD component Mbd3 is required for pluripotency of embryonic stem cells. Nature Cell Biol. 8, 285–292 (2006)
Pritsker, M., Ford, N. R., Jenq, H. T. & Lemischka, I. R. Genomewide gain-of-function genetic screen identifies functionally active genes in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 6946–6951 (2006)
Cai, J. et al. Assessing self-renewal and differentiation in human embryonic stem cell lines. Stem Cells 24, 516–530 (2006)
Zhang, J. et al. Sall4 modulates embryonic stem cell pluripotency and early embryonic development by the transcriptional regulation of Pou5f1 . Nature Cell Biol. 8, 1114–1123 (2006)
Acknowledgements
We would like to thank the National Institutes of Health for funding (grants CA97124 and CA81255 to S.M.). M.N.K. and S.K.S. were both recipients of the Dodie Hawn Fellowship in Cancer Genetics.
Author Contributions S.K.S. and M.N.K. participated in all aspects including manuscript preparation; J.P.-T. conducted blastocyst experiments; H.A. carried out immunofluorescence studies; S.M. did project planning and manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Methods, Supplementary Discussion with additional references, Supplementary Figure 1 and Supplementary Tables 1-3. The Supplementary information consists of the detailed methods with references. A discussion on the possible role of REST in the blastocyst is included. MiRNA profile is indicated in Supplementary Figure 1 and the values are shown in Table 1. Table 2 shows prediction of miRNA target sites for self renewal genes. Table 3 shows predicted NRSE sites on various miRNAs. Word File. Size: 444 KB. (PDF 190 kb)
Rights and permissions
About this article
Cite this article
Singh, S., Kagalwala, M., Parker-Thornburg, J. et al. REST maintains self-renewal and pluripotency of embryonic stem cells. Nature 453, 223–227 (2008). https://doi.org/10.1038/nature06863
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06863
This article is cited by
-
CRISPR/Cas9-based genome-wide screening of the deubiquitinase subfamily identifies USP3 as a protein stabilizer of REST blocking neuronal differentiation and promotes neuroblastoma tumorigenesis
Journal of Experimental & Clinical Cancer Research (2023)
-
The Potential of miR-21 in Stem Cell Differentiation and its Application in Tissue Engineering and Regenerative Medicine
Stem Cell Reviews and Reports (2023)
-
REST in the Road Map of Brain Development
Cellular and Molecular Neurobiology (2023)
-
High glucose and palmitic acid induces neuronal senescence by NRSF/REST elevation and the subsequent mTOR-related autophagy suppression
Molecular Brain (2022)
-
CSN6–TRIM21 axis instigates cancer stemness during tumorigenesis
British Journal of Cancer (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.