Abstract
Self-perpetuating changes in the conformations of amyloidogenic proteins play vital roles in normal biology and disease. Despite intense research, the architecture and conformational conversion of amyloids remain poorly understood. Amyloid conformers of Sup35 are the molecular embodiment of the yeast prion known as [PSI], which produces heritable changes in phenotype through self-perpetuating changes in protein folding. Here we determine the nature of Sup35's cooperatively folded amyloid core, and use this information to investigate central questions in prion biology. Specific segments of the amyloid core form intermolecular contacts in a ‘Head-to-Head’, ‘Tail-to-Tail’ fashion, but the ‘Central Core’ is sequestered through intramolecular contacts. The Head acquires productive interactions first, and these nucleate assembly. Variations in the length of the amyloid core and the nature of intermolecular interfaces form the structural basis of distinct prion ‘strains’, which produce variant phenotypes in vivo. These findings resolve several problems in yeast prion biology and have broad implications for other amyloids.
Similar content being viewed by others
Main
Prion proteins share an unusual property: they adopt distinct functional and conformational states that self-perpetuate through protein-conformational chain reactions1,2,3,4. The first known prion, PrP, facilitates the transmission of a fatal neurodegenerative disease in mammals (spongiform encephalopathy) by converting non-prion conformers to the prion state4,5. Fungal prions, however, are not generally pathogenic. Instead, they act as protein-only elements of inheritance. Their prion conformers produce new phenotypes—often beneficial phenotypes3—by changing processes as diverse as translation termination, nitrogen metabolism and heterokaryon formation6. These phenotypes are heritable because mother cells pass prion conformers on to their daughters, perpetuating the cycle of conversion1. In the marine snail Aplysia, a neuronal form of CPEB, a protein implicated in long-term memory7, can also switch to a self-perpetuating prion conformation. In this case, conformational switching activates the protein, suggesting that CPEB's self-perpetuating prion conformation functions in the long-term maintenance of synapses8. Most proteins can form amyloids under specific, unusual conditions9. Prion proteins access such states under normal physiological conditions, and some have been conserved for hundreds of millions of years10. Prions play a much broader role in biology than initially suspected11,12,13.
To understand these self-perpetuating conformational changes, we investigated [PSI+], a highly conserved prion in Saccharomyces cerevisiae. [PSI+] confers a wide variety of novel phenotypes by facilitating the read-through of nonsense codons11,12,13. Read-through occurs when Sup35, a translation termination factor, is inactivated by conversion to an amyloid with a self-sustaining structure1,2,6.
Sup35 has three distinct regions14,15: C, a GTP-binding domain at the carboxy terminus; M, a highly charged middle region; and N, a glutamine/asparagine-rich amino-terminal region containing oligopeptide repeats. C facilitates translation termination, and N and M govern prion status. N is essential for converting Sup35 to the prion state in vivo16 and for converting soluble protein into amyloid fibres in vitro17. M confers solubility in the non-prion state and stabilizes the prion during mitosis and meiosis18. When N and M are removed from C and fused to the glucocorticoid receptor, they create a new prion that confers a hormone-response phenotype on yeast but otherwise recapitulates all of the unusual physical and genetic behaviours of [PSI+]19. Thus, NM encodes prion function.
In vitro, the conversion of natively unfolded NM to β-rich amyloid fibres involves (1) a lag phase, in which part of the protein oligomerizes and converts to an amyloidogenic nucleus, and (2) an assembly phase, in which soluble proteins rapidly associate with mature nuclei and convert to amyloid17,20,21,22,23,24. NM can form distinct types of self-perpetuating conformers17,25. These produce distinct phenotypes (prion strains or variants) when used to transform non-prion [psi-] cells to the prion [PSI+] state26,27. Thus, NM amyloids also embody prion structure.
The β-strands in NM fibres run perpendicular to the main axis and are spaced ∼4.7 Å apart20,28. Several mutations in NM are known to affect prion maintenance and fibre assembly17,18,29,30. However, we do not yet know (1) the arrangement of individual NM molecules within the amyloid fibre, (2) the mechanisms of nucleation and conformational conversion, or (3) the structural basis of prion strains. To address these questions, we capitalized on the absence of cysteine residues in NM to create a large number of variants, each containing a single cysteine at a different location, which could be modified with fluorophores and crosslinkers.
Cysteine variants behave like wild-type NM
We created 37 individual cysteine-substitution mutations throughout NM (Fig. 1a). Each was used to replace the NM portion of the wild-type SUP35 gene in vivo, resulting in a single, full-length functional SUP35 gene. All NM variants retained the capacity to support the [PSI+] and [psi-] states (Supplementary Fig. S1a). None altered the stability of those states (data not shown).
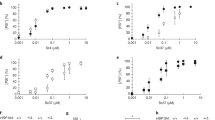
a, NM sequence, showing residues mutated to cysteine in green, the five imperfect oligopeptide repeats shaded in grey, and the boundary between N and M (123/124) where it is traditionally drawn. On the basis of amino acid composition, the boundary might equally be drawn at amino acids 137–139. b, Accessibility of cysteine residues after assembly. Fibres formed from single-cysteine substitution mutants were labelled with pyrene maleimide for 3 h. The ratio of labelled-to-unlabelled protein is plotted as a measure of accessibility. For all proteins, labelling approached 100% in denaturant (6 M GdmCl; data not shown). c, Denaturation profiles. Mutants were individually assembled at 25 °C, using 75% unlabelled and 25% acrylodan-labelled protein. Each preparation was then equilibrated for 30 min at 24 different concentrations of GdmCl and fluorescence spectra were recorded.
Next, each protein was expressed in and purified from Escherichia coli. All spontaneously assembled into amyloid at the same rate as wild-type (Supplementary Fig. S1b). Moreover, all fibres were indistinguishable from wild-type by electron microscopy and SDS (sodium dodecyl sulphate) solubility22 (data not shown). Finally, fibres made from each mutant seeded assembly as well as the wild type17,20 (Supplementary Fig. S1c). Having established that cysteine-substitution proteins recapitulate prion behaviour both in vivo and in vitro, we used them to explore NM structure and assembly.
Boundaries of the cooperatively folded amyloid
We used two independent approaches to investigate the general structure of NM fibres. First, in 37 sets of fibres assembled at 25 °C from each variant, we probed the accessibility of cysteine residues to labelling with pyrene maleimide, and to a more hydrophilic reagent, Lucifer yellow (Fig. 1b and data not shown). Proteins carrying a cysteine residue between amino acid residues 25 and 58 were sparsely labelled with either pyrene maleimide or Lucifer yellow. Proteins with a cysteine residue in amino acids 2–21 and 68–112 showed partial accessibility. All cysteines in M were highly accessible.
Next, we used acrylodan to report on the conformational status of different segments of NM. Acrylodan shows an increase in fluorescence intensity and a blue shift in λmax when sequestered from solvent. When denatured, all cysteine-labelled proteins had a λmax around 530 nm. After assembly at 25 °C, proteins labelled with acrylodan at a cysteine between residues 21 and 121 had strongly blue-shifted emissions (λmax 486–488 nm), indicating sequestration from solvent (no guanidine hydrochloride (GdmCl) in Fig. 1c and data not shown). Proteins labelled in regions adjoining residues 21–121 (residue 7 N-terminally and residues 137, 158 and 167 C-terminally) had partially blue-shifted emission maxima (λmax 493–525 nm). Proteins labelled at residues 2, 184, 225 or 234 had no significant blue shift, indicating that these residues remained exposed to solvent pre- and post-assembly (Fig. 1c and data not shown).
To determine which residues participate in the same cooperatively folded structure, we assessed their post-assembly GdmCl denaturation profiles, using 24 different GdmCl concentrations for each of the fibres. All fibres labelled between residues 21 and 121 showed a similar drop in fluorescence intensity and a corresponding red shift in emission maxima, with an inflection at 2.5 M ± 0.15 M GdmCl (Fig. 1c). These profiles fitted a monophasic unfolding transition that corresponds to the major unfolding transition of wild-type NM protein (A. Cashikar and S. L. L., unpublished results). Adjacent cysteine variants that had intermediate blue shifts upon fibrillization also had distinct unfolding transitions in GdmCl.
These studies establish that (1) a large portion of N is sequestered from solvent; (2) a sub-portion of N constitutes a distinct domain (formed by contiguous amino acids, including 21–121) with an unusually stable structure and a single cooperative unfolding transition; (3) flanking sequences are structurally heterogeneous, with residues 137 and 158 having distinct, but cooperative, unfolding transitions, and residues 2 and 7 having multiphasic transitions; and (4) M is flexible and solvent exposed from residue 158 onwards. Residues in the N/M transition zone (amino acids 121, 137 and 158) were fully accessible to post-assembly cysteine labelling but according to acrylodan fluorescence and GdmCl denaturation were partially sequestered and structured. This probably reflects different sensitivities of the two techniques used to detect structural instability. For example, if residue 121 occasionally adopts an open structure, it would show a strong blue shift with acrylodan but still be accessible to prolonged labelling.
Identifying intermolecular contacts
To determine which regions of NM make intermolecular contacts in fibres, we exploited the ability of pyrene-labelled proteins to form excimers (excited-state dimers). When two pyrene molecules lie within 4–10 Å of each other, the long fluorescence lifetime of pyrene allows an excited residue to interact with an unexcited residue before energy is emitted. This produces a strong red shift in fluorescence. Excimer fluorescence will occur between residues at β-strands that form the interface between two monomers, but not between residues that are distant from the interface. One caveat is that the pyrenes might alter the structure formed, but control experiments eliminated this concern (see Supplementary Fig. S2c).
In denaturant, each of the pyrene-labelled proteins had multiple emission maxima between 384 and 405 nm (Supplementary Fig. S2a, b). After assembly at 25 °C, most fibres labelled in the N region showed a blue shift in fluorescence (Supplementary Fig. S2b), indicating that they were sequestered from intermolecular contacts. Six different fibre preparations individually labelled at a residue in one of two distinct regions (residues 25, 31 or 38, or residues 91, 96 or 106) produced strong red-shifted fluorescence (λmax ∼465 nm; see Fig. 2a and Supplementary Fig. S2a). These residues must lie at or near a contact between two NM molecules. We will refer to these intermolecular contact regions as the ‘Head’ (residues 25–38) and ‘Tail’ (residues 91–106). Residues between them, the ‘Central Core’ (43–85), are also part of the cooperatively folded amyloid but are sequestered from intermolecular contacts.
a, Proximity analysis assessed by excimer fluorescence in fibres carrying pyrene labels at single sites. Fibres were assembled at 25 °C with 2.5 µM NM (60% labelled), either by gently rotating the samples (black circles), or by adding 4% sonicated seed (grey squares) formed at 25 °C. The ratio of excimer fluorescence to non-excimer fluorescence (I465/I375) is plotted. b, Excimer fluorescence in fibres assembled from mixtures of two different pyrene-labelled cysteine variants. Fibres were assembled at 25 °C with gentle rotation, in reactions containing equimolar mixtures of each variant (25% labelled and 75% unlabelled). Actual excimer values are provided in Supplementary Fig. S3. Head and Tail region residues are marked with lines. c, Effects of crosslinks on amyloid assembly. Proteins crosslinked under denaturing conditions with either a 2-Å disulphide bond (black bars) or a flexible 11-Å BMB crosslink (grey bars) were diluted 200-fold into buffer, to a final concentration of 2.5 µM NM, and assembled with rotation at 25 °C. Thioflavin T (ThT) fluorescence values for wild-type fibres, soluble NM, and a denatured standard (bovine serum albumin (BSA) treated with DTT to induce denaturation) are also shown. Values represent mean ± s.d. (n = 3 experiments).
Pyrene fluorescence patterns were virtually identical in repeat experiments (data not shown). They were also nearly identical in seeded and unseeded reactions (grey squares and black circles, respectively; Fig. 2a). The remarkable reproducibility of these excimer patterns establishes that the intermolecular contacts in seeded assembly quite precisely recapitulate those of spontaneous assembly.
Next, we mixed two proteins, each individually labelled at one cysteine residue, in all possible pair-wise combinations (Fig. 2b and Supplementary Fig. S3). Confirming that residues in the Central Core do not contribute to intermolecular contacts, all fibres in which either one or both of the proteins were labelled in this region produced low excimer signals (residues 51, 58 and 73; see Fig. 2b and Supplementary Fig. S3). The strongest signals were again from the Head and Tail regions, but only when both proteins were labelled in the same region (Fig. 2b, orange boxes). Fibres with one protein labelled in the Head and another in the Tail produced weak signals. We conclude that contacts between monomers in the fibre occur in a Head-to-Head and Tail-to-Tail fashion.
These data are not compatible with the parallel, super-pleated sheet model for the Sup35 prion domain31, in which individual NM molecules fold into long serpentine arrays, stacked in parallel along their entire length. They are, however, compatible with a β-helix model28 and other models in which a contiguous stretch of amino acids forms a cooperative amyloid fold, and a Central Core within this region is sequestered from intermolecular contacts. Furthermore, individual subunits must form contacts in a Head-to-Head and Tail-to-Tail fashion (see Fig. 3).
a–d, One model for NM assembly is provided to illustrate the constraints that our data place on the nature of NM fibre structures. In the cooperatively folded amyloid core, Head residues (red) in one NM molecule are in close proximity to Head residues of their neighbours; the same is true for the Tail residues (green) (c). Central Core residues (blue) are sequestered from intermolecular interactions. M (dashed line) is largely unstructured, but the segment of M proximal to N becomes structured when amyloid forms. In the initial stages, NM molecules rapidly acquire a collapsed state (a) that retains a molten character until the Head regions of two molecules (b) come into proximity with each other and nucleate assembly. We do not know how fibrils are arranged within fibres (d).
Constraints on inter-subunit relationships
To provide an independent assessment of inter-subunit interactions, we introduced two types of crosslinks into each of the individual cysteine-substituted proteins under denaturing conditions. Reaction with oxidized dithiothreitol (DTT) produced disulphide bonds with a bond length of ∼2 Å. Reaction with 1,4-bis-maleimidobutane (BMB), a homobifunctional agent, produced crosslinks with a 10.9-Å flexible linker.
Disulphide crosslinks inhibited fibre formation at every position tested between residues 21 and 121 (black bars in Fig. 2c; thioflavinT fluorescence levels equivalent to those of BSA aggregates). Disulfides in the extreme N terminus of N and in M had little effect on assembly. In contrast, with the flexible BMB linker, NM molecules crosslinked at the Head or Tail formed fibres very efficiently (grey bars in Fig. 2c). BMB crosslinks severely impeded fibre formation only in the Central Core.
The ∼2-Å bond length of a disulphide bond is closer than the inter-strand distances of ∼4.7 Å that characterize NM fibres20,28. The distributions of residues for which disulphide bonds inhibit amyloid formation support our earlier conclusion that a contiguous linear segment of amino acids, including residues 21–121, constitute a cooperatively folded unit. Improper intersubunit alignments of any two residues in this region prevent folding of the rest of the domain. Residues in the extreme N terminus of N and in M are outside this domain and have little influence on its capacity to fold.
With the longer linker, apposition of two NM proteins in the Head or of two NM proteins in the Tail permits fibre formation, but apposition of Central Core residues prevents it. Thus, the separation of Central Core regions is not only a general characteristic of NM fibres but is essential for fibre formation.
We next asked whether the structural information and the tools we had assembled could be used to address two of the most enigmatic questions in prion biology. How is prion assembly nucleated? And what is the structural basis of distinct prion strains?
Early events during nucleation and assembly
First, we monitored kinetic changes in the fluorescence of proteins labelled with acrylodan. All tested proteins that were labelled in the cooperatively folded amyloid region (21–106) showed a very rapid increase in fluorescence, characteristic of a first-order reaction with no lag phase (Fig. 4a). This fluorescence increase preceded conversion to amyloid: when amyloid formation was monitored by the acquisition of an SDS-insoluble state, each acrylodan-labelled protein had the same lag and assembly phase as wild-type protein (Supplementary Fig. S4). In contrast, molecules labelled at residues 158 or 167 changed fluorescence simultaneously with amyloid formation (Fig. 4a and data not shown). Proteins labelled at residue 184, 203 or 225 showed no change in fluorescence (Fig. 4a and data not shown). We conclude that: (1) residues that form the cooperatively folded amyloid core rapidly enter a collapsed but non-amyloid state, (2) M residues proximal to N become structured only when N residues convert to amyloid, and (3) the distal region of M remains largely unstructured and exposed to solvent after amyloid assembly.
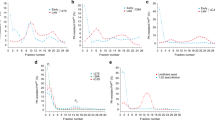
a, Formation of a collapsed intermediate. An increase in acrylodan fluorescence at positions N21 (black squares), Q38 (blue squares), G43 (red triangles), Y77 (green triangles), G96 (green hexagons) and Y106 (blue diamonds) occurred very rapidly after proteins were diluted into buffer, before the formation of amyloid (see also Supplementary Fig. S4). Amyloid formation was assessed by SDS insolubility because acrylodan and ThT fluorescence could not be analysed in the same reaction. Acrylodan fluorescence for mutants T158 (black triangles) and K184 (red circles) in the M domain, and ThT binding profiles of wild-type NM (blue circles) are plotted for comparison. b, Susceptibility of different cysteine residues to spontaneous crosslinking during assembly under non-reducing conditions. Crosslinks were assessed by non-reducing SDS–PAGE. c, Addition of a single charge in the Head region severely impedes fibre assembly. Assembly of negatively charged iodoacetate-labelled (blue) and uncharged iodoacetamide-labelled (red) proteins was monitored by ThT fluorescence. d, Assembly kinetics of cysteine variants crosslinked with BMB at position N21 (black triangles), G25 (black circles), Q38 (black squares), wild-type NM (blue triangles), G96 (blue circles), Y106 (blue squares), G43 (red squares) and Y73 (red circles), monitored by ThT fluorescence.
To determine which segments of the cooperatively folded amyloid region are the first to undergo conformational commitment, we took advantage of the fact that disulphide bonds anywhere in this region prohibit assembly (Fig. 2c). We reasoned that segments of NM that are the first to assume productive spatial relationships would also be the first to be protected from the spontaneous formation of disulphide bonds. Representative cysteine mutants were allowed to assemble in buffer without DTT (to facilitate the formation of disulphide bonds) and later analysed for the formation of crosslinks (dimers) on SDS gels without DTT. Cysteines in the Head region (21, 25 and 31) formed fewer disulphide bonds than cysteines at other positions (Fig. 4b). Thus, during conformational conversion, strand spacings compatible with a productive fold (and incompatible with disulphide formation) are achieved in the Head region more rapidly than in other regions.
Next, we investigated the effects of adding a single charge at various positions in the amyloid region. In some β-structures, such as the β-helix, alternating residues point towards or away from solvent, and the structures formed are very stable32. Although the introduction of a single charge would be unlikely to perturb such structures once they form, charge repulsions in early nucleating segments of the molten, collapsed intermediate would reduce the frequency with which these segments come into proximity, thereby slowing nucleation. Labelling individual NM cysteine residues with uncharged iodoacetamide had little effect on the quantity of protein converting to amyloid (Fig. 4c) or the kinetics of assembly (Supplementary Fig. S5). In contrast, iodoacetate labelling, which introduces a negatively charged moiety similar in size to acetamide, strongly inhibited assembly in the Head region (Fig. 4c and Supplementary Fig. S5).
Finally, if Head-to-Head interactions are not only characteristic of early productive amyloid conformations, but actually cause a commitment to it, bringing Head regions in proximity with each other should promote nucleation. We compared assembly kinetics for several NM variants that had been crosslinked with BMB under denaturing conditions and then transferred to assembly buffer. Crosslinks in the Central Core (residues 43 and 73) blocked assembly entirely, confirming that these regions must be separated from each other to form amyloid (Fig. 4d). Crosslinks in the Tail (residues 96 and 106) had little effect. Crosslinks in the Head (residues 21, 25 and 38) virtually eliminated the lag phase. Thus, the juxtaposition of residues in the Head region is an early event in amyloid formation and is, indeed, sufficient to nucleate it.
Structural distinctions between prion strains
To address the second critical question in prion biology—the basis of prion ‘strains’ or variants—within the structural framework we generated for NM fibres, we assembled the protein under conditions previously known to produce fibres enriched in different different strains32 (25 °C versus 4 °C). We confirmed that fibres produced at 4 °C assembled much more rapidly26 (data not shown), were less resistant to GdmCl denaturation (Fig. 5a), and produced mostly strong prion strains when used to transform cells from the [psi-] non-prion state to the [PSI+] prion state32 (Fig. 5c, left and Supplementary Fig. S7).
a, Amyloid core length and stability differ in fibres assembled at 4°C and 25°C. Acrylodan-labelled fibres assembled at 4 °C (blue) and 25 °C (red) were denatured as in Fig. 1c. D1/2 values were obtained from full GdmCl denaturation profiles (see Fig. 1c). b, Intersubunit interfaces change in fibres assembled at different temperatures. Excimer fluorescence of pyrene-labelled mutants assembled in rotated, unseeded reactions at 25°C (red), 4°C (blue) or in seeded, unrotated reactions with 4% seed formed at 4 °C (open circles). Pyrene fluorescence is similar in seeded and unseeded reactions. c, BMB crosslinks at different positions bias assembly towards different prion strains. NM proteins crosslinked in denaturant were diluted into buffer, assembled into fibres, and used to transform [psi-] cells to the [PSI+] state. Left columns show controls: un-crosslinked wild-type fibres assembled at 25 °C or at 4°C are biased towards the production of weak or strong strains, respectively27. Soluble NM, buffer alone, and proteins crosslinked in the Central Core did not induce [PSI+] above background and were not scored. Values represent means ± s.d. (n = 5 experiments). d, Strain biases created by crosslinking overcome the biases created by assembly at different temperatures. Whether they were assembled at 4 °C or at 25 °C, fibres cross-linked in the Head region (G25 or G31) produced mostly strong [PSI+] strains, and fibres crosslinked in the Tail region (G96 and Y106) produced mostly weak [PSI+] strains. Values represent means ± s.d. (n = 5 experiments).
Do fibres enriched in different prion strains have distinct cooperatively folded amyloid domains? Denaturation profiles of fibres, independently assembled from 16 acrylodan-labelled proteins at either 4 °C or 25 °C, were determined as in Fig. 1c. In fibres at 4 °C, residues 31–86 had strong blue shifts in fluorescence upon assembly and showed a single cooperative unfolding transition at D1/2 ∼ 1.5 M GdmCl (Fig. 5a and data not shown). Flanking residues 21, 25, 96, 112 and 121 had smaller blue shifts in fluorescence upon assembly, and heterogeneous denaturation profiles after assembly, as had residues flanking the amyloid domain at 25 °C (2 and 7, 137 and 158; Figs 1c and 5a, and data not shown). Thus, the cooperatively folded amyloid core has a similar character at both temperatures: it is formed by a contiguous stretch of amino acids and is flanked by residues that are structurally heterogeneous. However, the length of the region incorporated into the cooperative amyloid fold is much shorter in fibres at 4 °C than in fibres at 25 °C, and is consequently more easily denatured. Shorter, less stable amyloid cores probably produce stronger, more stably inherited prion strains because the fibres are more easily fragmented and transmitted to daughter cells, promoting the cycle of inheritance.
Finally, we asked if intermolecular contacts differ in fibres assembled at 4 °C and 25 °C. The excimer fluorescence of residues in the Head region and in flanking residues (2, 7 and 16) changed modestly but in an extremely reproducible manner (seeded and unseeded reactions; Fig. 5b). Excimer fluorescence in the Tail showed a larger shift. Thus, as the number of residues constituting the cooperatively folded amyloid domain changes, the intersubunit interfaces change as well.
The structural basis of prion strains
In fibres enriched for distinct prion strains, we confirmed27 differences in the rate at which assembly occurred, and discovered differences in both the length of the amyloid core and the intersubunit contacts. But are these features determinative? If so, proteins crosslinked in different places should strongly bias assembly reactions towards different prion strains.
Proteins crosslinked with BMB (as for Fig. 2c) were assembled at 25 °C or 4 °C and used to transform cells from the [psi-] to the [PSI+] state26,27. Proteins crosslinked in the Central Core did not induce [PSI+] above background levels (Fig. 5c), consistent with their failure to undergo amyloid assembly (Fig. 2c). Crosslinks in the Head, which caused very rapid assembly, biased fibres towards the production of strong strains. Conversely, crosslinks in the Tail, which should cause a longer segment to be incorporated into the cooperative amyloid fold, biased fibres towards weak strains (Fig. 5c). Notably, the strain biases produced by different crosslink positions overcame those of different assembly temperatures. Whether the fibres used for transformation had been assembled at 4 °C or at 25 °C, proteins crosslinked in the Head produced primarily strong strains, and proteins crosslinked in the Tail produced primarily weak strains (Fig. 5d). We conclude that strain distinctions are due to differences in the length of the amyloid core of individual NM molecules, as well as the nature of NM–NM interfaces.
Discussion
Our results provide a framework for the structure of NM fibres, define rate-limiting events that govern their nucleation and establish a physical basis for prion strains. The data provide many insights regarding prion initiation and propagation, and are likely to shed light on amyloid biology in other systems.
The first step in assembly is the formation of a collapsed intermediate, in which residues in the N region are sequestered from solvent but have not yet locked into their final structures (Fig. 4a and Supplementary Fig. S4). The search for structure in globular proteins also begins with a collapsed intermediate, but these are dominated by hydrophobic interactions. The collapse of N must be governed by polar interactions and/or backbone interactions, because polar residues outweigh hydrophobic residues by 16 to 1. Hence, proteins governed by very different interactions, and achieving very different final structures, both do so through a molten, collapsed intermediate.
Within this collapsed state, it is the Head region that commits most rapidly to amyloid-compatible spatial relationships (Fig. 4b). Moreover, simply bringing the Head regions of two NM molecules into close proximity triggers nucleation (Fig. 4d). We suggest that Head-to-Head contacts are a principal route for NM nucleation, funnelling the protein away from other folding pathways that might still be productive but are much slower. The lag phase of assembly would then represent the time required to search out these productive contacts in the collapsed molten state. The importance of Head-to-Head contacts in nucleation clarifies why the ‘species barrier’, which prevents cross-species seeding between S. cerevisiae and Candida albicans NM proteins, maps to this region2,33,34.
After nucleation, fibres grow through Head-to-Head and Tail-to-Tail interactions (Figs 2a, b and 3). This explains why they grow bi-directionally21. Moreover, the fact that both ends of NM are involved in prion propagation explains why rare cross-seeding events (between NM proteins from diverse species and between entirely different prions in the same species) rapidly convert from heterotypic to homotypic interactions35,36,37. Once the heterologous protein has joined the fibre, it will immediately present a homotypic interface for either Head-to-Head or Tail-to-Tail interactions.
Our work also sheds light on the role played by the 5.5 degenerate oligopeptide repeats (consensus sequence P/QQGGYQQ/SYN). Increasing the number of repeats greatly increases spontaneous prion formation in vivo and fibre nucleation in vitro38. Reducing them eliminates the protein's ability to propagate in vivo as a prion on its own. However, even a single repeat allows efficient incorporation into pre-existing wild-type prions38,39,40. We find that most repeats are in the Central Core, sequestered from intermolecular contacts (Figs 2 and 4d), and that the length of the Central Core determines fibre stability (Fig. 5a). However, intermolecular Tail-to-Tail contacts include the final 1.5 to 2.5 repeats. Repeat-to-repeat contacts are therefore accommodated in both intramolecular and intermolecular amyloid interactions. With a single repeat, the stability of NM amyloid might fall below the threshold required to maintain the prion state on its own (perhaps through increased susceptibility to Hsp104 disaggregation23). However, with this single repeat, the molecule could still join pre-existing wild-type prions through Head-to-Head and repeat-to-repeat interactions. Noting that the repeat sequence is able to form an amyloid on its own41, we further propose that the closely spaced repeat sequences of individual NM molecules readily collapse into transient pre-amyloid structures within the molten intermediates where nucleation occurs. The more repeats, the more likely this transient structure is to form, reducing the number of contacts the Head region must sample to initiate nucleation, and thereby increasing the spontaneous rate of prion formation.
What structural elements are responsible for enciphering prion strains? Previous data are perplexing. On one hand, amyloids assembled in vitro from a fragment containing only the first 65 residues of N can induce a broad spectrum of distinct heritable prion strains when used to transform [psi-] cells to [PSI+]26. On the other hand, during in vivo propagation, residues 1–137 are required to maintain several [PSI+] strains42. The extremely reproducible pyrene excimer signals in our experiments (Figs 2a and 5b) provide a key. Different intermolecular contacts in both the Head and Tail create distinct strains. Residues 1–65 contain the Head region and two repeats. This fragment should be able to form the full spectrum of Head region interactions that distinguish different strains, and be able to propagate these to wild-type protein. However, differences in the length of the Central Core and the position of Tail-to-Tail contacts are the major distinctions between prion strains. Hence, residues 65–137 are required for the maintenance of distinct strains in vivo.
Does our data have relevance to other prions and amyloids? Although it is not yet clear that molten oligomeric species are obligate intermediates in the formation of other amyloids (as it is for NM), they are present in the initial stages of many amyloid assemblies43,44. Evidence also attests to the importance of dimeric interactions in the assembly of diverse amyloids45,46,47. Recent molecular dynamics simulations with Aβ peptide suggest that assembly begins with a molten oligomeric species that allows the rapid sampling of multiple intermolecular contacts, with two individual molecules precipitously finding the right contact and nucleating assembly44.
Several other similarities with the mammalian prion determinant, PrP, are apparent. These include (1) electron crystallography data supporting a β-helix-like structure in the prion state48,49; (2) the importance of PHGGGWGQ oligopeptide repeats (an increase in repeat number causes spontaneous prion formation, and eliminating repeats inhibits propagation); and (3) the ability of PrP to form a variety of phenotypically distinct strains with different structures (that is, different sensitivities to GdmCl denaturation, protease cleavage sites and glycosylation states)50.
For simplicity, we distinguished only strong and weak strains of NM but, in fact, each category includes a multiplicity of strains (ref. 27 and data not shown). We suggest that promiscuity in intermolecular Head and Tail contacts, together with the low sequence complexity of the N domain (72% Q, N, G and Y) allow differences in intermolecular contacts to ratchet the amyloid core into different registers, creating an ensemble of structures with closely related but distinct β-strand contacts. If the length of the amyloid core and the residues involved in contact interfaces differ for PrP strains as they do for NM, it would readily explain variations in transmission within and between species. One fold might display an interface that is polymorphic between species, another might display an invariant interface. One fold might restrict the incorporation of a glycosylated residue because the modified side chain would point inward; another might permit it because the residue points outward. Similarly, transient intramolecular β-structures formed by expanded oligopeptide repeats would reduce the number of intermolecular contacts that must be sampled to nucleate prion formation, even though the prion-enhancing repeats of PrP lie outside the final amyloid core4,5. There will undoubtedly be many variations in the structure and dynamics of amyloid assemblies, but several of the principles we have uncovered, as well as the methods we used, will probably be applicable to other systems.
Note added in proof: A study focusing on three residues (amino acids 36, 76 and 108) has recently reported that paragmagnetic spin resonance spectroscopy provides evidence for strain-specific differences that affect the species barrier between S. cerevisiae and C. albicans51.
Methods
Mutagenesis, protein purification and yeast strains
The site-specific substitution of individual amino-acid codons for cysteine codons, the production of integrating genomic constructs, the strategy for replacing wild-type SUP35 with the cysteine mutants, and overexpression and purification of cysteine mutants is described in the Supplementary Information.
Cysteine labelling
Post-assembly cysteine labelling: 5 µM NM fibres were incubated with 15 µM of pyrene maleimide or 15 µM Lucifer yellow for 3 h. Fibres were washed three times with 20% methanol containing 5 mM DTT, and redissolved in 6 M GdmCl. Labelling efficiencies were calculated according to the Molecular probes website (http://www.probes.com).
Pre-assembly cysteine labelling: proteins were incubated in 6 M GdmCl with acrylodan or pyrene maleimide as recommended by Molecular Probes (see Supplementary Information). Labelling efficiencies for acrylodan (> 90%) and pyrene (70–80%) were also determined according to the manufacturer's protocol.
Iodoacetate and iodoacetamide labelling in 6 M GdmCl used 25 µM protein and a fivefold excess of iodoacetate or iodoacetamide for 2 h at 25 °C. After using a PD10 desalting column (Pharmacia) to remove free label, efficiencies were determined from the number of free thiol groups still accessible to 5,5′-dithiobis-2-nitrobenzoic acid.
BMB (1,4-bis-maleimidobutane) was used at a protein-to-reagent ratio of 1:2. O-DTT (oxidized dithiothreitol) was used at a ratio of 5:1. Reactions in 6 M GdmCl were terminated after 3 h with 5 mM DTT. Crosslinking efficiencies (all 70–85%) were assessed by SDS–PAGE.
Protein transformation
Yeast cells containing an ADE1 mutation suppressible by [PSI+] were co-transformed with NM proteins and a URA3 plasmid as described27. Ura + transformants were picked before colour development (three days instead of seven), to avoid bias. Approximately 200 transformants for each mutant (obtained from five independent transformations in each case), were patched onto uracil-deficient medium and replica plated to both adenine-deficient medium and 25% rich medium (YPD; yeast extract, peptone and dextrose) supplemented with 20 µg ml-1 adenine. Transformation efficiencies were calculated (Supplementary Fig. S6) and strong and weak strains were scored27 by colour on YPD (Supplementary Fig. S1), and rates of growth were scored on adenine-deficient medium.
References
Tuite, M. F. & Cox, B. S. Propagation of yeast prions. Nature Rev. Mol. Cell Biol. 4, 878–890 (2003)
Chien, P., Weissman, J. S. & DePace, A. H. Emerging principles of conformation-based prion inheritance. Annu. Rev. Biochem. 73, 617–656 (2004)
Shorter, J. & Lindquist, S. Prions as adaptive conduits of memory and inheritance. Nature Rev. Genet. doi:10.1038/nrg1616 (in the press)
Caughey, B. Transmissible spongiform encephalopathies, amyloidoses and yeast prions: common threads? Nature Med. 6, 751–754 (2000)
Prusiner, S. B. Prions. Proc. Natl Acad. Sci. USA 95, 13363–13383 (1998)
Uptain, S. M. & Lindquist, S. Prions as protein-based genetic elements. Annu. Rev. Microbiol. 56, 703–741 (2002)
Si, K. et al. A neuronal isoform of CPEB regulates local protein synthesis and stabilizes synapse-specific long-term facilitation in Aplysia. Cell 115, 893–904 (2003)
Si, K., Lindquist, S. & Kandel, E. R. A neuronal isoform of the Aplysia CPEB has prion-like properties. Cell 115, 879–891 (2003)
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890 (2003)
Jensen, M. A., True, H. L., Chernoff, Y. O. & Lindquist, S. Molecular population genetics and evolution of a prion-like protein in Saccharomyces cerevisiae. Genetics 159, 527–535 (2001)
Eaglestone, S. S., Cox, B. S. & Tuite, M. F. Translation termination efficiency can be regulated in Saccharomyces cerevisiae by environmental stress through a prion-mediated mechanism. EMBO J. 18, 1974–1981 (1999)
True, H. L. & Lindquist, S. L. A yeast prion provides a mechanism for genetic variation and phenotypic diversity. Nature 407, 477–483 (2000)
True, H. L., Berlin, I. & Lindquist, S. L. Epigenetic regulation of translation reveals hidden genetic variation to produce complex traits. Nature 431, 184–187 (2004)
Kushnirov, V. V. et al. Divergence and conservation of SUP2 (SUP35) gene of yeast Pichia pinus and Saccharomyces cerevisiae. Yeast 6, 461–472 (1990)
Ter-Avanesyan, M. D. et al. Deletion analysis of the SUP35 gene of the yeast Saccharomyces cerevisiae reveals two non-overlapping functional regions in the encoded protein. Mol. Microbiol. 7, 683–692 (1993)
Chernoff, Y. O., Derkach, I. L. & Inge-Vechtomov, S. G. Multicopy SUP35 gene induces de-novo appearance of psi-like factors in the yeast Saccharomyces cerevisiae. Curr. Genet. 24, 268–270 (1993)
Glover, J. R. et al. Self-seeded fibers formed by Sup35, the protein determinant of [PSI + ], a heritable prion-like factor of S. cerevisiae. Cell 89, 811–819 (1997)
Liu, J. J., Sondheimer, N. & Lindquist, S. L. Changes in the middle region of Sup35 profoundly alter the nature of epigenetic inheritance for the yeast prion [PSI + ]. Proc. Natl Acad. Sci. USA 99(suppl. 4), 16446–16453 (2002)
Li, L. & Lindquist, S. Creating a protein-based element of inheritance. Science 287, 661–664 (2000)
Serio, T. R. et al. Nucleated conformational conversion and the replication of conformational information by a prion determinant. Science 289, 1317–1321 (2000)
Scheibel, T., Kowal, A. S., Bloom, J. D. & Lindquist, S. L. Bidirectional amyloid fiber growth for a yeast prion determinant. Curr. Biol. 11, 366–369 (2001)
Scheibel, T., Bloom, J. & Lindquist, S. L. The elongation of yeast prion fibers involves separable steps of association and conversion. Proc. Natl Acad. Sci. USA 101, 2287–2292 (2004)
Shorter, J. & Lindquist, S. Hsp104 catalyzes formation and elimination of self-replicating Sup35 prion conformers. Science 304, 1793–1797 (2004)
Collins, S. R., Douglass, A., Vale, R. D. & Weissman, J. S. Mechanism of prion propagation: amyloid growth occurs by monomer addition. PLoS Biol. 2, e321 (2004)
DePace, A. H. & Weissman, J. S. Origins and kinetic consequences of diversity in Sup35 yeast prion fibers. Nature Struct. Biol. 9, 389–396 (2002)
King, C. Y. & Diaz-Avalos, R. Protein-only transmission of three yeast prion strains. Nature 428, 319–323 (2004)
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J. S. Conformational variations in an infectious protein determine prion strain differences. Nature 428, 323–328 (2004)
Kishimoto, A. et al. β-Helix is a likely core structure of yeast prion Sup35 amyloid fibers. Biochem. Biophys. Res. Commun. 315, 739–745 (2004)
DePace, A. H., Santoso, A., Hillner, P. & Weissman, J. S. A critical role for amino-terminal glutamine/asparagine repeats in the formation and propagation of a yeast prion. Cell 93, 1241–1252 (1998)
King, C. Y. Supporting the structural basis of prion strains: induction and identification of [PSI] variants. J. Mol. Biol. 307, 1247–1260 (2001)
Kajava, A. V., Baxa, U., Wickner, R. B. & Steven, A. C. A model for Ure2p prion filaments and other amyloids: the parallel superpleated β-structure. Proc. Natl Acad. Sci. USA 101, 7885–7890 (2004)
Kamen, D. E., Griko, Y. & Woody, R. W. The stability, structural organization, and denaturation of pectate lyase C, a parallel β-helix protein. Biochemistry 39, 15932–15943 (2000)
Santoso, A., Chien, P., Osherovich, L. Z. & Weissman, J. S. Molecular basis of a yeast prion species barrier. Cell 100, 277–288 (2000)
Chien, P., DePace, A. H., Collins, S. R. & Weissman, J. S. Generation of prion transmission barriers by mutational control of amyloid conformations. Nature 424, 948–951 (2003)
Bradley, M. E., Edskes, H. K., Hong, J. Y., Wickner, R. B. & Liebman, S. W. Interactions among prions and prion “strains” in yeast. Proc. Natl Acad. Sci. USA 99 (suppl. 4), 16392–16399 (2002)
Derkatch, I. L. et al. Effects of Q/N-rich, polyQ, and non-polyQ amyloids on the de novo formation of the [PSI + ] prion in yeast and aggregation of Sup35 in vitro. Proc. Natl Acad. Sci. USA 101, 12934–12939 (2004)
Nakayashiki, T., Ebihara, K., Bannai, H. & Nakamura, Y. Yeast [PSI + ] “prions” that are crosstransmissible and susceptible beyond a species barrier through a quasi-prion state. Mol. Cell 7, 1121–1130 (2001)
Liu, J. J. & Lindquist, S. Oligopeptide-repeat expansions modulate ’protein-only’ inheritance in yeast. Nature 400, 573–576 (1999)
Parham, S. N., Resende, C. G. & Tuite, M. F. Oligopeptide repeats in the yeast protein Sup35p stabilize intermolecular prion interactions. EMBO J. 20, 2111–2119 (2001)
Osherovich, L. Z., Cox, B. S., Tuite, M. F. & Weissman, J. S. Dissection and design of yeast prions. PLoS Biol. 2, E86 (2004)
Balbirnie, M., Grothe, R. & Eisenberg, D. S. An amyloid-forming peptide from the yeast prion Sup35 reveals a dehydrated β-sheet structure for amyloid. Proc. Natl Acad. Sci. USA 98, 2375–2380 (2001)
Bradley, M. E. & Liebman, S. W. The Sup35 domains required for maintenance of weak, strong or undifferentiated yeast [PSI + ] prions. Mol. Microbiol. 51, 1649–1659 (2004)
Eakin, C. M., Attenello, F. J., Morgan, C. J. & Miranker, A. D. Oligomeric assembly of native-like precursors precedes amyloid formation by β-2 microglobulin. Biochemistry 43, 7808–7815 (2004)
Hwang, W., Zhang, S., Kamm, R. D. & Karplus, M. Kinetic control of dimer structure formation in amyloid fibrillogenesis. Proc. Natl Acad. Sci. USA 101, 12916–12921 (2004)
Lee, S. & Eisenberg, D. Seeded conversion of recombinant prion protein to a disulfide-bonded oligomer by a reduction-oxidation process. Nature Struct. Biol. 10, 725–730 (2003)
Garzon-Rodriguez, W. et al. A conformation change in the carboxyl terminus of Alzheimer's Aβ(1–40) accompanies the transition from dimer to fibril as revealed by fluorescence quenching analysis. J. Biol. Chem. 275, 22645–22649 (2000)
Barghorn, S. & Mandelkow, E. Toward a unified scheme for the aggregation of tau into Alzheimer paired helical filaments. Biochemistry 41, 14885–14896 (2002)
Wille, H. et al. Structural studies of the scrapie prion protein by electron crystallography. Proc. Natl Acad. Sci. USA 99, 3563–3568 (2002)
Govaerts, C., Wille, H., Prusiner, S. B. & Cohen, F. E. Evidence for assembly of prions with left-handed β-helices into trimers. Proc. Natl Acad. Sci. USA 101, 8342–8347 (2004)
Safar, J. et al. Eight prion strains have PrPSc molecules with different conformations. Nature Med. 4, 1157–1165 (1998)
Tanaka, M., Chien, P., Yonekura, K. & Weissman, J. S. Mechanism of cross-species prion transmission: an infectious conformation compatible with two highly divergent yeast prion proteins. Cell 121, 49–62 (2005)
Acknowledgements
We thank J. Weissman for providing us with the fibre transformation protocols, members of the Lindquist, Langen and Berger laboratories for helpful discussions, and R. Latek and T. DiCesare for help with the figures. This research was supported by the DuPont-MIT Alliance and an NIH grant.Author Contributions Experimental work was performed by R.K., data analysis by R.K. and S.L.L., and writing primarily by S.L.L.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This contains Supplementary Notes, Legends to accompany Supplementary Figures S1-S6 and Supplementary References (PDF 142 kb)
Supplementary Figure S1
Single cysteine substitution mutations behave like wild-type NM (PDF 64 kb)
Supplementary Figure S2
Analysis of pyrene fluorescence (PDF 53 kb)
Supplementary Figure S3
Excimer values for the experiment in Fig. 2b employing fibres assembled with two labelled proteins, in all pairwise interactions (PDF 13 kb)
Supplementary Figure S4
Acrylodan-labelling of NM proteins does not affect amyloid formation (PDF 82 kb)
Supplementary Figure S5
Assembly kinetics (PDF 24 kb)
Supplementary Figure S6
The efficiency with which cells are converted from the [psi-] to the [PSI+] state after transformation with various preparations of NM (PDF 18 kb)
Supplementary Figure S7
Strong and weak strains produced by fiber-mediated protein transformation. (PDF 4484 kb)
Rights and permissions
About this article
Cite this article
Krishnan, R., Lindquist, S. Structural insights into a yeast prion illuminate nucleation and strain diversity. Nature 435, 765–772 (2005). https://doi.org/10.1038/nature03679
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03679
This article is cited by
-
Generic nature of the condensed states of proteins
Nature Cell Biology (2021)
-
Generalizing a mathematical model of prion aggregation allows strain coexistence and co-stability by including a novel misfolded species
Journal of Mathematical Biology (2019)
-
Heterologous prion-forming proteins interact to cross-seed aggregation in Saccharomyces cerevisiae
Scientific Reports (2017)
-
Melanosomal formation of PMEL core amyloid is driven by aromatic residues
Scientific Reports (2017)
-
Detergent-induced aggregation of an amyloidogenic intrinsically disordered protein
Journal of Chemical Sciences (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.