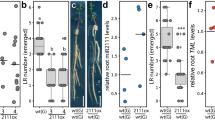
Abstract
The roots of most higher plants form arbuscular mycorrhiza, an ancient, phosphate-acquiring symbiosis with fungi, whereas only four related plant orders are able to engage in the evolutionary younger nitrogen-fixing root-nodule symbiosis with bacteria1. Plant symbioses with bacteria and fungi require a set of common signal transduction components that redirect root cell development2,3. Here we present two highly homologous genes from Lotus japonicus, CASTOR and POLLUX, that are indispensable for microbial admission into plant cells and act upstream of intracellular calcium spiking4, one of the earliest plant responses to symbiotic stimulation. Surprisingly, both twin proteins are localized in the plastids of root cells, indicating a previously unrecognized role of this ancient endosymbiont in controlling intracellular symbioses that evolved more recently.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Smith, S. E. & Read, D. J. Mycorrhizal Symbiosis (Academic, London, 1997)
Oldroyd, G. E. D. Dissecting symbiosis: developments in Nod factor signal transduction. Ann. Bot. 87, 709–718 (2001)
Kistner, C. & Parniske, M. Evolution of signal transduction in intracellular symbiosis. Trends Plant Sci. 7, 511–518 (2002)
Ehrhardt, D., Wais, R. & Long, S. Calcium spiking in plant root hairs responding to Rhizobium nodulation signals. Cell 85, 673–681 (1996)
Truchet, G. et al. Sulphated lipooligosaccharide signals from Rhizobium meliloti elicit root nodule organogenesis in alfalfa. Nature 351, 670–673 (1991)
Radutoiu, S. et al. Plant recognition of symbiotic bacteria requires two LysM receptor-like kinases. Nature 425, 585–592 (2003)
Cárdenas, L. et al. Ion changes in legume root hairs responding to Nod factors. Plant Physiol. 123, 443–452 (2000)
Senoo, K. et al. Isolation of two different phenotypes of mycorrhizal mutants in the model legume plant Lotus japonicus after EMS-treatment. Plant Cell Physiol. 41, 726–732 (2000)
Szczyglowski, K. et al. Nodule organogenesis and symbiotic mutants of the model legume Lotus japonicus. Mol. Plant Microbe Interact. 11, 684–697 (1998)
Bonfante, P. et al. The Lotus japonicus LjSym4 gene is required for the successful symbiotic infection of root epidermal cells. Mol. Plant Microbe Interact. 13, 1109–1120 (2000)
Novero, M. et al. Dual requirement of the LjSym4 gene for mycorrhizal development in epidermal and cortical cells of Lotus japonicus roots. New Phytol. 154, 741–749 (2002)
Harris, J. M., Wais, R. & Long, S. R. Rhizobium-induced calcium spiking in Lotus japonicus. Mol. Plant Microbe Interact. 16, 335–341 (2003)
Hayashi, M. et al. Construction of a genetic linkage map of the model legume Lotus japonicus using an intraspecific F2 population. DNA Res. 8, 301–310 (2001)
Nakamura, Y. et al. Structural analysis of a Lotus japonicus genome. II. Sequence features and mapping of sixty-five TAC clones which cover the 6.5-mb regions of the genome. DNA Res. 9, 63–70 (2002)
Kawasaki, S. & Murakami, Y. Genome analysis of Lotus japonicus. J. Plant Res. 113, 497–506 (2000)
Kawaguchi, M. et al. Providing the basis for genomics in Lotus japonicus: the accessions Miyakojima and Gifu are appropriate crossing partners for genetic analyses. Mol. Gen. Genomics 266, 157–166 (2001)
Stracke, S. et al. A plant receptor-like kinase required for both fungal and bacterial symbiosis. Nature 417, 959–962 (2002)
Ane, J. M. et al. Medicago truncatula DMI1 required for bacterial and fungal symbioses in legumes. Science 303, 1364–1367 (2004)
Köhler, R. H. et al. Exchange of protein molecules through connections between higher plant plastids. Science 276, 2039–2042 (1997)
Shi, J., Blundell, T. L. & Mizuguchi, K. FUGUE: sequence-structure homology recognition using environment-specific substitution tables and structure-dependent gap penalties. J. Mol. Biol. 310, 243–257 (2001)
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002)
Jiang, Y. et al. Structure of the RCK domain from the E. coli K+ channel and demonstration of its presence in the human BK channel. Neuron 29, 593–601 (2001)
Kwok, E. Y. & Hanson, M. R. Plastids and stromules interact with the nucleus and cell membrane in vascular plants. Plant Cell Rep. 23, 188–195 (2004)
Kawaguchi, M. et al. Root, root hair, and symbiotic mutants of the model legume Lotus japonicus. Mol. Plant Microbe Interact. 15, 17–26 (2002)
Perry, J. A. et al. A TILLING reverse genetics tool and a web-accessible collection of mutants of the legume Lotus japonicus. Plant Physiol. 131, 866–871 (2003)
Schauser, L. et al. Symbiotic mutants deficient in nodule establishment identified after T-DNA transformation of Lotus japonicus. Mol. Gen. Genet. 259, 414–423 (1998)
Niwa, S. et al. Responses of a model legume Lotus japonicus to lipochitin oligosaccharide nodulation factors purified from Mesorhizobium loti JRL501. Mol. Plant Microbe Interact. 14, 848–856 (2001)
Broughton, W. J. & Dilworth, M. Y. Control of leghemoglobin synthesis in snake beans. Biochem. J. 125, 1075–1080 (1971)
Firmin, J. L. et al. Resistance to nodulation of cv. Afghanistan peas is overcome by nodX, which mediates an O-acetylation of the Rhizobium leguminosarum lipo-oligosaccharide nodulation factor. Mol. Microbiol. 10, 351–360 (1993)
Isono, K. et al. Leaf-specifically expressed genes for polypeptides destined for chloroplasts with domains of σ70 factors of bacterial RNA polymerases in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 94, 14948–14953 (1997)
Acknowledgements
We thank K. Szczyglowski, J. Webb and J. Stougaard for providing mutant seeds; M. Hayashi for help with mapping; T. Kojima and R. Ohtomo for mycorrhiza analysis; Y. Niwa for providing pUC18-CaMV35S-sGFP (S65T)-nos vector; G. Oldroyd and J. Sun for help with Ca-spiking assays; J. Krüger and B. B. H. Wulff for critical reading of the manuscript; J. Soll for providing the pea root transformation protocol before publication; and M. Durrant for help with modelling the CASTOR pore structure. Part of this work was supported by the fund of Promotion of Basic Research Activities for Innovative Biosciences (BRAIN), and Core Research for Evolutional Science and Technology (CREST), Japan Science and Technology Agency. Research at the Sainsbury Laboratory is funded by the Gatsby Charitable Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Positional cloning of CASTOR and POLLUX genes, and legend. (PPT 86 kb)
Supplementary Figure 1 legend
Additional copy of the legend for Supplementary Figure S1. (DOC 21 kb)
Supplementary Table 1
Castor and pollux mutant alleles. (DOC 41 kb)
Rights and permissions
About this article
Cite this article
Imaizumi-Anraku, H., Takeda, N., Charpentier, M. et al. Plastid proteins crucial for symbiotic fungal and bacterial entry into plant roots. Nature 433, 527–531 (2005). https://doi.org/10.1038/nature03237
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03237
This article is cited by
-
Wild species rice OsCERK1DY-mediated arbuscular mycorrhiza symbiosis boosts yield and nutrient use efficiency in rice breeding
Molecular Breeding (2024)
-
Nutrient regulation of lipochitooligosaccharide recognition in plants via NSP1 and NSP2
Nature Communications (2022)
-
Endophytic fungi: understanding complex cross-talks
Symbiosis (2021)
-
Calcium spikes, waves and oscillations in plant development and biotic interactions
Nature Plants (2020)
-
An ancestral signalling pathway is conserved in intracellular symbioses-forming plant lineages
Nature Plants (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.