Abstract
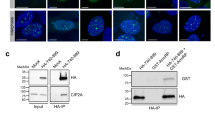
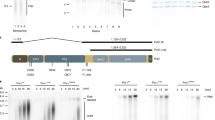
The checkpoint regulatory mechanism has an important role in maintaining the integrity of the genome1,2,3,4,5. This is particularly important in S phase of the cell cycle, when genomic DNA is most susceptible to various environmental hazards3,6,7. When chemical agents damage DNA, activation of checkpoint signalling pathways results in a temporary cessation of DNA replication. A replication-pausing complex is believed to be created at the arrested forks to activate further checkpoint cascades, leading to repair of the damaged DNA. Thus, checkpoint factors are thought to act not only to arrest replication but also to maintain a stable replication complex at replication forks6,7,8,9. However, the molecular mechanism coupling checkpoint regulation and replication arrest is unknown. Here we demonstrate that the checkpoint regulatory proteins Tof1 and Mrc1 interact directly with the DNA replication machinery in Saccharomyces cerevisiae. When hydroxyurea blocks chromosomal replication, this assembly forms a stable pausing structure that serves to anchor subsequent DNA repair events.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Zhou, B. S. & Elledge, S. J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439 (2000)
Melo, J. & Toczyski, D. A unified view of the DNA-damage checkpoint. Curr. Opin. Cell Biol. 14, 237–245 (2002)
Shirahige, K. et al. Regulation of DNA-replication origins during cell-cycle progression. Nature 395, 618–621 (1998)
Kondo, T., Wakayama, T., Naiki, T., Matsumoto, K. & Sugimoto, K. Recruitment of Mec1 and Ddc1 checkpoint proteins to double-strand breaks through distinct mechanisms. Science 294, 867–870 (2001)
Melo, J., Cohen, J. & Toczyski, D. Two checkpoint complexes are independently recruited to sites of DNA damage in vivo. Genes Dev. 15, 2809–2821 (2001)
Lopes, M. et al. The DNA replication checkpoint response stabilizes stalled replication forks. Nature 412, 557–561 (2001)
Tercero, J. A. & Diffley, J. F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557 (2001)
Tercero, J. A., Longhese, M. P. & Diffley, J. F. A central role for DNA replication forks in checkpoint activation and response. Mol. Cell 11, 1323–1336 (2003)
Sogo, J. M., Lopes, M. & Foiani, M. Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science 297, 599–602 (2002)
Shirahige, K., Iwasaki, T., Rashid, M. B., Ogasawara, N. & Yoshikawa, H. Location and characterization of autonomously replicating sequences from chromosome VI of Saccharomyces cerevisiae. Mol. Cell. Biol. 13, 5043–5056 (1993)
Yamashita, M. et al. The efficiency and timing of initiation of replication of multiple replicons of Saccharomyces cerevisiae chromosome VI. Genes Cells 2, 655–665 (1997)
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000)
Iyer, V. R. et al. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 409, 533–538 (2001)
Lengronne, A., Pasero, P., Bensimon, A. & Schwob, E. Monitoring S phase progression globally and locally using BrdU incorporation in TK(+ ) yeast strains. Nucleic Acids Res. 29, 1433–1442 (2001)
Kurdistani, S. K., Robyr, D., Tavazoie, S. & Grunstein, M. Genome-wide binding map of the histone deacetylase Rpd3 in yeast. Nature Genet. 31, 248–254 (2002)
Alcasabas, A. A. et al. Mrc1 transduces signals of DNA replication stress to activate Rad53. Nature Cell Biol. 3, 958–965 (2001)
Zhao, X., Muller, E. G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998)
Labib, K., Tercero, J. A. & Diffley, J. F. Uninterrupted MCM2–7 function required for DNA replication fork progression. Science 288, 1643–1647 (2000)
Foss, E. J. Tof1p regulates DNA damage responses during S phase in Saccharomyces cerevisiae. Genetics 157, 567–577 (2001)
Tanaka, K. & Russell, P. Mrc1 channels the DNA replication arrest signal to checkpoint kinase Cds1. Nature Cell Biol. 3, 966–972 (2001)
De Antoni, A. & Gallwitz, D. A novel multi-purpose cassette for repeated integrative epitope tagging of genes in Saccharomyces cerevisiae. Gene 246, 179–185 (2000)
Paulovich, A. G. & Hartwell, L. H. A checkpoint regulates the rate of progression through S phase in S. cerevisiae in response to DNA damage. Cell 82, 841–847 (1995)
Winzeler, E. A. et al. Direct allelic variation scanning of the yeast genome. Science 281, 1194–1197 (1998)
Cimbora, D. M. et al. Long-distance control of origin choice and replication timing in the human β-globin locus are independent of the locus control region. Mol. Cell. Biol. 20, 5581–5591 (2000)
Schubeler, D. et al. Genome-wide DNA replication profile for Drosophila melanogaster: a link between transcription and replication timing. Nature Genet. 32, 438–442 (2002)
Zou, L. & Stillman, B. Assembly of a complex containing Cdc45p, replication protein A, and Mcm2p at replication origins controlled by S-phase cyclin-dependent kinases and Cdc7p-Dbf4p kinase. Mol. Cell. Biol. 20, 3086–3096 (2000)
Acknowledgements
We thank A. De Antoni, E. Schwob, S. Bell and J. Hegemann for the plasmids; K. Tamai for the special preparation of BrdU antibody; H. Araki, K. Okumura, E. Winzeler, M Foiani, T. Yada and K. Umezu for their comments; E. Schwob, F. Uhlmann, R. Cha and H. Yoshikawa for critical reading of the manuscript; T. Itoh, Y. Nakao, A. Nakada and C. Kawagoe for technical assistance; and Y. Sakaki and all the members of the Genome Structure and Function team for their support. This work was supported partly by grants-in-aid on priority areas from the Ministry of Education, Culture, Sports, Science and Technology, Japan, to K.S. Y. Katou is a Junior Research Associate of the RIKEN GSC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Additional information
Data presented in this paper can be obtained from GEO (http://www.ncbi.nlm.nih.gov/geo) with accession number GSE486.
Rights and permissions
About this article
Cite this article
Katou, Y., Kanoh, Y., Bando, M. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083 (2003). https://doi.org/10.1038/nature01900
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01900
This article is cited by
-
Replisome-cohesin interactions provided by the Tof1-Csm3 and Mrc1 cohesion establishment factors
Chromosoma (2023)
-
SWI/SNF and the histone chaperone Rtt106 drive expression of the Pleiotropic Drug Resistance network genes
Nature Communications (2022)
-
Histones on fire: the effect of Dun1 and Mrc1 on origin firing and replication of hyper-acetylated genomes
Current Genetics (2021)
-
Characterisation of unessential genes required for survival under conditions of DNA stress
Journal of Genetic Engineering and Biotechnology (2020)
-
Ethanol exposure increases mutation rate through error-prone polymerases
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.