Abstract
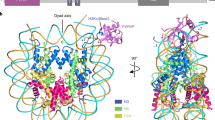
Specific modifications to histones are essential epigenetic markers1—heritable changes in gene expression that do not affect the DNA sequence. Methylation of lysine 9 in histone H3 is recognized by heterochromatin protein 1 (HP1), which directs the binding of other proteins to control chromatin structure and gene expression2,3,4. Here we show that HP1 uses an induced-fit mechanism for recognition of this modification, as revealed by the structure of its chromodomain bound to a histone H3 peptide dimethylated at Nζ of lysine 9. The binding pocket for the N-methyl groups is provided by three aromatic side chains, Tyr 21, Trp 42 and Phe 45, which reside in two regions that become ordered on binding of the peptide. The side chain of Lys 9 is almost fully extended and surrounded by residues that are conserved in many other chromodomains. The QTAR peptide sequence preceding Lys 9 makes most of the additional interactions with the chromodomain, with HP1 residues Val 23, Leu 40, Trp 42, Leu 58 and Cys 60 appearing to be a major determinant of specificity by binding the key buried Ala 7. These findings predict which other chromodomains will bind methylated proteins and suggest a motif that they recognize.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000)
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001)
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001)
Nakayama, J., Rice, J. C., Strahl, B. D., Allis, C. D. & Grewal, S. I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001)
Zhang, Y. & Reinberg, D. Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev. 15, 2343–2360 (2001)
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000)
Noma, K., Allis, C. D. & Grewal, S. I. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293, 1150–1155 (2001)
Litt, M. D., Simpson, M., Gaszner, M., Allis, C. D. & Felsenfeld, G. Correlation between histone lysine methylation and developmental changes at the chicken β-globin locus. Science 293, 2453–2455 (2001)
Nielsen, S. J. et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 412, 561–565 (2001)
Ball, L. J. et al. Structure of the chromatin binding (chromo) domain from mouse modifier protein 1. EMBO J. 16, 2473–2481 (1997)
Brasher, S. V. et al. The structure of mouse HP1 suggests a unique mode of single peptide recognition by the shadow chromo domain dimer. EMBO J. 19, 1587–1597 (2000)
Horita, D. A., Ivanova, A. V., Altieri, A. S., Klar, A. J. & Byrd, R. A. Solution structure, domain features, and structural implications of mutants of the chromo domain from the fission yeast histone methyltransferase Clr4. J. Mol. Biol. 307, 861–870 (2001)
Cowieson, N. P., Partridge, J. F., Allshire, R. C. & McLaughlin, P. J. Dimerisation of a chromo shadow domain and distinctions from the chromodomain as revealed by structural analysis. Curr. Biol. 10, 517–525 (2000)
Akhtar, A., Zink, D. & Becker, P. B. Chromodomains are protein–RNA interaction modules. Nature 407, 405–409 (2000)
Jacobs, S. A. et al. Specificity of the HP1 chromo domain for the methylated N-terminus of histone H3. EMBO J. 20, 5232–5241 (2001)
Platero, J. S., Hartnett, T. & Eissenberg, J. C. Functional analysis of the chromo domain of HP1. EMBO J. 14, 3977–3986 (1995)
Means, G. E. & Feeney, R. E. Reductive alkylation of amino groups in proteins. Biochemistry 7, 2192–2201 (1968)
Ferentz, A. E. & Wagner, G. NMR: A multifaceted approach to macromolecular structure. Q. Rev. Biophys. 33, 29–65 (2000)
Zwahlen, C. et al. Methods for measurement of intermolecular NOEs by multinuclear NMR spectroscopy: Application to a bacteriophage λ N-peptide/boxB RNA complex. J. Am. Chem. Soc. 119, 6711–6721 (1997)
Melacini, G. Separation of intra- and intermolecular NOEs through simultaneous editing and J-compensated filtering: A quadrature-free constant-time J-resolved approach. J. Am. Chem. Soc. 122, 9735–9738 (2000)
Kraulis, P. J. et al. The solution structure and dynamics of ras p21.GDP determined by heteronuclear three- and four-dimensional NMR spectroscopy. Biochemistry 33, 3515–3531 (1994)
Brunger, A. T. et al. Crystallography and NMR system (CNS): A new software system for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Linge, J. P., O'Donoghue, S. J. & Nilges, M. Automated assignment of ambiguous NOEs with ARIA. Methods Enzymol. 339, 71–90 (2001)
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biol. NMR 13, 289–302 (1999)
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991)
Merrit, E. A. & Murphy, M. E. P. Raster3D Version 2.0. A program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994)
Barton, G. J. ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng. 6, 37–40 (1993)
Acknowledgements
We thank A. Verreault for the pGEX-H3 construct; R. Turner, P. Sharratt and the PNAC facility for mass spectrometry and amino-acid analysis; A. Thiru for comments on the manuscript; the Medical Research Council for a Career Development Award (to H.R.M.); and the Wellcome Trust for financial support. The Cambridge Centre for Molecular Recognition is supported by the Biotechnology and Biological Sciences Research Council and the Wellcome Trust.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests
Rights and permissions
About this article
Cite this article
Nielsen, P., Nietlispach, D., Mott, H. et al. Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 416, 103–107 (2002). https://doi.org/10.1038/nature722
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature722
This article is cited by
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
Keep quiet: the HUSH complex in transcriptional silencing and disease
Nature Structural & Molecular Biology (2024)
-
Structural insights into the binding mechanism of Clr4 methyltransferase to H3K9 methylated nucleosome
Scientific Reports (2024)
-
Methylation across the central dogma in health and diseases: new therapeutic strategies
Signal Transduction and Targeted Therapy (2023)
-
Dynamics of the HP1 Hinge Region with DNA Measured by Site-Directed Spin Labeling-EPR Spectroscopy
Applied Magnetic Resonance (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.