Abstract
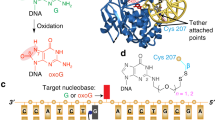
Spontaneous oxidation of guanine residues in DNA generates 8-oxoguanine (oxoG). By mispairing with adenine during replication, oxoG gives rise to a G·C → T·A transversion, a frequent somatic mutation in human cancers. The dedicated repair pathway for oxoG centres on 8-oxoguanine DNA glycosylase (hOGG1), an enzyme that recognizes oxoG·C base pairs, catalysing expulsion of the oxoG and cleavage of the DNA backbone. Here we report the X-ray structure of the catalytic core of hOGG1 bound to oxoG·C-containing DNA at 2.1 Å resolution. The structure reveals the mechanistic basis for the recognition and catalytic excision of DNA damage by hOGG1 and by other members of the enzyme superfamily to which it belongs. The structure also provides a rationale for the biochemical effects of inactivating mutations and polymorphisms in hOGG1. One known mutation, R154H, converts hOGG1 to a pro-mutator by relaxing the specificity of the enzyme for the base opposite oxoG.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Lindahl, T. & Wood, R. D. Quality control by DNA repair. Science 286, 1897–1905 (1999).
Michaels, M. L. & Miller, J. H. The GO system protects organisms from the mutagenic effect of the spontaneous lesion 8-hydroxyguanine (7,8-dihydro-8-oxoguanine). J. Bacteriol. 174, 6321–6325 (1992).
Grollman, A. P. & Moriya, M. Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet. 9, 246–249 (1993).
Hollstein, M. et al.Somatic point mutations in the p53 gene of human tumors and cell lines: updated compilation. Nucleic Acids Res. 24, 141–146 (1996).
Lu, R., Nash, H. M. & Verdine, G. L. A mammalian DNA repair enzyme that excises oxidatively damaged guanines maps to a locus frequently lost in lung cancer. Curr. Biol. 7, 397–407 (1997).
Arai, K. et al.Cloning of a human homolog of the yeast OGG1 gene that is involved in the repair of oxidative DNA damage. Oncogene 14, 2857–2861 (1997).
Roldan-Arjona, T. et al.Molecular cloning and functional expression of a human cDNA encoding the antimutator enzyme 8-hydroxyguanine-DNA glycosylase. Proc. Natl Acad. Sci. USA 94, 8016–8020 (1997).
Bjoras, M. et al.Opposite base-dependent reactions of a human base excision repair enzyme on DNA containing 7,8-dihydro-8-oxoguanine and abasic sites. EMBO J. 16, 6314–6322 (1997).
Rosenquist, T. A., Zharkov, D. O. & Grollman, A. P. Cloning and characterization of a mammalian 8-oxoguanine DNA glycosylase. Proc. Natl Acad. Sci. USA 94, 7429–7434 (1997).
Radicella, J. P., Dherin, C., Desmaze, C., Fox, M. S. & Boiteux, S. Cloning and characterization of hOGG1, a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 94, 8010–8015 (1997).
Kuo, F. C. & Sklar, J. Augmented expression of a human gene for 8-oxoguanine DNA glycosylase (MutM) in B lymphocytes of the dark zone in lymph node germinal centers. J. Exp. Med. 186, 1547–1556 (1997).
Aburatani, H. et al.Cloning and characterization of mammalian 8-hydroxyguanine-specific DNA glycosylase/apurinic, apyrimidinic lyase, a functional mutM homologue. Cancer Res 57, 2151–2156 (1997).
van der Kemp, P. A., Thomas, D., Barbey, R., de Oliveira, R. & Boiteux, S. Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc. Natl Acad. Sci. USA 93, 5197–5202 (1996).
Bruner, S. D., Nash, H. M., Lane, W. S. & Verdine, G. L. Repair of oxidatively damaged guanine in Saccharomyces cerevisiae by an alternative pathway. Curr. Biol. 8, 393–403 (1998).
Klungland, A. et al.Accumulation of premutagenic DNA lesions in mice defective in removal of oxidative base damage. Proc. Natl Acad. Sci. USA 96, 13300–13305 (1999).
Takao, M., Aburatani, H., Kobayashi, K. & Yasui, A. Mitochondrial targeting of human DNA glycosylases for repair of oxidative DNA damage. Nucleic Acids Res. 26, 2917–2922 (1998).
Nishioka, K. et al.Expression and differential intracellular localization of two major forms of human 8-oxoguanine DNA glycosylase encoded by alternatively spliced OGG1 mRNAs. Mol. Biol. Cell 10, 1637–1652 (1999).
Nash, H. M. et al.Cloning of a yeast 8-oxoguanine DNA glycosylase reveals the existence of a base-excision DNA-repair protein superfamily. Curr. Biol. 6, 968–980 (1996).
Shinmura, K., Kasai, H., Sasaki, A., Sugimura, H. & Yokota, J. 8-Hydroxyguanine (7,8-dihydro-8-oxoguanine) DNA glycosylase and AP lyase activities of hOGG1 protein and their substrate specificity. Mutat. Res. 385, 75–82 (1997).
Chevillard, S. et al.Mutations in OGG1, a gene involved in the repair of oxidative DNA damage, are found in human lung and kidney tumours. Oncogene 16, 3083–3086 (1998).
Dherin, C., Radicella, J. P., Dizdaroglu, M. & Boiteux, S. Excision of oxidatively damaged DNA bases by the human alpha-hOgg1 protein and the polymorphic alpha-hOgg1(Ser326Cys) protein which is frequently found in human populations. Nucleic Acids Res. 27, 4001–4007 (1999).
Shinmura, K. et al.Infrequent mutations of the hOGG1 gene, that is involved in the excision of 8-hydroxyguanine in damaged DNA, in human gastric cancer. Jpn. J. Cancer Res. 89, 825–828 (1998).
Blons, H. et al.Frequent allelic loss at chromosome 3p distinct from genetic alterations of the 8-oxoguanine DNA glycosylase 1 gene in head and neck cancer. Mol. Carcinog. 26, 254–260 (1999).
Nash, H. M., Lu, R., Lane, W. S. & Verdine, G. L. The critical active-site amine of the human 8-oxoguanine DNA glycosylase, hOgg1: direct identification, ablation and chemical reconstitution. Chem. Biol. 4, 693–702 (1997).
Thayer, M. M., Ahern, H., Xing, D., Cunningham, R. P. & Tainer, J. A. Novel DNA binding motifs in the DNA repair enzyme endonuclease III crystal structure. EMBO J. 14, 4108–4120 (1995).
Labahn, J. et al.Structural basis for the excision repair of alkylation-damaged DNA. Cell 86, 321–329 (1996).
Kuo, C. F. et al.Atomic structure of the DNA repair [4Fe-4S] enzyme endonuclease III. Science 258, 434–440 (1992).
Guan, Y. et al.MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nature Struct. Biol. 5, 1058–1064 (1998).
Yamagata, Y. et al.Three-dimensional structure of a DNA repair enzyme, 3-methyladenine DNA glycosylase II, from Escherichia coli. Cell 86, 311–319 (1996).
Dodson, M. L., Michaels, M. L. & Lloyd, R. S. Unified catalytic mechanism for DNA glycosylases. J. Biol. Chem. 269, 32709–32712 (1994).
Kohno, T. et al.Genetic polymorphisms and alternative splicing of the hOGG1 gene, that is involved in the repair of 8-hydroxyguanine in damaged DNA. Oncogene 16, 3219–3225 (1998).
Choi, J. Y. et al.Thermolabile 8-hydroxyguanine DNA glycosylase with low activity in senescence-accelerated mice due to a single-base mutation. Free Radical Biol. Med. 27, 848–854 (1999).
Barrett, T. E. et al.Crystal structure of a thwarted mismatch glycosylase DNA repair complex. EMBO J. 18, 6599–6609 (1999).
Parikh, S. S. et al.Base excision repair initiation revealed by crystal structures and binding kinetics of human uracil-DNA glycosylase with DNA. EMBO J. 17, 5214–5226 (1998).
Lau, A. Y., Scharer, O. D., Samson, L., Verdine, G. L. & Ellenberger, T. Crystal structure of a human alkylbase-DNA repair enzyme complexed to DNA: mechanisms for nucleotide flipping and base excision. Cell 95, 249–258 (1998).
Verdine, G. L. & Bruner, S. D. How do DNA repair proteins locate damaged bases in the genome? Chem. Biol. 4, 329–334 (1997).
Van Duyne, G. D., Standaert, R. F., Karplus, P. A., Schreiber, S. L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Otwinowski, Z. M. W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1996).
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D Biol. Crystallogr. 55, 849-61 (1999).
Brunger, A. T. et al.Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Brunger, A. Assessment of phase accuracy by cross validation: the free R value. Methods and applications. Acta Crystallogr. 49, 24–36 (1993).
Carson, M. J. Ribbons 2.0. J. Appl. Crystallogr. 24, 379–384 (1991).
Nicholls, A., Sharp, K. A., Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Laskowski, R. J., Macarthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–290 (1993).
Acknowledgements
We thank E. Lawson for assisting in DNA synthesis and protein expression; H. Huang, Q. Fan, L. Chen, S. Ealick, A. Lau, S. Liemann, H. Nash and members of the Verdine, S. C. Harrison and D.C. Wiley laboratories for helpful discussions; C. Schafmeister, R. Crouse, M. Przetakiewicz, R. Sweet, C. Metz and the staff of the CHESS A1 and NSLS X12C synchrotron beamlines for invaluable help with data collection; and K. Haushalter and X. Zhang for critical reading of this manuscript. This research was funded by the NIH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bruner, S., Norman, D. & Verdine, G. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000). https://doi.org/10.1038/35002510
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35002510
This article is cited by
-
Differential Binding of NLRP3 to non-oxidized and Ox-mtDNA mediates NLRP3 Inflammasome Activation
Communications Biology (2023)
-
CRISPR/Cas12a trans-cleavage triggered by cleavage ligation of dumbbell DNA for specific detection of human 8-oxoguanine DNA glycosylase activity
Microchimica Acta (2023)
-
Copper-mediated DNA damage caused by purpurin, a natural anthraquinone
Genes and Environment (2022)
-
CRISPR–Cas9 bends and twists DNA to read its sequence
Nature Structural & Molecular Biology (2022)
-
8-Oxoguanine: from oxidative damage to epigenetic and epitranscriptional modification
Experimental & Molecular Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.