Abstract
Identification and characterization of molecular mechanisms that connect genetic risk factors to initiation and evolution of disease pathophysiology represent major goals and opportunities for improving therapeutic and diagnostic outcomes in Alzheimer's disease (AD). Integrative genomic analysis of the human AD brain transcriptome holds potential for revealing novel mechanisms of dysfunction that underlie the onset and/or progression of the disease. We performed an integrative genomic analysis of brain tissue–derived transcriptomes measured from two lines of mice expressing distinct mutant AD-related proteins. The first line expresses oligomerogenic mutant APPE693Q inside neurons, leading to the accumulation of amyloid beta (Aβ) oligomers and behavioral impairment, but never develops parenchymal fibrillar amyloid deposits. The second line expresses APPKM670/671NL/PSEN1Δexon9 in neurons and accumulates fibrillar Aβ amyloid and amyloid plaques accompanied by neuritic dystrophy and behavioral impairment. We performed RNA sequencing analyses of the dentate gyrus and entorhinal cortex from each line and from wild-type mice. We then performed an integrative genomic analysis to identify dysregulated molecules and pathways, comparing transgenic mice with wild-type controls as well as to each other. We also compared these results with datasets derived from human AD brain. Differential gene and exon expression analysis revealed pervasive alterations in APP/Aβ metabolism, epigenetic control of neurogenesis, cytoskeletal organization and extracellular matrix (ECM) regulation. Comparative molecular analysis converged on FMR1 (Fragile X Mental Retardation 1), an important negative regulator of APP translation and oligomerogenesis in the post-synaptic space. Integration of these transcriptomic results with human postmortem AD gene networks, differential expression and differential splicing signatures identified significant similarities in pathway dysregulation, including ECM regulation and neurogenesis, as well as strong overlap with AD-associated co-expression network structures. The strong overlap in molecular systems features supports the relevance of these findings from the AD mouse models to human AD.
Similar content being viewed by others
Introduction
Integrative genomic analysis of the human Alzheimer’s disease (AD) brain transcriptome offers potential for revealing patterns of molecule or pathway dysfunction that underlie the onset and progression of AD.1 One would predict that these different stages of disease pathogenesis might display spreading and evolving molecular pathology in the same way that Braak and Braak stages define spreading and evolving histological pathology.2 Previous genetics and integrative genomics studies of human AD brain tissues converged on components of the microglial phagocytic system specified, respectively, by either the TREM2 cell surface protein3, 4 or by its intracellular adaptor, DAP12/TYROBP.5 With significant integrative genomic efforts underway to map networks underlying the onset and progression of human AD, there is a need to map molecular signatures and networks of AD animal models. Further, there is a need to develop a ‘systems understanding’ of animal models of AD and to understand molecular networks and activities shared and distinct between both individual models and also between animal models and human AD.
We undertook a study of the transcriptomes of the brains of two lines of transgenic mice expressing mutant AD-related proteins. The first line of mice expresses oligomerogenic mutant APPE693Q driven in neurons by the Thy-1 promoter, leading to the accumulation of amyloid beta (Aβ) oligomers and marked intracellular accumulation of APP/Aβ-like immunoreactivity.6 This amyloid mutation, also known as the Dutch mutation, causes cerebral amyloid angiopathy and accumulation of diffuse Aβ deposits in humans.7 These mice develop behavioral impairment as a function of the levels of Aβ oligomers.6 There are structural abnormalities of synapses8 but parenchymal amyloid plaques are never observed in these mice up to 24 months of age. The second line of mice expresses APPKM670/671NL/PSEN1Δexon9 in neurons and accumulates fibrillar amyloid in the interstitial spaces of the brain that goes on to form typical amyloid plaques accompanied by neuritic dystrophy and abnormalities in spatial memory.9 Notably, the APPKM670/671NL/PSEN1Δexon9 mouse accumulates both oligomers and fibrils composed of Aβ1–42, with the level of inflammogens correlating to the levels of the oligomers and not the levels of plaques and/or neurites.10
We chose these mouse lines because they each display impaired learning behavior despite the presence of quite distinct Aβ conformations and pathologies.6, 9 Accumulation of human Aβ1-42 in APPKM670/671NL/PSEN1Δexon9 transgenic mice is associated with robust deposition of parenchymal amyloid plaques surrounded by neuritic dystrophy as well as cerebral amyloid angiopathy.9 Oligomerogenic APPE693Q contains the pro-oligomerogenic E to Q mutation at position 22 of Aβ. It should be noted that owing to this E to Q mutation, the oligomers associated with each strain are not exactly comparable. Our Dutch APPE693Q transgenic mice express the epitopes of oligomeric Aβ recognized by antibodies A11, OC and NU4 (ref. 11, 12) (Knight et al., unpublished data). Aβ oligomers warrant particular attention because they may be the most important mediators of neuronal cell loss in the human AD brain.13 Although neither line of mice from the present study displays neuronal loss, both show impairment in learning behavior, indicating that amyloid plaques are not required for the appearance of clinically detectable behavioral changes.
We describe a comparative integrative genomic analysis of brain-derived transcriptomic profiles of mice accumulating oligomers of AβE22Q when compared with those of mice accumulating both oligomers and fibrils of Aβ1-42. We compare these data with existing knowledge of the molecular and histological pathogenesis for each model. For example, dysregulation of extracellular matrix (ECM) regulating genes was observed in APPKM670/671NL/PSEN1Δexon9 transgenic mice but not in APPE693Q transgenic mice, consistent with the presence of the obvious physical ECM perturbations associated with amyloid fibril accumulation. We also explore the concordance of our transcriptome profiles with differential expression (DE) and splicing profiles, gene co-expression and Bayesian networks that were built from human late onset AD (LOAD) postmortem brain samples.5 We identified significant overlap with LOAD-associated transcriptional and subnetwork changes, as well as similarities in the pathways that were implicated. We discuss this and other evidence for differential activation of transcriptomic pathways in association with accumulation of either AβE22Q oligomers or Aβ1-42 fibrils plus oligomers.
Materials and methods
Animals
The creation of the APPE693Q mouse was previously described.6 APPKM670/671NL/PSEN1Δexon9 mice were obtained from Jackson Laboratories (Bar Harbor, ME, USA). Both lines were maintained on a C57Bl6/J background. Animals were maintained on a 12:12 light:dark cycle with ad libitum access to food and water throughout the course of the entire experiment. The experimental protocol was conducted within NIH guidelines for animal research and was approved by the Institutional Animal Care and Use Committee (IACUC) at Icahn School of Medicine at Mount Sinai. Male 12-month-old mice were killed by CO2 asphyxiation and decapitation. DG and ECs were collected, snap frozen and stored at −80 °C prior to RNA isolation.
The minimum number of samples necessary to adequately detect DE was determined using RNASeqPower.14 Estimating that the average depth of coverage from RNA sequencing would be 20 reads per gene, the biological coefficient of variation as 0.1 and setting alpha to 0.05, we determined that at least three samples per comparison group would be necessary to detect expression differences with an effect-size of 2, with a power of 0.9.
RNA isolation
Snap-frozen DG and ECs were homogenized in QIAzol Lysis Reagent (Qiagen, Valencia, CA, USA). Total RNA purification was performed with the RNeasy Micro kit (Qiagen), and was carried out according to the manufacturer’s instructions. RNA quantification and quality was evaluated by Agilent BioAnalyzer and processed for RNA library preparation. RNA integrity was checked by either the Fragment Analyzer (Advanced Analytical, Ankeny, IA, USA) or the 2100 Bioanalyzer using the RNA 6000 Nano assay (Agilent, Santa Clara, CA, USA). All processed total RNA samples had an RNA quality number/RNA integrity number value of 6.1 or greater.
Real-time qPCR
Quantitative real-time PCR (qRT-PCR) was performed with the same RNA extracts that were used for RNA-Seq analysis. Top six differentially expressed (DE) genes identified by RNAseq in entorhinal cortex (EC) of APPE693Q and dentate gyrus (DG) of APPKM670/671NL/PSEN1Δexon9 were selected for qRT-PCR. Five hundred nanograms of RNA were reversed-transcribed using the High Capacity RNA-to-cDNA Kit (Applied Biosystems, Foster City, CA, USA). The cDNA solution was subjected to real-time qPCR in a Step-One Plus system (Applied Biosystems) using the PerfeCTa SYBR Green FastMix ROX (Quanta BioSciences, Gaithersburg, MD, USA). Quantitative PCR consisted of 40 cycles, 15 s at 95 °C and 30 s at 60 °C each, followed by dissociation curve analysis.
Library preparation
The sequencing library was prepared with the standard TruSeq RNA Sample Prep Kit v2 protocol (Illumina, San Diego, CA, USA). Briefly, ribosomal RNA was removed using the Ribo-Zero rRNA Removal Kit (Human/Mouse/Rat) (Illumina). The remaining RNA was then fragmented, and the cDNA was synthesized using random hexamers, end-repaired and ligated with appropriate adaptors for sequencing. The library then underwent size selection and purification using AMPure XP beads (Beckman Coulter, CA, USA). The appropriate Illumina-recommended 6 bp barcode bases were introduced at one end of the adaptors during the PCR amplification step. The size and concentration of the RNAseq libraries was measured by Bioanalyzer and Qubit fluorometry (Life Technologies, Grand Island, NY, USA) before loading onto the sequencer. The rRNA-depleted libraries were sequenced on the Illumina HiSeq 2500 System with 100 nucleotide single-end reads, according to the standard manufacturer's instructions (Illumina).
Read alignment and gene expression counts
Single end fastq files for 28 samples were aligned to a modified version of Mouse Reference genome (mm10),15 using STAR read aligner.16 To allow discrimination between the human APP transgene, and the mouse App, we appended the human APP sequence as a dummy chromosome to the mm10.fasta file, and appended the ensemble v70 APP gene models to the gtf used to build the genome index used by STAR. Accepted mapped reads were summarized separately to gene and exon levels using the featureCounts function of subread.17, 18
Differential expression analysis
Gene count matrices were produced separately for each of the six primary comparisons (oligomerogenic APPE693Q vs wild type (EC and DG), fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild type (EC and DG) and fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q mouse (EC and DG). In each case, the assembled count matrix was filtered to retain transcripts with a summed count across samples of at least 7 (the quantity of samples in each comparison). The count matrix was corrected for library size, batch status and normalized using DESeq2.19 DE analysis was performed also using DESeq2, and P-values were adjusted using the Benjamini-Hochberg method.20
Gene set enrichment analysis
Gene set enrichments were calculated using two different methods, as indicated throughout the text. GAGE21 was used for whole-transcriptome region-based comparisons between groups of samples. Gene count matrices were normalized using DESeq2,19 and gene set enrichments were calculated using settings same.dir=FALSE, and compare=’unpaired’. Gene set enrichments for discrete groups of genes (that is, significance thresholded DE results, genes containing differentially spliced exons and human LOAD co-expression networks) were calculated using Fisher’s exact text, and one-sided P-values (to identify overrepresentation of gene sets) were adjusted using the Benjamini-Hochberg method.20
Gene sets used throughout the enrichment analysis were derived from a combination of publicly available sources, such as the molecular signatures database,22 brain-specific gene sets curated from publicly available data,23 protein-protein hubs interactor sets24 and ChipSeq-based transcription factor target sets.25
All gene sets were derived from human data. For comparison against the mm10 ensembl transcriptome, we converted from mouse to available human orthologs using the MGI vertebrate homology classes,26 and to human ensembl gene identifiers using the R package org.Hs.eg.db.27
Differential exon expression analysis
Exon count matrices were produced separately for each of the six primary comparisons (oligomerogenic APPE693Q vs wild type (EC and DG), fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild type (EC and DG) and fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q mouse (EC and DG). For comparison of mutant line samples to wild type, we used DEXSeq,28 which allows detailed control over visualization of results. The comparisons of fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q mice required more detailed matrix models to account for our batch structure co-varying with phenotype. Each line was derived from a separate batch (where each batch also included wild-type samples). This required correction of batch status while including wild-type samples (to distinguish between the batch effect and the strain effect), and later subsetting to only the oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse samples for the differential exon analysis. For this, we used diffSplice function in the limma R package.29, 30, 31
DEX profiles from human LOAD were generated using exon count matrices from postmortem samples collected across three brain regions. Each region was processed separately, and in each case, comparison was made between samples annotated as ‘Definite AD’ vs ‘Normal’ (CERJ score of 2 vs 1), including age and gender as covariates. Differentially spliced genes were identified using the diffSplice function in the limma R package.29, 30, 31
Code availability
Code used to generate DE and DEX results is available upon request.
Results
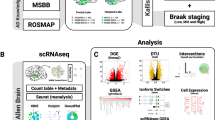
We generated transcriptomic profiles for oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice and characterized each model in comparison with wild-type mice and each other (Figure 1). We performed next-generation RNA sequencing on 28 samples from EC and DG.
Schematic overview of mouse AD transcriptome analysis. RNA sequencing was performed on the entorhinal cortex and dentate gyrus for three groups of animals (oligomerogenic APPE693Q, fibrillogenic APPKM670/671NL/PSEN1Δexon9 and wild type) comprising a total of 28 samples (Transgenic n=3 and Wild type n=4 samples per comparison). Region-based differential gene and exon expression analysis was performed between all mouse lines, and results were annotated with diverse functional molecular data.
Differential gene expression analysis of transgenic vs wild-type mice
We assessed the regional DE between oligomerogenic APPE693Q vs wild type, APPKM670/671NL/PSEN1Δexon9 vs wild type and APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q mice. In this comparison against the wild-type animals, we identified 354 DE genes (false discovery rate (FDR)<0.05) in the DG of the fibrilllogenic APPKM670/671NL/PSEN1Δexon9 mice, and 22 DE genes (FDR<0.05) in the EC of the oligomerogenic APPE693Q mice (Figure 2). We did not observe any DE genes in the EC of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice or in the DG of the oligomerogenic APPE693Q mice (see Supplementary Table 1 for full DE results). To investigate whether this difference in the amount of detected DE across comparisons related to differential transgene expression, we examined the expression for the human APP sequence across mouse strains. We found that in both regions and lines, there was evidence of increased expression of reads that mapped to the human APP sequence (see Supplementary Figure S1), though we did not see a clear relationship between human APP expression and the quantity of genes identified as being differentially expressed.
Differential gene expression and enrichment analysis summary. (a) Differentially expressed genes in the entorhinal cortex of oligomerogenic APPE693Q vs wild-type mice. (b) Top differentially expressed genes in the dentate gyrus of fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mice. (c) Quantity of differentially expressed genes and selected GO term enrichments shared across regional comparisons of fibrillogenic APPKM670/671NL/PSEN1Δexon9 and oligomerogenic APPE693Q vs wild-type mice. (d) Quantity of differentially expressed genes and selected GO term and KEGG pathway enrichments shared across regional comparisons of fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q mice. Enrichments shown were selected for known or suspected relevance to AD pathophysiology, and bolding highlights enrichments that relate to the main biological themes also implicated by the differential exon analysis findings. (Differential expression and gene set enrichments thresholded at FDR<0.05.) FDR, false discovery rate.
We performed qPCR on the top six DE genes for the EC of oligomerogenic APPE693Q and the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9. We saw good overall concordance between the log2 fold change as detected by RNAseq and qPCR, with a Pearson correlation of 0.95 for the EC of oligomerogenic and 0.62 for the DG of fibrillogenic mice (see Supplementary Figure S2).
To identify biological pathways that may be differentially regulated across mouse lines, we used GAGE20 and performed gene set enrichment analysis. We identified Gene Ontology32 and KEGG pathway33, 34 enrichments that are either shared or exclusive to either brain regions and/or mouse line (Figure 2c, see Supplementary Table 2 for full enrichment results). Similar biological themes were implicated in both brain regions as well as in both mutant mouse lines in comparison with wild type, including cytoskeletal organization, microtubule processes, glutamatergic signaling and neuronal differentiation. We also observed a dysregulation of protein-protein interactions involving APP and/or Aβ in the DG of both lines that was not present in the EC of either line. We did not observe shared EC-specific enrichments between the two transgenic mouse lines.
We performed an additional enrichment analysis of the DE gene sets comparing against an expanded library of public and privately curated gene sets, including brain-specific gene signatures,23 protein-protein hub interactions,24 ChipSeq transcription factor gene targets,25 and the molecular signatures database.22 (See Supplementary Table 3 for full enrichment results). Intersection of the DE gene sets with curated brain-specific gene sets23 revealed that the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse had significant overlap with multiple hippocampal regional signatures as well as with a mixture of glutamatergic neuronal and microglial gene sets. We also observed enrichment for genes reported as upregulated in the CA1 hippocampal region of AD patients as compared against age-matched non-demented subjects35 (see Supplementary Table 3).
Analysis of the DE genes for the oligomerogenic APPE693Q transgenic mouse EC identified significant overlaps with multiple hippocampal, microglial, glutamatergic and interneuronal gene sets. We also observed enrichment for genes reported in multiple studies as upregulated in the CA1 hippocampal region of postmortem brain from AD patients as compared with age-matched, non-demented subjects35, 36, 37 (see Supplementary Table 3).
Integration of DE genes with a library of protein-protein interactions24 highlighted APP as a protein hub with an enrichment of DE genes among its known interactors, in both the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse DG and the oligomerogenic APPE693Q transgenic mouse EC gene sets (Figure 3b). The DE analysis of fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse DG also highlighted multiple other transcripts of hub proteins (see Supplementary Table 3), including ITGB1, an integrin molecule with important roles in cell adhesion, as well as forming a microglia receptor complex that mediates detection, phagocytosis and clearance of fibrillar Aβ.38, 39, 40 This is especially notable in view of recent evidence linking AD to TREM2 and its adaptor protein, DAP12/TYROBP.5
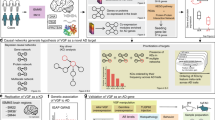
Multiregion transcriptome comparisons between fibrillogenic, oligomerogenic and wild-type mice implicates amyloid/Aβ processing, extracellular matrix (ECM) regulation and neurogenesis (a, b–i) Fragile X Mental Retardation 1 (FMR1) gene is differentially spliced in fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q dentate gyrus (also vs wild type), as well as multiple brain regions in LOAD and (b-ii) is a known regulator of APP, binding to mRNA in the post-synaptic neuron in an mGluR5 stimulation-dependent manner. (b-iii) DE genes in both comparisons with wild type (see Figure 2), are enriched for known protein interactors of APP. APP interactors that are DE in the fibrillogenic APPKM670/671NL/PSEN1Δexon9DG vs wild type are shown. (b-iv) Adaptor protein GRB2 is differentially spliced in fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q dentate gyrus, and interacts with APP and PSEN1, localized to the centrosomes, resulting in ERK1/2 activation, and potentiation of oligomer-induced toxicity. (c) ECM regulation was a recurring theme of the pathway analysis following differential gene and exon expression analysis. (c-i) Known ECM regulators that are differentially expressed in fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mice (dentate gyrus) suggest mechanisms of perturbation and compensation. (c-ii) Gene Ontology (GO) enrichment analysis of the 354 genes that are differentially expressed in fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mice (dentate gyrus) demonstrate that the trend toward ECM disruption is particularly strong in this comparison. (d) Pathway enrichment analysis of the differentially expressed genes in fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mice (dentate gyrus) indicates perturbation of stem cell, neural progenitor cell and neurogenesis pathways. (d-i) SUZ12 is a key member of the polycomb repressive complex 2 (PRC2), and is differentially spliced in fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q dentate gyrus (and also vs wild type). (d-ii) A functional role for SUZ12 is strongly supported by enrichment analysis of ChipSeq-based transcription factor gene targets, with the 354 differentially expressed genes in fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mice (dentate gyrus). (d-iii) SUZ12 function within the PRC2 is associated with regulation of neurogenic differentiation of stem cells via histone H3K27 and H3K9 methylation. (e-i) Zinc finger gene SP1 was identified as the transcription factor most strongly enriched for DEX genes (APPKM670/671NL/PSEN1Δexon9 vs wild-type comparison). (e-ii) SP1 is a transcriptional regulator of multiple AD-associated genes, and forms a potential link between these molecular nodes and the main DEX themes we have discussed, including perturbations in neurogenesis, amyloid processing and ECM regulation.
Differential gene expression analysis of oligomerogenic APPE693Q vs fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice
A direct comparison of the transcriptomes of fibrillogenic APPKM670/671NL/PSEN1Δexon9 against those of the oligomerogenic APPE693Q transgenic mice identified a single gene, cortexin-3 (CTXN3), that was upregulated in the DG of fibrilllogenic APPKM670/671NL/PSEN1Δexon9 mice. A recent genome-wide association meta-analysis aimed at finding genes linked to plasma Aβ identified CTXN3.41 In that study, CTXN3 was the gene most strongly associated with Aβ plasma levels in the high-risk population (that is, over age 65 years) and was subsequently validated as a modulator of Aβ metabolism.41
To identify relative perturbations that might help inform an understanding of the fibrillar pathology observed in APPKM670/671NL/PSEN1Δexon9 transgenic mice, we performed a comparison of Gene Ontology32 and KEGG pathway set33, 34 enrichments between the transcriptomes of APPKM670/671NL/PSEN1Δexon9 and oligomerogenic APPE693Q transgenic mice. In both the DG and the EC, we detected robust evidence for increased dysregulation of multiple pathways implicated in human LOAD, including GTPase/Ras protein regulation,42 cytoskeletal organization, glutamatergic receptor activity, neuronal differentiation and microtubule processes in the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice (Figure 2d). Cytoskeletal organization and cell junction/adhesion processes were also highlighted by the KEGG analysis. We observed additional EC-specific enrichments for other neurodegenerative diseases, including Huntington’s disease, amyotrophic lateral sclerosis and prion disease. In the DG, we observed specific enrichment for cell adhesion molecules and for olfactory transduction molecules (Figure 2d, see Supplementary Table 4 for full enrichment results).
Differential gene splicing analyses
We performed differential exon expression (DEX) analysis using DEXseq28 and Limma29, 30, 31 (Figure 4a) comparing oligomerogenic APPE693Q vs wild type, fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild type and fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q. Overall, the number of genes identified as containing DE exons mirrored the trends seen in the DE analysis, with the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice dominating the results, followed by the EC of oligomerogenic APPE693Q transgenic mice (Figure 4b). No differentially spliced genes were detected for the EC of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 nor for the DG of the oligomerogenic APPE693Q transgenic mouse line (see Supplementary Table 5 for full results). For any given comparison, we detected relatively little overlap between genes that are differentially spliced and those that are differentially expressed.
Differential exon expression and enrichment analysis summary. (a) Differential exon analysis comparisons between all three mouse lines. (b) Quantity of genes with differentially expressed exons (DEX) for fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice and oligomerogenic APPE693Q mice vs wild type. Enrichments for DEX genes in a library of curated brain-focused gene sets, highlighted for connection to Alzheimer's disease (AD), other neuropsychiatric disease (NPD) and cell/brain regional signatures (Cell) for (c) APPE693Q oligomerogenic vs wild-type mouse entorhinal cortex and (d) fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs wild-type mouse. (e) Quantity of DEX genes for fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice vs oligomerogenic APPE693Q mice across dentate gyrus (DG) and entorhinal cortex. (f) DEX genes for fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q mice in the DG, highlighted for multiple themes, including amyloid/Aβ regulation and processing (APP), association with AD risk, extracellular matrix regulation (ECM) and multiple others (MULTI). Concordance with DEX in human postmortem LOAD samples across the frontal pole (FP), superior temporal gyrus (STG) and parahippocampal gyrus (PHG) are also shown. (DEX genes with FDR<0.05 and gene set enrichments with FDR<0.1 are shown). FDR, false discovery rate.
Gene set enrichment analysis of each DEX gene set (fibrillogenic APPKM670/671NL/PSEN1Δexon9 DG and oligomerogenic APPE693Q EC vs wild type, FDR<0.05) highlighted several enrichments relevant to AD pathophysiology. Intersection of the datasets with brain-specific gene sets23 identified enrichments for DEX genes from oligomerogenic EC (Figure 4c), among genes that are upregulated in the hippocampal CA1 region of AD patients vs age-matched, non-demented subjects, and a mixture of microglial, oligodendrocyte, interneuronal and glutamatergic neuronal marker sets. The DEX genes from fibrillogenic DG (Figure 4d) revealed enrichment of these same signatures as well as an additional astrocyte gene signature. We also found that DEX from the fibrillogenic APPKM670/671NL/PSEN1Δexon9 DG vs wild type were enriched for genes containing the binding motifs for multiple zinc finger transcription factors (Figure 3e, see Supplementary Table 6 for full results), most notably SP1, MAZ, ZF5/ZBTB14, EGR1 and E4F1.
DEX analysis between fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice vs oligomerogenic APPE693Q transgenic mice identified 5 DEX genes in the EC and 23 DEX genes in the DG (Figures 4e and f, see Supplementary Table 7 for full list). The set of 23 DEX genes from the DG highlighted a number of genes with known or possible risk associations with AD, including TRPC4AP (identified in an extended pedigree haplotype analysis43) and CUL2 (part of multiple ubiquitin ligase complexes, located near an AD risk-associated locus on chromosome 10 (ref. 44).
Enrichment analysis of the DEX genes of these fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q transgenic mice (Figure 4f) revealed significant overlaps in the DG with genes downregulated in AD in the CA1 region of the hippocampus, as well as a mixture of glutamatergic and interneuronal cell signatures. DEX genes from the EC enriched for a co-expression module comprised of genes downregulated in AD, as well as a parvalbumin-positive interneuron-enriched co-expression module (see Supplementary Table 6 for full enrichment results). We observed that many of the DEX genes identified in the DG have well-documented associations with APP metabolism and processing (Figure 4f and Figure 3b). These include differential splicing of (i) FMR1, which encodes FMRP, a translational repressor protein associated with Fragile X syndrome, known to bind and regulate APP mRNA in synaptoneurosomes containing mGluR5 receptor;45 (ii) GRB2, an adaptor protein enriched around amyloid plaques in the AD brain, which, through its interactions with APP and PSEN1 (ref. 46) activates ERK1/2 signaling, thus mediating Aβ oligomer-induced toxicity;47 (iii) ANKS1B, a multi-domain protein expressed in brain and testes, known to bind and regulate APP processing in an isoform-specific manner, and, in some cases, thereby diminishing Aβ generation through inhibition of γ-secretase;48 and (iv) the sumoylation regulator SENP2, which catalyzes the conjugation/deconjugation of SUMO1 with SUMO target proteins, affecting Aβ regulation via a splicing-dependent interaction with BACE1.49
Some of the most striking enrichment themes that we identified involve dysregulation of histone methylation, neurogenesis, neural progenitor cell and stem cell pathways. We observed these enrichments across all comparisons, but most strongly in the DE genes of fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice (DG). SUZ12 was differentially spliced in the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice when compared against either oligomerogenic APPE693Q transgenic mice or against wild-type mice, suggesting it may have a key role. SUZ12 forms part of the Polycomb-repressive complex 2 (PRC2), exerting a repressive role on target gene expression and is critical for establishing programs required for differentiation of embryonic stem cells and regulation of neurogenic potential.50 To explore the possible functional role for SUZ12 as a regulator of transcriptional changes in the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice, we performed Chip-X enrichment analysis,25 overlapping the DE gene sets with ChipSeq derived transcription factor target gene sets. We found that the top four significant enrichments were for SUZ12 gene target sets derived from four separate experiments, strongly implicating SUZ12 as the transcription factor that best explained the DE signature (Figure 3d).
One of the pervasive pathway enrichment themes we observed across both regions and mutant mouse lines was the dysregulation of genes involved in cell adhesion, ECM regulation and cytoskeletal organization. This finding was particularly robust in the context of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse DG (Figure 2d and Figures 3e and f). A number of genes involved in ECM metabolism were identified in the DE analysis, including MMP17, CHSY3, ADAMTSL3, ADAM10, ADAMTS16 and ADAM12, and these pathways were highlighted in the DEX genes as well. We found that ABL2 was differentially spliced in the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice when compared with oligomerogenic APPE693Q transgenic mice (Figure 4f). ABL2 is a non-receptor tyrosine kinase, and increasingly considered to be an essential integrator of diverse extracellular cues into patterns of cytoskeletal rearrangements, cell motility and cell adhesion.51 EXT2 was also differentially spliced in the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice. EXT2 encodes a glycosyltransferase that complexes with EXT1 to catalyze the chain elongation step of heparan-sulfate proteoglycan synthesis. Heparan-sulfate proteoglycans bind to cell membranes and to the ECM, where they interact with a diverse array of protein ligands, regulating axonal guidance, ECM metabolism, cell adhesion, morphogenesis and cell migration (for review, see Sarrazin et al.52).
Comparison of differential gene splicing in mouse AD models vs human LOAD
Given the potential of the mouse DEX analysis to inform on novel aspects of human disease pathophysiology, we sought to compare our results with patterns of differential splicing across three brain regions (superior temporal gyrus, frontal pole and parahippocampal gyrus) in postmortem LOAD samples collected through the Mount Sinai NIH Brain and Tissue Repository.53 We found that of the 23 DEX genes identified in the fibrillogenic APPKM670/671NL/PSEN1Δexon9 vs oligomerogenic APPE693Q transgenic mice comparison, 8 were also differentially spliced in at least one of the three brain regions examined (Figure 4f, see Supplementary Table 8 for all significant DEX results). Perhaps most notably, FMR1, discussed above as a synaptic regulator of APP, was differentially spliced in LOAD in the frontal pole and superior temporal gyrus. We also note that ANKS1B, a master regulator of microtubule activity54 was differentially spliced in the superior temporal gyrus, and the parahippocampal gyrus, and AD risk-associated gene CUL2 (ref. 44) was differentially spliced in the superior temporal gyrus.
Comparison of gene expression in mouse AD models vs human LOAD
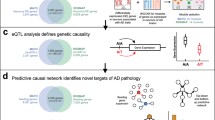
We accessed publicly available, postmortem gene expression profiles for six brain regions from a comparison across 34 individuals diagnosed with LOAD, along with 14 age-matched, non-demented controls37 to evaluate how the oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse transcriptomes approximate changes observed in human LOAD. We mapped human genes to mouse orthologs where available, and calculated the Spearman correlations between the gene expression log-fold-change (Figure 5a). We observed significant global similarity between all pairs of mouse DE, across all six LOAD brain regions. The oligomerogenic APPE693Q transgenic mouse EC was the profile most highly correlated with five of the six human LOAD signatures, and of equal similarity along with the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic EC, to the human LOAD hippocampal signature.
Enrichments for differentially expressed genes and differentially spliced genes with human LOAD signatures. (a) Similarity of AD model transcriptional changes with postmortem LOAD gene expression collected from six brain regions (AD vs non-demented controls). Spearman’s rho (shown in heatmap cells) reflects correlation between log2 fold change of orthologous mouse-human genes. All correlations were positive and significant. (b) Gene co-expression modules constructed from human postmortem brain samples (AD and non-demented controls) were intersected with DE and DEX gene sets, identifying multiple significant overlaps, including modules that are significantly differentially connected in LOAD. (c) Bayesian network built from human LOAD postmortem prefrontal cortex samples, subset by fibrillogenic APPKM670/671NL/PSEN1Δexon9 DE genes (FDR<0.1), and their immediate neighbors. TYROBP, the key driver in the subnetwork most strongly associated with LOAD status, remained the most strongly connected gene in this induced subnetwork and is shown here with its local network neighborhood (first and second degree neighbors). (DE and DEX genes with FDR<0.05 -unless otherwise stated- and gene co-expression module enrichments with FDR<0.1 are shown). FDR, false discovery rate.
We intersected the DE and DEX gene sets with gene networks built from over 1600 postmortem brain samples from LOAD and non-demented donors.5 Zhang et al.5 used paired gene expression and genotype data to build integrated co-expression and Bayesian networks, identifying key drivers responsible for regulating large parts of the network in a general and also in a disease-specific manner. A number of our DE and DEX signatures were overrepresented in the reported co-expression modules (Figure 5b), including the DE genes of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse DG, in the Zhang et al.5 ‘immune and microglia’ (Yellow) co-expression module, the module most strongly associated with LOAD status. We also found strong overlap with the LOAD-associated ‘extracellular matrix’ (Tan) module, and the ‘neurogenesis’ (Midnight-blue) modules. In addition, DEX genes in the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse DG (vs wild type), and the oligomerogenic APPE693Q transgenic mouse EC (vs wild type) are enriched for genes contained in multiple 'synaptic transmission’ annotated modules, including the LOAD-associated salmon module from Zhang et al.5 These data suggesting that the strong themes that we observed in these biological domains in the mouse models are consistent with those reported by Zhang et al.5 in their studies of human LOAD transcriptomes.
To characterize the relationship between the mouse AD transcriptome and LOAD network topology, we projected the DE genes of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse DG (at a relaxed threshold of FDR<0.1) on to the Zhang et al.5 ‘case-only’ Bayesian network, extracting a sub-graph of these genes and their immediate neighbors (Figure 5c). We further annotated these genes with DEX status (FDR<0.05), as well as association with ECM regulation and SUZ12, and visualized them in network context using Cytoscape.55, 56 TYROBP, which was identified as the strongest key driver of the LOAD network, remained the most strongly connected gene in this induced sub-graph, connecting to multiple other DE genes, as well SUZ12 targets and ECM-associated genes.
Discussion
We performed an integrative genomic study aimed at comparing the transcriptome in the DG and EC of an oligomerogenic APPE693Q transgenic mouse and of a second transgenic mouse model that accumulates both Aβ oligomers and neuritic amyloid plaques (fibrillogenic APPKM690/671NL/PSEN1Δexon9). Our goal was to characterize transcriptional perturbations that are either shared by both lines or unique to one line, and to compare these mouse model transcriptomes with corresponding human AD postmortem brain transcriptomes. Our approach involved the use of next-generation RNA sequencing to perform regional differential gene (DE) and exon (DEX) expression analysis, followed by comprehensive integrative genomic and network analysis to provide insight to the biological context for these results.
To identify biological processes that may be impacted across both the oligomer and plaque-laden clinicopathological syndromes, we performed gene set enrichment analysis. While we observed few common DE genes shared by the two models and the two brain regions when comparing against wild-type mice, we noted a number of functional enrichments that were shared across all comparisons, including many human AD-associated pathways such as cytoskeletal organization, glutamatergic signaling and neurogenesis. Furthermore, when we performed DE comparing the transcriptomes of fibrillogenic mice against those of oligomerogenic mice, although we only identified a single DE gene, we found that the pathway enrichments that had been shared by both lines when compared with wild-type mice (such as cytoskeletal organization and regulation of neuron differentiation) tended to be more severely dysregulated in the fibrillogenic mouse, supporting the relevance of these pathways to disease.
Differential splicing highlights mediators of APP/Aβ metabolism, neurogenesis, ECM regulators and microglial activation
The DEX results generated by comparison between the transcriptomes of fibrillogenic and oligomerogenic transgenic mice were particularly interesting, and because of their potential to inform on mechanisms that may mediate the neuropathology and behavioral changes of the fibrillogenic mouse, these formed the main focus of our study. We observed a number of robust themes that were concordant between the DEX and DE analysis, in particular APP/Aβ processing (Figures 4f and 3b), epigenetic dysregulation of neurogenesis (Figure 3d), cytoskeletal organization (Figure 3b) and ECM regulation (Figures 4f, 5b and 3c). It has been suggested that abnormal splicing may be a major mediator of AD pathophysiology.49
The overlap of many of these DEX genes with human LOAD differential splicing patterns (Figure 4f) is encouraging because it suggests that these findings reflect more than the immediate impact of human transgenes (where potential relevance to LOAD may be limited), but instead indicate a response to secondary network changes that have been induced by genetic perturbation. This also supports the utility of these mouse strains as model systems for translational studies and for further characterization of these molecules in the context of LOAD.
Mediators of APP/Aβ metabolism
DEX analysis identified multiple potential mediators of differential amyloid processing, including FMR1, which encodes a translational repressor protein associated with Fragile X syndrome, an inheritable form of intellectual disability. Westmark and Malter.45 reported that the protein product of FMR1 binds to APP mRNA in synaptoneurosomes containing the post-synaptic mGluR5 receptor and negatively regulates APP translation in a stimulation-dependent manner. FMR1-KO mice are also known to display elevated basal levels of both APP and Aβ.57 Given these interesting molecular overlaps between Fragile X syndrome and AD, Renoux et a l.58 recently studied FMR1 expression in AD but did not identify alteration of cortical or subcortical expression of FMR1 in APPKM670/671NL/PSEN1Δexon9 mice nor in the prefrontal cortex or cerebellum of human AD postmortem samples. We also did not detect DE of FMR1 in our comparisons, but we observed an overexpression of FMR1 exon 5, suggesting a role for the alternate splicing of FMR1 in mediating the alteration in amyloid metabolism observed in fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice, particularly in the context of glutamatergic signaling which was also reflected in the pathway enrichment analysis. Moreover, FMR1 is transcriptionally repressed by ZF5/ZBTB14,59 a zinc finger transcription factor that was strongly enriched for binding site motifs among the DEX genes of the fibrillogenic APPKM670/671NL/PSEN1Δexon9 DG. We also found that FMR1 was differentially spliced across multiple brain regions in postmortem LOAD samples, supporting its translational relevance, and also supporting the utility of the fibrillogenic mouse as a means to interrogate it further. To the best of our knowledge, this study is the first to report differential splicing of FMR1 in the context of AD. As discussed above, ANKS1B may alter APP metabolism and is a central player in microtubule tracking and regulation,54 and is also seen to be differentially spliced in human LOAD across multiple brain regions. SENP2 (differentially spliced in the frontal pole in human LOAD), and by extension its splicing regulator HNRNPLL, is potentially linked with both APP processing and glucose metabolism, the latter of which may also be associated with AD.60 This theme of differential regulation of APP/Aβ metabolism was also reflected at the DE level, where we observed an enrichment of known APP protein-protein interactors among the DE genes of the fibrillogenic mouse DG and the oligomerogenic mouse EC.
Mediators of neurogenesis
Some of the most striking enrichment themes that we identified across all comparisons included dysregulation of histone methylation, neurogenesis, neural progenitor cell and stem cell pathways, most strongly in the DE genes of fibrillogenic APPKM670/671NL/PSEN1Δexon9 mice (DG). SUZ12 emerged as a potential nexus for this following DEX analysis in which SUZ12 was found to be differentially spliced in the DG of fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mice when compared with either oligomerogenic APPE693Q transgenic mice or with wild-type mice. SUZ12 forms part of the Polycomb-repressive complex 2 (PRC2), exerting a repressive role on target gene expression and acting through methylation of histone-3 at lysine-9 and lysine-27. In addition, PRC2 functions as a recruitment protein for DNA methyltransferases, thus acting as a potential regulator of two epigenetic systems61 that are recognized as important mechanisms of gene expression dysregulation in AD.62, 63, 64 SUZ12 was also implicated as functionally relevant to our study at the transcriptional level, with strong enrichments for DE fibrillogenic genes among multiple ChipSeq-derived gene target sets for SUZ12. This concordance between DEX and DE suggests a novel mechanism underlying the dysregulation of neurogenesis and neural progenitor cells in the DG in these models. To our knowledge, this is the first report in which SUZ12 has been implicated in the pathogenesis of AD. We did not see evidence for differential splicing of SUZ12 in the human LOAD data utilized for this study, however, it is important to note that in the biological context implicated by the enrichment analysis, the links to the regulation of neurogenesis suggest that this may be specific to regions of neurogenesis (such as the DG), which were not included in the current study. We are working towards the profiling of neurogenesis-associated brain regions in human AD samples to validate the relevance of SUZ12 splicing alterations to human LOAD.
Mediators of ECM and cell adhesion processes
Enrichment for perturbation of genes involved in cell adhesion, ECM and extracellular attachments was a consistently strong theme, particularly in the fibrillogenic mouse DG (Figure 2d and Figure 3c). The ECM has a crucial role in cell migration and axon guidance. During normal aging and late-stage AD, the ECM indices in the brain are progressively upregulated.65 Using a proteomic analysis of hippocampal synaptosomes, Végh et al.66 recently described the dysregulation of ECM proteins in the fibrillogenic APPKM670/671NL/PSEN1Δexon9 transgenic mouse concurrent with the rise in levels of Aβ oligomers, and the decline in hippocampal LTP and contextual memory, but prior to histologically detectable amyloid deposition. We identified a number of genes involved in ECM metabolism in the DE analysis, including MMP17, CHSY3, ADAMTSL3, ADAM10, ADAMTS16 and ADAM12. The metallopeptidase ADAM10 is of particular note. This protease regulates multiple Notch-1 pathway genes, and is associated with surface shedding of neuronal proteins, including APP, and is the most important basal α-secretase in the brain.67 Postina et al.68 demonstrated that the activation of ADAM10 had a beneficial effect in APP transgenic mice, reducing amyloid burden and improving cognition, and that expression of a catalytically inactive mutant ADAM10 increased plaque deposition in these same APP transgenic mice. Suh et al.69 recently discovered two rare mutations in the prodomain of ADAM10 that associate with LOAD. Upregulation of ADAM10 in the fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse DG may represent a compensatory mechanism for cleaving APP and degrading Aß fibrils.
DEX analysis also identified multiple potential mediators of ECM perturbations, including ABL2, which is associated with cytoskeletal organization, neural development and synaptic plasticity, and AD-associated tau phosphorylation.70 The concordance of our results with those described by Végh et al.66 demonstrates that this finding is robust across experiments, and our study may assist in providing a transcriptional context for these data, as well as highlighting candidate molecular drivers of this ECM dysregulation, including ADAM10, ABL2, EXT2 and SUZ12.
Mediators of microglial activation
We identified enrichment for multiple microglial signatures in the DE signatures of fibrillogenic mouse DG, oligomerogenic mouse EC and in the DEX genes of oligomerogenic mouse EC. The role of microglia in AD pathology has long been of interest,71 and recent genetic studies have renewed and focused interest in microglial dysfunction in AD, including the identification of new risk loci at HLA-DRB5–HLA-DRB1 (ref. 72) and the role for AD risk-associated gene CD33 in regulation of microglial amyloid uptake and clearance.73 An exciting convergence was apparent in the identification of TREM2 (triggering receptor expressed on myeloid cells 2) as AD risk-associated,3, 4 which is bound by the DAP12/TYROBP adaptor protein, highlighted in a computational network biology approach5 as central to LOAD pathophysiology. In our study, we did not observe DE of TREM2, but did note an upregulation of TYROBP in fibrillogenic mouse DG.
The identification of differentially spliced SENP1 and HNRNPLL in the fibrillogenic mouse DG is also intriguing, in light of the role of HNRNPLL in regulation of CD45 splicing patterns, and the beneficial influence of SENP1 on HNRNPLL-mutant T cells. Given the overexpression of CD45 in AD microglia,71 the regulatory role of CD45 in microglial activation74 and the dynamic regulatory interactions between T cells and microglia,75 a role for SENP1 and HNRNPLL regulation of CD45 in AD microglia should be further explored.
IL10 is recently implicated in AD, following the observations of elevated IL10 signaling in human AD subjects, the improvement in Aß clearance and cognitive deficits in IL10-deficient AD mice76 and the inhibitory role of IL10 on microglial Aß phagocytosis.77 We did not observe DE of IL10 in our study but do note that IL10 receptor subunit beta (IL10RB) is differentially spliced in the fibrillogenic mouse DG (vs wild type). In addition, using binding motif-based gene target sets, the zinc finger transcription factor SP1, known as a key mediator of IL10 induction78 was identified as the top transcription factor associated with DEX genes in the fibrillogenic mouse DG (vs wild type) (Figure 3e). In addition to its role in IL10 regulation, SP1 is also a co-activator of APP,79 BACE1 (ref. 80) and MAPT,81 positioning it as an intriguing molecular node at the crossroads of microglial activation, amyloid and tau regulation. Zawia et al.82 have reported on SP1 as a potential therapeutic target in AD, demonstrating that degradation of SP1 in AD mice is associated with improvement of cognitive deficits and Aβ. These data may provide a biological context for the role of SP1 in AD.
Comparison of mouse AD model networks vs human LOAD gene networks
We identified multiple significant overlaps between the DE and DEX signatures of this study, and LOAD-associated gene co-expression modules identified by Zhang et al.,5 including the ‘immune and microglia’ module that contains TYROBP. A recent large study describing gene co-expression of multiple AD mouse strains across three brain regions and multiple time points,83 reported very strong correlation between the expression of an ‘Immune' co-expression module, and amyloid plaque density. This module (‘M1’), which includes TYROBP and TREM2 as highly connected genes, is in line with the findings of Zhang et al.5 and transitively suggests overlap with our findings for genes DE in the fibrillogenic mouse DG.
We also note that in the Bayesian network built from case-only samples by Zhang et al., TYROBP remained the most strongly connected node in the immediate neighborhood of the DE fibrillogenic DG genes, with connections to multiple other DE genes, as well SUZ12 targets and ECM-associated genes.
Additional connections were made to LOAD-associated co-expression modules with the top Gene Ontology terms ‘Neurogenesis’ and ‘Extracellular Matrix’, indicating that these themes, and their molecular correlates in these mice have a meaningful co-expression analog in human LOAD.
Summary
The novel data and analysis presented by this study offer several new conclusions concerning molecular systems perturbations in AD mouse strains. First, the broader corroboration of mouse model molecular pathology by that occurring in human LOAD validates the use of genetically manipulated mice to model the pathogenesis and/or treatment of LOAD. This is especially noteworthy given the recent speculation that current AD drug discovery has failed, at least in part, because of the reliance on mouse models.84 Although neither of these mouse strains represent a true model of LOAD, and therefore would not be expected to inform on the molecular basis of altered Aβ clearance that is seen in LOAD, they do offer a valuable opportunity for detailed study of the damaging consequences of Aβ accumulation, such as mechanisms of oligomer-induced toxicity. We found that the integrated analysis of differential gene and exon expression also highlighted multiple candidate mediators of disease pathophysiology, such as the association of SUZ12 with dysregulated neurogenesis in the dentate gyrus, and FMR1 as a known suppressor of APP translation. When the data herein are taken together with the recent realization that amyloid pathology may precede clinical cognitive impairment by up to 30 years,85 as well as with the recent success of aducanumab in modulating the clinical and pathological features of LOAD,86 the aggregated observations tend to support the amyloid hypothesis. Specifically, the formulation that the prior failure of human clinical trials is primarily attributable to the focus on treatment of symptomatic disease when the better application of amyloid-reducing strategies would be toward prevention of the initial phases of amyloidosis by initiating intervention in middle age.85, 87, 88, 89 Finally, the combination of genetic linkage of TREM2 to AD and the identification of several microglial phagocytic pathways through integrative analyses of both the human LOAD brain19 and the mouse AD model brain (this study) further strengthen the attractiveness of microglia as targets for discovery of drugs that modulate AD pathogenesis at the levels of both disease initiation and disease progression.
Data accessibility
Gene expression and exon expression data are available at the Accelerating Medicines Partnership - Alzheimer's Disease (AMP-AD) Knowledge Portal (https://www.synapse.org/#!Synapse:syn2580853), and can be accessed with synapse ID syn4560807.
References
Schadt EE . Molecular networks as sensors and drivers of common human diseases. Nature 2009; 461: 218–223.
Braak H, Braak E, Yilmazer D, de Vos RA, Jansen EN, Bohl J . Pattern of brain destruction in Parkinson's and Alzheimer's diseases. J Neural Transm 1996; 103: 455–490.
Jonsson T, Stefansson H, Steinberg S, Jonsdottir I, Jonsson PV, Snaedal J et al. Variant of TREM2 associated with the risk of Alzheimer's disease. N Engl J Med 2013; 368: 107–116.
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E et al. TREM2 variants in Alzheimer's disease. N Engl J Med 2013; 368: 117–127.
Zhang B, Gaiteri C, Bodea LG, Wang Z, McElwee J, Podtelezhnikov AA et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer's disease. Cell 2013; 153: 707–720.
Gandy S, Simon AJ, Steele JW, Lublin AL, Lah JJ, Walker LC et al. Days to criterion as an indicator of toxicity associated with human Alzheimer amyloid-beta oligomers. Ann Neurol 2010; 68: 220–230.
Levy E, Carman MD, Fernandez-Madrid IJ, Power MD, Lieberburg I, van Duinen SG et al. Mutation of the Alzheimer's disease amyloid gene in hereditary cerebral hemorrhage, Dutch type. Science 1990; 248: 1124–1126.
Price KA, Varghese M, Sowa A, Yuk F, Brautigam H, Ehrlich ME et al. Altered synaptic structure in the hippocampus in a mouse model of Alzheimer's disease with soluble amyloid-beta oligomers and no plaque pathology. Mol Neurodegener 2014; 9: 41.
Jankowsky JL, Fadale DJ, Anderson J, Xu GM, Gonzales V, Jenkins NA et al. Mutant presenilins specifically elevate the levels of the 42 residue beta-amyloid peptide in vivo: evidence for augmentation of a 42-specific gamma secretase. Hum Mol Genet 2004; 13: 159–170.
Lopez-Gonzalez I, Schluter A, Aso E, Garcia-Esparcia P, Ansoleaga B, LL F et al. Neuroinflammatory signals in Alzheimer disease and APP/PS1 transgenic mice: correlations with plaques, tangles, and oligomeric species. J Neuropathol Exp Neurol 2015; 74: 319–344.
Kim SH, Steele JW, Lee SW, Clemenson GD, Carter TA, Treuner K et al. Proneurogenic Group II mGluR antagonist improves learning and reduces anxiety in Alzheimer Abeta oligomer mouse. Mol Psychiatry 2014; 19: 1235–1242.
Sebollela A, Mustata GM, Luo K, Velasco PT, Viola KL, Cline EN et al. Elucidating molecular mass and shape of a neurotoxic Abeta oligomer. ACS Chem Neurosci 2014; 5: 1238–1245.
Walsh DM, Klyubin I, Fadeeva JV, Rowan MJ, Selkoe DJ . Amyloid-beta oligomers: their production, toxicity and therapeutic inhibition. Biochem Soc Trans 2002; 30: 552–557.
Hart SN, Therneau TM, Zhang Y, Poland GA, Kocher JP . Calculating sample size estimates for RNA sequencing data. J Comput Biol 2013; 20: 970–978.
Mouse Genome Sequencing C Mouse Genome Sequencing C Waterston RH Mouse Genome Sequencing C Lindblad-Toh K Mouse Genome Sequencing C Birney E Mouse Genome Sequencing C Rogers J Mouse Genome Sequencing C Abril JF et al. Initial sequencing and comparative analysis of the mouse genome. Nature 2002; 420: 520–562.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013; 29: 15–21.
Liao Y, Smyth GK, Shi W . The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res 2013; 41: e108.
Liao Y, Smyth GK, Shi W . FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014; 30: 923–930.
Love MI, Huber W, Anders S . Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014; 15: 550.
Benjamini Y, Hochberg Y . Controlling the false discovery rate - a practical and powerful approach to multiple testing. J Roy Stat Soc B Met 1995; 57: 289–300.
Luo W, Friedman MS, Shedden K, Hankenson KD, Woolf PJ . GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 2009; 10: 161.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Miller JA, Cai C, Langfelder P, Geschwind DH, Kurian SM, Salomon DR et al. Strategies for aggregating gene expression data: the collapseRows R function. BMC Bioinformatics 2011; 12: 322.
Chen EY, Xu H, Gordonov S, Lim MP, Perkins MH, Ma'ayan A . Expression2Kinases: mRNA profiling linked to multiple upstream regulatory layers. Bioinformatics 2012; 28: 105–111.
Lachmann A, Xu H, Krishnan J, Berger SI, Mazloom AR, Ma'ayan A . ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 2010; 26: 2438–2444.
Eppig JT, Blake JA, Bult CJ, Kadin JA, Richardson JE, Mouse Genome Database G. The Mouse Genome Database (MGD): facilitating mouse as a model for human biology and disease. Nucleic Acids Res 2015; 43: D726–D736.
Carlson M, Falcon S, Pages H, Li N . org.Hs.eg.db: Genome wide annotation for Human, 2013.
Anders S, Reyes A, Huber W . Detecting differential usage of exons from RNA-seq data. Genome Res 2012; 22: 2008–2017.
Berkeley CA . Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. E-book available at: http://www.bepress.com/sagmb/vol3/iss1/art3, 2004.
Law CW, Chen Y, Shi W, Smyth GK . voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 2014; 15: R29.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015; 43: e47.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25: 25–29.
Kanehisa M, Goto S, Sato Y, Kawashima M, Furumichi M, Tanabe M . Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res 2014; 42: D199–D205.
Kanehisa M, Goto S . KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 2000; 28: 27–30.
Blalock EM, Geddes JW, Chen KC, Porter NM, Markesbery WR, Landfield PW . Incipient Alzheimer's disease: microarray correlation analyses reveal major transcriptional and tumor suppressor responses. Proc Natl Acad Sci USA 2004; 101: 2173–2178.
Colangelo V, Schurr J, Ball MJ, Pelaez RP, Bazan NG, Lukiw WJ . Gene expression profiling of 12633 genes in Alzheimer hippocampal CA1: transcription and neurotrophic factor down-regulation and up-regulation of apoptotic and pro-inflammatory signaling. J Neurosci Res 2002; 70: 462–473.
Liang WS, Dunckley T, Beach TG, Grover A, Mastroeni D, Ramsey K et al. Altered neuronal gene expression in brain regions differentially affected by Alzheimer's disease: a reference data set. Physiol Genomics 2008; 33: 240–256.
Bamberger ME, Harris ME, McDonald DR, Husemann J, Landreth GE . A cell surface receptor complex for fibrillar beta-amyloid mediates microglial activation. J Neurosci 2003; 23: 2665–2674.
Coraci IS, Husemann J, Berman JW, Hulette C, Dufour JH, Campanella GK et al. CD36, a class B scavenger receptor, is expressed on microglia in Alzheimer's disease brains and can mediate production of reactive oxygen species in response to beta-amyloid fibrils. Am J Pathol 2002; 160: 101–112.
Koenigsknecht J, Landreth G . Microglial phagocytosis of fibrillar beta-amyloid through a beta1 integrin-dependent mechanism. J Neurosci 2004; 24: 9838–9846.
Chouraki V, De Bruijn RF, Chapuis J, Bis JC, Reitz C, Schraen S et al. A genome-wide association meta-analysis of plasma Abeta peptides concentrations in the elderly. Mol Psychiatry 2014; 19: 1326–1335.
Gartner U, Holzer M, Arendt T . Elevated expression of p21ras is an early event in Alzheimer's disease and precedes neurofibrillary degeneration. Neuroscience 1999; 91: 1–5.
Poduslo SE, Huang R, Huang J, Smith S . Genome screen of late-onset Alzheimer's extended pedigrees identifies TRPC4AP by haplotype analysis. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 50–55.
Grupe A, Li Y, Rowland C, Nowotny P, Hinrichs AL, Smemo S et al. A scan of chromosome 10 identifies a novel locus showing strong association with late-onset Alzheimer disease. Am J Hum Genet 2006; 78: 78–88.
Westmark CJ, Malter JS . FMRP mediates mGluR5-dependent translation of amyloid precursor protein. PLoS Biol 2007; 5: e52.
Nizzari M, Venezia V, Repetto E, Caorsi V, Magrassi R, Gagliani MC et al. Amyloid precursor protein and Presenilin1 interact with the adaptor GRB2 and modulate ERK 1,2 signaling. J Biol Chem 2007; 282: 13833–13844.
Chong YH, Shin YJ, Lee EO, Kayed R, Glabe CG, Tenner AJ . ERK1/2 activation mediates Abeta oligomer-induced neurotoxicity via caspase-3 activation and tau cleavage in rat organotypic hippocampal slice cultures. J Biol Chem 2006; 281: 20315–20325.
Ghersi E, Noviello C, D'Adamio L . Amyloid-beta protein precursor (AbetaPP) intracellular domain-associated protein-1 proteins bind to AbetaPP and modulate its processing in an isoform-specific manner. J Biol Chem 2004; 279: 49105–49112.
Yun SM, Cho SJ, Song JC, Song SY, Jo SA, Jo C et al. SUMO1 modulates Abeta generation via BACE1 accumulation. Neurobiol Aging 2013; 34: 650–662.
Pasini D, Bracken AP, Hansen JB, Capillo M, Helin K . The polycomb group protein Suz12 is required for embryonic stem cell differentiation. Mol Cell Biol 2007; 27: 3769–3779.
Bradley WD, Koleske AJ . Regulation of cell migration and morphogenesis by Abl-family kinases: emerging mechanisms and physiological contexts. J Cell Sci 2009; 122: 3441–3454.
Sarrazin S, Lamanna WC, Esko JD . Heparan sulfate proteoglycans. Cold Spring Harb Perspect Biol 2011; 3: pii: a004952.
Synapse. Accelerating Medicines Partnership (AMP) Alzheimer’s Disease Knowledge Portal. https://www.synapse.org/#!Synapse:syn2580853/wiki/ doi:10.7303/syn2580853, 2015.
Lansbergen G, Akhmanova A . Microtubule plus end: a hub of cellular activities. Traffic 2006; 7: 499–507.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13: 2498–2504.
Morris JH, Kuchinsky A, Ferrin TE, Pico AR . enhancedGraphics: a Cytoscape app for enhanced node graphics. F1000Res 2014; 3: 147.
Westmark CJ, Westmark PR, O'Riordan KJ, Ray BC, Hervey CM, Salamat MS et al. Reversal of fragile X phenotypes by manipulation of AbetaPP/Abeta levels in Fmr1KO mice. PloS One 2011; 6: e26549.
Renoux AJ, Carducci NM, Ahmady AA, Todd PK . Fragile X mental retardation protein expression in Alzheimer's disease. Front Genet 2014; 5: 360.
Orlov SV, Kuteykin-Teplyakov KB, Ignatovich IA, Dizhe EB, Mirgorodskaya OA, Grishin AV et al. Novel repressor of the human FMR1 gene - identification of p56 human (GCC)(n)-binding protein as a Kruppel-like transcription factor ZF5. FEBS J 2007; 274: 4848–4862.
Lane RF, Raines SM, Steele JW, Ehrlich ME, Lah JA, Small SA et al. Diabetes-associated SorCS1 regulates Alzheimer's amyloid-beta metabolism: evidence for involvement of SorL1 and the retromer complex. J Neurosci 2010; 30: 13110–13115.
Ko CY, Hsu HC, Shen MR, Chang WC, Wang JM . Epigenetic silencing of CCAAT/enhancer-binding protein delta activity by YY1/polycomb group/DNA methyltransferase complex. J Biol Chem 2008; 283: 30919–30932.
Gjoneska E, Pfenning AR, Mathys H, Quon G, Kundaje A, Tsai LH et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer's disease. Nature 2015; 518: 365–369.
Yu L, Chibnik LB, Srivastava GP, Pochet N, Yang J, Xu J et al. Association of Brain DNA methylation in SORL1, ABCA7, HLA-DRB5, SLC24A4, and BIN1 with pathological diagnosis of Alzheimer disease. JAMA Neurol 2015; 72: 15–24.
Lunnon K, Smith R, Hannon E, De Jager PL, Srivastava G, Volta M et al. Methylomic profiling implicates cortical deregulation of ANK1 in Alzheimer's disease. Nat Neurosci 2014; 17: 1164–1170.
Lendvai D, Morawski M, Negyessy L, Gati G, Jager C, Baksa G et al. Neurochemical mapping of the human hippocampus reveals perisynaptic matrix around functional synapses in Alzheimer's disease. Acta Neuropathol 2013; 125: 215–229.
Vegh MJ, Heldring CM, Kamphuis W, Hijazi S, Timmerman AJ, Li KW et al. Reducing hippocampal extracellular matrix reverses early memory deficits in a mouse model of Alzheimer's disease. Acta Neuropathol Commun 2014; 2: 76.
Jorissen E, Prox J, Bernreuther C, Weber S, Schwanbeck R, Serneels L et al. The disintegrin/metalloproteinase ADAM10 is essential for the establishment of the brain cortex. J Neurosci 2010; 30: 4833–4844.
Postina R, Schroeder A, Dewachter I, Bohl J, Schmitt U, Kojro E et al. A disintegrin-metalloproteinase prevents amyloid plaque formation and hippocampal defects in an Alzheimer disease mouse model. J Clin Invest 2004; 113: 1456–1464.
Suh J, Choi SH, Romano DM, Gannon MA, Lesinski AN, Kim DY et al. ADAM10 missense mutations potentiate beta-amyloid accumulation by impairing prodomain chaperone function. Neuron 2013; 80: 385–401.
Tremblay MA, Acker CM, Davies P . Tau phosphorylated at tyrosine 394 is found in Alzheimer's disease tangles and can be a product of the Abl-related kinase, Arg. J Alzheimer's Dis 2010; 19: 721–733.
Gandy S, Heppner FL . Microglia as dynamic and essential components of the amyloid hypothesis. Neuron 2013; 78: 575–577.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat Genet 2013; 45: 1452–1458.
Griciuc A, Serrano-Pozo A, Parrado AR, Lesinski AN, Asselin CN, Mullin K et al. Alzheimer's disease risk gene CD33 inhibits microglial uptake of amyloid beta. Neuron 2013; 78: 631–643.
Tan J, Town T, Mori T, Wu Y, Saxe M, Crawford F et al. CD45 opposes beta-amyloid peptide-induced microglial activation via inhibition of p44/42 mitogen-activated protein kinase. J Neurosci 2000; 20: 7587–7594.
Ebner F, Brandt C, Thiele P, Richter D, Schliesser U, Siffrin V et al. Microglial activation milieu controls regulatory T cell responses. J Immunol 2013; 191: 5594–5602.
Guillot-Sestier MV, Doty KR, Gate D, Rodriguez J Jr, Leung BP, Rezai-Zadeh K et al. Il10 deficiency rebalances innate immunity to mitigate Alzheimer-like pathology. Neuron 2015; 85: 534–548.
Chakrabarty P, Li A, Ceballos-Diaz C, Eddy JA, Funk CC, Moore B et al. IL-10 alters immunoproteostasis in APP mice, increasing plaque burden and worsening cognitive behavior. Neuron 2015; 85: 519–533.
Tone M, Powell MJ, Tone Y, Thompson SA, Waldmann H . IL-10 gene expression is controlled by the transcription factors Sp1 and Sp3. J Immunol 2000; 165: 286–291.
Docagne F, Gabriel C, Lebeurrier N, Lesne S, Hommet Y, Plawinski L et al. Sp1 and Smad transcription factors co-operate to mediate TGF-beta-dependent activation of amyloid-beta precursor protein gene transcription. Biochem J 2004; 383: 393–399.
Christensen MA, Zhou W, Qing H, Lehman A, Philipsen S, Song W . Transcriptional regulation of BACE1, the beta-amyloid precursor protein beta-secretase, by Sp1. Mol Cell Biol 2004; 24: 865–874.
Heicklen-Klein A, Ginzburg I . Tau promoter confers neuronal specificity and binds Sp1 and AP-2. J Neurochem 2000; 75: 1408–1418.
Subaiea GM, Adwan LI, Ahmed AH, Stevens KE, Zawia NH . Short-term treatment with tolfenamic acid improves cognitive functions in Alzheimer's disease mice. Neurobiol Aging 2013; 34: 2421–2430.
Matarin M, Salih DA, Yasvoina M, Cummings DM, Guelfi S, Liu W et al. A genome-wide gene-expression analysis and database in transgenic mice during development of amyloid or tau pathology. Cell Rep 2015; 10: 633–644.
Burns TC, Li MD, Mehta S, Awad AJ, Morgan AA . Mouse models rarely mimic the transcriptome of human neurodegenerative diseases: A systematic bioinformatics-based critique of preclinical models. Eur J Pharmacol 2015; 759: 101–117.
Jansen WJ, Ossenkoppele R, Knol DL, Tijms BM, Scheltens P, Verhey FR et al. Prevalence of cerebral amyloid pathology in persons without dementia: a meta-analysis. JAMA 2015; 313: 1924–1938.
Sevigny J, Chiao P, Williams L, Chen T, Ling Y, O’Gorman J et al. Randomized, Double-blind, Placebo-controlled, Phase 1b Study of Aducanumab (BIIB037), an Anti-Aβ Monoclonal Antibody, in Patients With Prodromal or Mild Alzheimer’s Disease: Interim Results by Disease Stage and ApoE ɛ4 Status. 67th Annual Meeting of the American Academy of Neurology 18-25 April 2015 Washington DC.
Gandy S . Lifelong management of amyloid-beta metabolism to prevent Alzheimer's disease. N Engl J Med 2012; 367: 864–866.
Gandy S . Perspective: prevention is better than cure. Nature 2011; 475: S15.
Gandy S, DeKosky ST . Toward the treatment and prevention of Alzheimer's disease: rational strategies and recent progress. Annu Rev Med 2013; 64: 367–383.
Acknowledgements
We thank the Mount Sinai Genomics Core Facility for carrying out the RNA sequencing. We would like to recognize the federal and foundation support that made this research possible. NIH support for this work included multi-PI U01 AG046170 (to EES, SG, VH, MEE), multi-PI R34 AG049649 (JTD, SG), R01 NS075685 (SG), P50 AG005138 to Mary Sano (SG), R01 DK098242 to JTD, U54 CA189201 to JTD, and HHSN271201300031C to VH. VA MERIT support included RRD MERIT I01RX000684 (SG) and BLRD MERIT I01BX000348 (SG). The Cure Alzheimer's Fund Research Consortium also supported this work (SG). The computational resources and staff expertise provided by the Department of Scientific Computing at the Icahn School of Medicine at Mount Sinai also contributed to the performance of this research. Data for the human LOAD DEX analysis were generated from postmortem brain tissue collected through the Mount Sinai VA Medical Center Brain Bank (generously provided by Dr. Eric Schadt), and the Mount Sinai NIH Brain and Tissue Repository (generously provided by Dr. Vahram Haroutunian). The results published here are based on data obtained from the Accelerating Medicines Partnership for Alzheimer's Disease (AMP-AD) Target Discovery Consortium data portal and can be accessed at doi:10.7303/syn2580853.
Author contributions
MEE, JVHM, SG, BR and JTD designed the study; MEE and JVHM carried out the dissections; MEE and JVHM prepared and extracted the RNA; BR performed the computational analysis; MEE, JVHM, SG, BR and JTD wrote the paper; EES, BZ and VH revised the analysis critically for important intellectual content. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Readhead, B., Haure-Mirande, JV., Zhang, B. et al. Molecular systems evaluation of oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse models identifies shared features with human Alzheimer’s brain molecular pathology. Mol Psychiatry 21, 1099–1111 (2016). https://doi.org/10.1038/mp.2015.167
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.167
This article is cited by
-
Integrative approach to sporadic Alzheimer’s disease: deficiency of TYROBP in a tauopathy mouse model reduces C1q and normalizes clinical phenotype while increasing spread and state of phosphorylation of tau
Molecular Psychiatry (2019)
-
Integrative approach to sporadic Alzheimer’s disease: deficiency of TYROBP in cerebral Aβ amyloidosis mouse normalizes clinical phenotype and complement subnetwork molecular pathology without reducing Aβ burden
Molecular Psychiatry (2019)
-
PRDX6 Inhibits Neurogenesis through Downregulation of WDFY1-Mediated TLR4 Signal
Molecular Neurobiology (2019)
-
Fragile X and APP: a Decade in Review, a Vision for the Future
Molecular Neurobiology (2019)
-
Deficiency of TYROBP, an adapter protein for TREM2 and CR3 receptors, is neuroprotective in a mouse model of early Alzheimer’s pathology
Acta Neuropathologica (2017)