Abstract
Leiomyomas associated with hereditary leiomyomatosis and renal cell carcinoma syndrome and leiomyomas with bizarre nuclei often show overlapping morphological features, in particular cells with prominent eosinophilic nucleoli, perinucleolar halos, and eosinophilic cytoplasmic inclusions. Although hereditary leiomyomatosis and renal cell carcinoma syndrome is defined by fumarate hydratase (FH) germline mutations, resulting in S-(2-succino)-cysteine (2SC) formation, it is unknown whether leiomyomas with bizarre nuclei show similar alterations. In this study, we evaluated the morphology and FH/2SC immunoprofile of 31 leiomyomas with bizarre nuclei. DNA from tumor and normal tissues from 24 cases was subjected to massively parallel sequencing targeting 410 key cancer genes. Somatic genetic alterations were detected using state-of-the-art bioinformatics algorithms. No patient reported a personal history of renal neoplasia or cutaneous leiomyomas, but one had a family history of renal cell carcinoma while another had a family history of uterine leiomyomas. Aberrant FH/2SC expression was noted in 17 tumors (16 FH-negative/2SC-positive, 1 FH-positive/2SC-positive). On univariate analysis, staghorn vessels, eosinophilic cytoplasmic inclusions, diffuse distribution of prominent eosinophilic nucleoli with perinucleolar halos, and an ‘alveolar pattern of edema’ were associated with an abnormal immunoprofile, but only staghorn vessels remained significant on multivariate analysis. Massively parallel sequencing analysis (n=24) revealed that 13/14 tumors with aberrant FH/2SC immunoprofile harbored somatic FH somatic genetic alterations, including homozygous deletions (n=9), missense mutations coupled with loss of heterozygosity (n=3), and a splice site mutation (n=1), whereas no somatic FH mutations/deletions were found in tumors with normal immunoprofile (n=10; P<0.0001). Leiomyomas with bizarre nuclei with normal FH/2SC staining pattern more frequently harbored TP53 and/or RB1 alterations than those with aberrant FH/2SC immunoprofile (60 vs 14%; P=0.032). These data demonstrate that leiomyomas with bizarre nuclei are morphologically and genetically heterogeneous and that hereditary leiomyomatosis and renal cell carcinoma syndrome-related morphological features, abnormal FH/2SC staining, and somatic FH mutations/deletions can be seen in a subset of sporadic tumors.
Similar content being viewed by others
Main
In hereditary leiomyomatosis and renal cell carcinoma syndrome, germline mutations in the fumarate hydratase (FH) gene predispose individuals to renal cell carcinoma as well as to uterine and cutaneous leiomyomas.1 FH plays a key role in the tricarboxylic acid cycle by catalyzing the conversion of fumarate to malate. In FH-mutant cells, however, intracellular levels of fumarate accumulate and can react with cysteine residues on various proteins to form S-(2-succino)-cysteine (2SC).2, 3 This process, termed protein succination, has been identified in several key proteins involved in metabolic and regulatory functions,4, 5 and recently an antibody has been developed to detect this biochemical modification.3
Several morphological features have been reported to be distinctive of uterine leiomyomas from patients with hereditary leiomyomatosis and renal cell carcinoma syndrome, including increased cellularity, staghorn vascularity, eosinophilic cytoplasmic inclusions, and large prominent eosinophilic nucleoli with perinucleolar halos.6, 7, 8 On immunohistochemistry, these hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas demonstrate strong and diffuse 2SC expression with absence of FH staining.8 Although leiomyomas with bizarre nuclei may show overlapping morphological features,9 it is unknown whether these tumors display similar immunohistochemical profiles and if they harbor alterations affecting the FH gene. Herein, we reviewed a series of leiomyomas with bizarre nuclei and aimed to evaluate morphological features that have been reported in hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas, study their FH and 2SC expression patterns, correlate the morphological and immunohistochemical findings, and analyze their repertoire of somatic genetic alterations, including FH gene alterations, using targeted massively parallel sequencing.
Materials and methods
Case Selection
The surgical pathology archives at the Massachusetts General Hospital, as well as the consultation files of two of the authors (EO and RHY), were searched from 1990 to 2014, and 31 leiomyomas with bizarre nuclei were identified. Clinical information, including age, personal or family history of renal or cutaneous tumors, and follow-up, were obtained from the electronic medical records and/or consulting pathologist. This study was approved by the Institutional Review Board of the Massachusetts General Hospital.
Morphological Analysis
The number of hematoxylin and eosin-stained tumor slides ranged from 1 to 18 (mean 7). The tumors were reviewed by two gynecological pathologists (JAB, EO) for morphological features previously reported in hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas, including cellularity compared to the adjacent myometrium (hypercellular, normocellular/hypocellular), staghorn vessels (present, absent), eosinophilic cytoplasmic inclusions (present, absent), and prominent eosinophilic nucleoli with perinucleolar halos (diffusely distributed, focally distributed, or absent).6, 7, 8 In addition, we evaluated the presence/absence of a striking pattern of edema, designated as ‘alveolar pattern of edema’ (due to its histological resemblance to pulmonary alveoli), that we have noted in a subset of leiomyomas with bizarre nuclei,10 as well as recorded the distribution of the bizarre nuclei (diffuse, focal).
Immunohistochemistry
Immunohistochemical analysis for FH (1:1000, Clone J13, Santa Cruz Technology) and 2SC (1:5000, not commercially available)11 was performed on 4-μm-thick sections from representative formalin-fixed, paraffin-embedded tissue blocks using the Ventana Discovery XT system as previously described.11, 12 Positive and negative controls were included in each slide run. The immunohistochemical results were reviewed by two pathologists (Y-BC, SC).12 FH staining was scored qualitatively as negative or positive when compared with internal positive controls (endothelial/stromal cells). 2SC staining was assessed for intensity (1+ to 3+) and staining pattern (nuclear and cytoplasmic versus cytoplasmic only), although only 3+ intensity nucleocytoplasmic staining was interpreted as positive, also as reported previously.11 Renal tumors from genetically confirmed hereditary leiomyomatosis and renal cell carcinoma syndrome patients were used as positive control for the 2SC immunohistochemical analysis.
Microdissection and Nucleic Acid Extraction
Eight-μm-thick tumor and matched normal formalin-fixed, paraffin-embedded tissue sections were stained with nuclear fast red and microdissected by two pathologists (RB, AMS) to ensure >80% of tumor cell content and that the normal tissue was devoid of any neoplastic cells as previously described.13 Genomic DNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen) and quantified using the Qubit Fluorometer (Life Technologies), as previously described.14 We obtained tumor and matched normal DNA of sufficient quantity and quality from 24 leiomyomas with bizarre nuclei.
Targeted Capture Massively Parallel Sequencing
Tumor and normal DNA samples from 24 cases were subjected to targeted massively parallel sequencing at the Memorial Sloan Kettering Cancer Center Integrated Genomics Operation using the Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT) assay targeting all exons and selected introns of 410 key cancer genes, as previously described.14, 15 Sequence reads generated on the Illumina HiSeq2000 were aligned to the human reference genome GRCh37 using the Burrows–Wheeler Aligner (v0.7.10),16 and local realignment, duplicate removal, and base quality recalibration were performed using the Genome Analysis Toolkit (GATK, v3.1.1).17
Variant calling was performed as described previously.14 Somatic single-nucleotide variants (SNVs) were identified using MuTect (v1.0),18 and small insertions and deletions (indels) using Strelka (v2.0.15)19 and VarScan 2 (v2.3.7).20 SNVs and indels with mutant allelic fraction (MAF) of <1% and/or those supported by ≤5 reads, those for which the tumor MAF was <5 times that of the matched normal MAF, as well as SNVs and indels found at >5% global minor allele frequency of dbSNP (build 137), were filtered out as previously described.21 Copy number alterations and loss of heterozygosity were defined using FACETS as previously described.22, 23 All areas of homozygous deletions and loss of heterozygosity were visually inspected using raw Log2 and allelic copy ratio plots. Cancer cell fractions of all mutations were inferred using ABSOLUTE (v1.0.6),24 as previously described.14, 23
To validate the presence/absence of somatic FH mutations identified using MSK-IMPACT, all cases were subjected to massively parallel sequencing using a custom bait set (Integrated DNA Technologies, IDT) targeting all exons and flanking intronic regions of the FH gene. Captured DNA libraries were sequenced on an Illumina HiSeq2000, and sequencing data were analyzed as described above.
The potential functional effect of each somatic SNV was investigated using a combination of MutationTaster, CHASM (uterus), and FATHMM25, 26, 27, 28 and of each somatic indel using MutationTaster and PROVEAN.29 In addition, we assessed whether the variants affected cancer genes as defined by Kandoth et al,30 the Cancer Gene Census,31 or Lawrence et al,32 as previously described.14 Hotspot SNVs were annotated according to Chang et al.33
Statistical Analysis
Statistical analysis was performed using Stata (Version 14.0, StataCorp, College Station, TX, USA). Univariate analysis was conducted using Fisher’s exact test. Multivariable-adjusted analysis was performed using exact logistic regression to identify independent predictors of aberrant FH/2SC expression. Statistically significant variables in univariate analysis were considered for inclusion in the multivariable model. Odds ratios and 95% confidence intervals were obtained. P-values<0.05 were considered statistically significant.
Results
Clinical Features
Patient age ranged from 26 to 77 (n=30, mean 46.7, median 44.5; n=1, unknown) years. Personal and family history was available for 27 and 24 patients, respectively. None of the patients included in this study had a personal history of renal neoplasia or cutaneous leiomyomas; however, one patient had a urethral leiomyoma (SCU29T) while another had multiple colonic leiomyomas (SCU4T). Patient SCU29T with the urethral leiomyoma also had a family history of renal cell carcinoma, whereas another patient (SCU19T) noted a family history of uterine leiomyomas. None of the patients reported a family history of cutaneous leiomyomas.
The majority of patients (25/31; 81%) included in this study presented with a known history of uterine leiomyomas and/or associated symptoms (typically menorrhagia or pelvic pain). In four others, indication for surgery included uterine prolapse, squamous cell carcinoma in situ of the cervix, endometrial carcinoma, and ovarian cysts, while clinical presentation was unknown in the remaining two patients. Hysterectomy and myomectomy were performed in 23 (74%) and 7 (23%), respectively; the surgical procedure was not known in 1 (3%) patient. Follow-up in 26 (84%) patients showed all were alive without any evidence of hereditary leiomyomatosis and renal cell carcinoma syndrome-associated disease (mean 7.4 years; range 2.5 months to 22 years).
Gross Features
Multiple leiomyomas were found in the uterus of 19 (61%) patients and a single leiomyoma in 8 (26%) (Table 1). In 3 (10%) patients, the number of leiomyomas could not be determined due to specimen fragmentation, and in the remaining 1 (3%), a gross description was unavailable. In all but 1 patient, the leiomyoma with bizarre nuclei was the largest leiomyoma, ranging from 1.4 to 18 cm (mean and median 9 cm; unavailable in 1) in maximum dimension, and was well demarcated from the surrounding myometrium. Most tumors (74%) displayed a tan-pink to white and firm, rubbery, whorled cut surface; however, 7 were soft and 2 yellow. Focal hemorrhage was noted in 10 (32%), cystification/cavitation in 4 (13%), necrosis in 2 (6%), and edema or calcification in 1 (3%) each. One tumor (SCU20T) was primarily yellow with focal necrosis and hemorrhage but also contained a discrete, well-circumscribed, 5.5 cm gray-white nodule.
Morphological and Immunohistochemical Features
At low magnification, 87% (27/31) of tumors were hypercellular compared to the adjacent myometrium (Figure 1a). Staghorn vessels were noted in 71% (22/31) (Figure 1b) while an ‘alveolar pattern of edema’ was observed in 42% (13/31) (Figure 1c) of tumors. Bizarre nuclei, characterized by enlarged, hyperchromatic, frequently multinucleated cells, showed a diffuse (Figure 1d) and focal distribution in 61% (19/31) and 39% (12/31) of the leiomyomas with bizarre nuclei studied, respectively. On higher magnification, eosinophilic cytoplasmic globules imparting a rhabdoid appearance (Figure 1e) were noted in 84% (26/31). Prominent eosinophilic nucleoli with perinucleolar halos (Figure 1f) were present with diffuse and focal distribution in 71% (22/31) and 19% (6/31) tumors, respectively, and were absent in 10% (3/31) (Table 1). Of note, these latter two features were primarily observed in mononuclear cells.
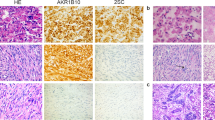
Morphological features of leiomyomas with bizarre nuclei. They were often hypercellular compared to the adjacent myometrium (a) and many had staghorn vessels (b). An ‘alveolar pattern of edema’ (c) was noted in 42% and a diffuse distribution of bizarre nuclei (d) in 61% of tumors. Most leiomyomas with bizarre nuclei, especially those with an aberrant FH/2SC immunoprofile, had eosinophilic cytoplasmic inclusions (e) and a diffuse distribution of cells with prominent eosinophilic nucleoli and perinucleolar halos (f, arrows).
Other observed features included prominent hydropic change in two tumors (SCU2T and SCU19T), in one of them being quite pronounced and focally resembled a dissecting cotyledonoid leiomyoma (SCU19T). A neurilemmoma-like pattern was noted in two leiomyomas with bizarre nuclei (SCU19T and SCU26T). Tumor SCU20T that was described as having a discrete nodule within the main lesion was diagnosed as a leiomyosarcoma arising in a leiomyoma with bizarre nuclei. The sarcoma made up approximately 40% of the sampled tumor and showed an abrupt transition from the leiomyoma. The former was characterized by pleomorphic spindle cells with brisk mitotic activity and tumor cell necrosis, and in contrast to the leiomyoma with bizarre nuclei, lacked eosinophilic cytoplasmic inclusions and perinucleolar halos. Brisk mitotic activity and tumor necrosis were not present in the adjacent leiomyoma with bizarre nuclei.
A normal pattern of staining, characterized by diffuse cytoplasmic positivity for FH and absence of 2SC staining, was observed in 45% (14/31) of the leiomyomas with bizarre nuclei studied (Figure 2a). Sixteen of the 17 remaining tumors did not express FH and were positive for 2SC (Figure 2b), whereas one tumor expressed both FH and 2SC (SCU14T) (Figure 2c; Table 1). There was strong and diffuse FH expression and absence of 2SC staining in the case demonstrating a leiomyosarcoma arising within a leiomyoma with bizarre nuclei (SCU20T). The FH and 2SC staining patterns in the malignant and benign areas were identical.
FH/2SC immunoprofiles observed in leiomyoma with bizarre nuclei. (a) Representative micrograph of a leiomyoma with bizarre nuclei (hematoxylin and eosin (H&E), top) showing a normal FH/2SC expression pattern with diffuse cytoplasmic positivity for FH (middle) and absence of 2SC staining (bottom). (b) Leiomyoma with bizarre nuclei (H&E, top) displaying an aberrant immunoprofile. Tumor cells lack FH expression (middle) and are 2SC-positive (bottom). (c) Leiomyoma with bizarre nuclei (SCU14T) (H&E, top) with an aberrant FH/2SC staining pattern. Tumor cells express both FH (middle) and 2SC (bottom). FH, fumarate hydratase; 2SC, S-(2-succino)-cysteine.
On univariate analysis, the presence of ‘alveolar pattern of edema’ (P=0.001), staghorn vessels (P=0.004), diffuse distribution of eosinophilic nucleoli with perinucleolar halos (P=0.004), and eosinophilic cytoplasmic inclusions (P=0.012) were all shown to be associated with an aberrant immunophenotype. Increased cellularity compared to the adjacent myometrium and a diffuse distribution of bizarre nuclei failed to reach statistical significance (P=1.000 and P=0.724, respectively; Table 2). In a multivariable-adjusted model, the presence of staghorn vessels was the only significant independent predictor of abnormal FH/2SC staining (P=0.031; Supplementary Table S1).
Repertoire of Somatic Genetic Alterations
For 24 leiomyomas with bizarre nuclei, targeted massively parallel sequencing of 410 key cancer genes was performed, with a median depth of 543 × (range 107 × –828 ×) and 401 × (78 × –958 ×) for tumor and normal samples, respectively (Supplementary Table S2). Leiomyomas with bizarre nuclei harbored a median of two non-synonymous somatic mutations (range 0–4) in the 410 genes tested, with the genes most frequently targeted by somatic mutations including FH, TP53, MED12, FOXA1 and KMT2A (Figure 3, Supplementary Table S3). Furthermore, in these tumors a median of 1.5 genes (range 0–13) of the 410 genes assayed were targeted by homozygous deletions, including recurrent homozygous deletions affecting FH, AKT3, RB1, HIST3H3, PARP1, H3F3A, DIS3, FOXO1 and NOTCH4 (Figure 3).
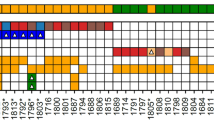
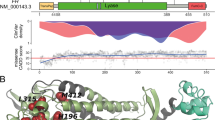
Somatic mutations and gene copy number alterations detected by targeted massively parallel sequencing in leiomyomas with bizarre nuclei. (a) Heatmap showing the non-synonymous somatic mutations and gene copy number alterations identified in two or more leiomyomas with bizarre nuclei. Mutation types and copy number alterations are color-coded according to the legend. The presence of loss of heterozygosity of the wild-type allele of a mutated gene is represented by a diagonal bar. Note that the combined analysis of loss of heterozygosity and copy number revealed that in cases SCU10T, SCU11T and SCU14T the loss of heterozygosity was caused by a physical deletion of the wild-type copy of FH. IHC, immunohistochemistry. (b) Representative genome plots of a leiomyoma with bizarre nuclei with aberrant FH/2SC immunoprofile and FH homozygous deletion (blue arrow) (top) and a leiomyoma with bizarre nuclei with normal FH/2SC immunoprofile and RB1 homozygous deletion (blue arrow) (bottom). The Log2 copy number ratios are plotted on the y axis according to the genomic positions on the x axis. Red denotes regions of copy number loss/deletion, green denotes regions of copy number gain. (c) Representative chromosome 1 plots of leiomyomas with bizarre nuclei with (top) and without (bottom) FH homozygous deletions (blue arrows) (top), where the Log2 copy number ratios are plotted on the y axis according to the segment index on the x axis. Segments of 250 base pairs of at least 35 × coverage obtained from the targeted massively parallel sequencing assay are shown. Red denotes regions of copy number loss/deletion, green denotes regions of copy number gain.
Leiomyomas with bizarre nuclei displaying an aberrant FH/2SC immunoprofile differed from those showing normal patterns of FH/2SC expression in that somatic mutations and homozygous deletions affecting the FH gene were significantly more frequent in the former (Figure 3). Of the 14 tumors with an aberrant FH/2SC immunoprofile, 9 harbored FH gene homozygous deletions (Figures 3a–c), and 3 (21%) harbored clonal (ie, present in virtually all tumor cells) likely pathogenic missense FH mutation coupled with loss of the wild-type allele (ie, loss of heterozygosity, Figure 3a, Supplementary Table S3). All three somatic FH mutations were validated using a custom FH massively parallel sequencing assay (see Methods section). In addition, 1 (7%) leiomyoma with bizarre nuclei harbored a subclonal FH splice site mutation of unknown pathogenic effect, identified using the custom FH sequencing assay (Figure 3a, Supplementary Table S3). No FH mutations were identified in the leiomyomas with bizarre nuclei with normal FH/2SC expression pattern using either sequencing assay. Taken together, 13/14 (93%) tumors with an aberrant FH/2SC immunoprofile harbored FH genetic alterations, whereas no FH mutations/homozygous deletions were found in tumors with normal FH/2SC immunoprofile (n=10; P<0.0001). We observed, however, that leiomyomas with bizarre nuclei with normal FH/2SC staining pattern more frequently harbored TP53 and/or RB1 alterations than those with an aberrant FH/2SC immunoprofile (60 vs 14%; P=0.032; Figure 3a). Hotspot MED12 mutations were rare in this series of leiomyomas with bizarre nuclei; 1 of the 14 (7%) tumors with aberrant FH/2SC immunoprofile and 1 of the 10 (10%) FH/2SC immunoprofile normal tumors harbored MED12 exon 2 G44D/V hotspot mutations. FOXA1 hotspot mutations were found in 2 of the 14 (14%) FH/2SC aberrant leiomyomas with bizarre nuclei but not in those with normal FH/2SC immunoprofile, whereas FOXO1 and NOTCH4 homozygous deletions were restricted to leiomyomas with bizarre nuclei with normal FH/2SC immunoprofile (20% each; Figure 3a).
Germline FH Mutations
The patient with a history of a urethral leiomyoma and a family history of renal cell carcinoma (SCU29) underwent FH germline testing, which did not reveal any FH germline mutations. Owing to institutional review board restrictions, germline mutations could not be evaluated in the other tumors.
Discussion
Herein we show that leiomyomas with bizarre nuclei are morphologically and genetically heterogeneous and that hereditary leiomyomatosis and renal cell carcinoma syndrome-associated morphological features, aberrant FH/2SC staining, and somatic FH gene mutations/homozygous deletions can be seen in a subset of sporadic tumors. In addition, we demonstrated that leiomyomas with bizarre nuclei with aberrant FH/2SC immunoprofile are distinct at the genetic level from those with a normal FH/2SC staining pattern.
When the morphology of renal cell carcinomas from patients with hereditary leiomyomatosis and renal cell carcinoma syndrome was first described, presence of a large nucleus with a very prominent orangiophilic/eosinophilic nucleolus surrounded by a clear halo was coined as the hallmark feature of these tumors.34 Several subsequent studies reported this finding in hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas and also expanded the morphological spectrum to include hypercellularity, staghorn vessels, and eosinophilic cytoplasmic inclusions.6, 7, 8 In conjunction with the appropriate clinical history, these specific histological features have been hypothesized to be a valuable screening tool to identify potential hereditary leiomyomatosis and renal cell carcinoma syndrome patients. Our findings have shown, however, that these morphological features are not specific to hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas but can also be seen in sporadic leiomyomas with bizarre nuclei.
Other studies have identified that these features can also be seen in sporadic leiomyomas.35, 36, 37, 38, 39 In a morphological-based study by Alsolami et al,35 6 pathologists evaluated a series of 30 uterine leiomyomas (19 from patients with germline-proven hereditary leiomyomatosis and renal cell carcinoma syndrome and 11 sporadic) for the above described features; however, interobserver variability was poor for most parameters. Another study observed at least one of these morphological findings in 92% (11/12) of uterine leiomyomas from non-hereditary leiomyomatosis and renal cell carcinoma syndrome patients, but for their 2 leiomyomas from patients with germline-proven mutations, only 1 showed all of the ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’.37 Likewise, although Harrison et al36 identified that hemangiopericytomatous vessels were a constant finding in their FH-deficient tumors, they noted that the other features were variably present. Furthermore, in their tumors from five patients with germline-proven or presumed hereditary leiomyomatosis and renal cell carcinoma syndrome, one leiomyoma was completely banal, whereas the remaining ones showed variability in morphological ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’. Herein we found staghorn vessels, diffuse distribution of eosinophilic nucleoli with prominent perinucleolar halos, eosinophilic cytoplasmic inclusions, and an alveolar pattern of edema to be predictive of an abnormal immunophenotype on univariate analysis, but only staghorn vessels remained significant on multivariate analysis. Such vessels were seen in all of our leiomyomas with bizarre nuclei with abnormal staining, a feature first described by Reyes et al8 and further confirmed by Harrison et al.36 Similarly, all tumors with an aberrant immunophenotype also had a diffuse distribution of prominent eosinophilic nucleoli with perinucleolar halos, a feature only seen in 43% of leiomyomas with bizarre nuclei with a normal immunoprofile.
Furthermore, Joseph et al40 found eosinophilic cytoplasmic inclusions to be a fairly sensitive and specific feature of FH-deficient tumors, occurring in 80% (4/5) of 2SC-positive uterine leiomyomas but in <1% (1/189) of 2SC-negative tumors. In contrast, in our study, eosinophilic cytoplasmic inclusions were noted not only in our 2SC-positive leiomyomas with bizarre nuclei but also in 65% (9/14) of 2SC-negative tumors. Several possible explanations exist for this discrepancy. First, our study was limited to leiomyomas with bizarre nuclei, whereas only three leiomyomas studied by Joseph et al40 had bizarre nuclei. Second, as we have reported previously, cytoplasmic inclusions are a frequent and characteristic feature of leiomyomas with bizarre nuclei regardless of FH gene mutation status.9 Third, they observed these inclusions at low magnification, whereas we found them more apparent on high magnification. Finally, tumor sampling may also be a contributing factor. Joseph et al40 stated as a potential caveat (misinterpretation as cytoplasmic inclusions) the presence of cytoplasmic eosinophilia within smooth muscle cells in areas of infarction, but in our opinion, this does not manifest as discrete globules. Even though, the finding of eosinophilic cytoplasmic inclusions was predictive of abnormal immunoprofile on univariate analysis, it failed to be predictive on multivariate analysis as they were often seen in those with a normal immunophenotype.
We introduced the term ‘alveolar pattern of edema’ in 201510 as anecdotally it was noted that a number of leiomyomas with bizarre nuclei had a peculiar form of edema that differed from typical hydropic change. In this pattern, edema caused thinning and elongation of the smooth muscle cells, resulting in variably sized cystic spaces that superficially resembled pulmonary alveoli. Although this appearance was only observed in 42% of leiomyomas with bizarre nuclei in the current study, it was seen in 71% of tumors with an aberrant immunoprofile. Similarly, Harrison et al36 identified this pattern in 62% of FH-deficient leiomyomas analyzed. Although Joseph et al40 described a ‘zonated’ appearance in their leiomyomas with FH gene alterations, review of the image depicting this change allowed us to conclude that it was analogous to what we have designated as ‘alveolar pattern of edema’ in these tumors.
Even though only a limited number of studies have been performed, 2SC and FH immunohistochemical analysis has thus far demonstrated a strong association between aberrant staining and germline FH mutations in renal cell carcinomas11, 41, 42; however, in uterine leiomyomas, the significance of an abnormal 2SC/FH immunoprofile is less clear. Reyes et al8 first reported an association between 2SC expression and FH germline mutations. They identified nine leiomyomas with ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’, all of which showed diffuse 2SC expression. Although only three of these tumors underwent FH germline testing, two were shown to harbor the mutation. These two patients later developed either renal cell carcinoma or cutaneous leiomyomas. The authors speculated that many 2SC-positive leiomyomas might in fact result from somatic rather than germline mutations. Nonetheless, given that the results of this study were similar to those of the renal cell carcinoma series,11, 41, 42 it was suggested that immunohistochemical analysis might be a valuable and cost-effective method in triaging tumors for germline testing and referral to clinical genetics.
Although molecular studies on leiomyomas with an aberrant immunoprofile are limited, especially those implementing germline testing, data thus far favors these tumors to harbor somatic mutations.36, 38, 40 In a series of 194 leiomyomas from patients <40 years, Joseph et al40 identified 5 tumors that were 2SC-positive (2 of them completely FH-negative), and 4 of these were classified as ‘highly suspicious for hereditary leiomyomatosis and renal cell carcinoma syndrome’ based upon morphology. Molecular analysis identified somatic mutations in four leiomyomas and homozygous deletion of the FH gene in the other, but germline testing was not performed. Interestingly, in their cohort, three tumors were classified as leiomyomas with bizarre nuclei, but only one showed suspicious ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’ with aberrant staining and FH gene deletion. Harrison et al36 evaluated FH immunohistochemistry in five patients with either germline-proven FH mutations or a presumed mutation based on personal and/or family history, and all five showed loss of FH expression. They subsequently performed FH immunohistochemical analysis on an unselected cohort of 1152 leiomyomas, 12 of which showed loss of FH expression. Although somatic FH mutations were identified in six of the FH-deficient leiomyomas, germline testing revealed normal FH status in five (unsuccessful in the sixth). Similarly, Miettinen et al38 reported that 5% of the uterine smooth muscle tumors analyzed (n=1583) were FH deficient by immunohistochemistry, with most (79%) being leiomyomas with bizarre nuclei. They noted that, while similar histological features were often seen in both FH-deficient and -proficient leiomyomas with bizarre nuclei, those that were FH deficient were more likely to have staghorn vessels, prominent eosinophilic nucleoli, and perinucleolar halos. Miettinen et al38 also performed molecular analysis on 16 FH-deficient tumors (not specified as to whether leiomyomas with bizarre nuclei) and identified FH deletions in 5 and frameshift mutations in 3 leiomyomas.
From these studies, one can speculate that the majority of leiomyomas with bizarre nuclei displaying an abnormal FH immunoprofile are the result of somatic rather than FH germline mutations in the absence of personal or family history of hereditary leiomyomatosis and renal cell carcinoma syndrome. Even though these mutations likely lack clinical significance, FH immunohistochemical screening (and 2SC if it becomes commercially available) on all leiomyomas with bizarre nuclei and the aforementioned morphological features may be helpful as an adjunct to investigate further into the patient’s relevant personal and/or family history until more refined testing is commonly available. If FH expression is lost and the patient has a concerning personal and/or family history, then referral for genetic counseling and mutational testing may be warranted.
Our study had similar findings to the previously described series with ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’ being more common in leiomyomas with bizarre nuclei with an aberrant FH/2SC immunoprofile. Furthermore, we expanded on previous observations and demonstrated that leiomyomas with bizarre nuclei with an aberrant FH/2SC immunoprofile are genetically distinct from those with a normal FH/2SC staining pattern, with somatic FH genetic alteration restricted to the former and TP53 and/or RB1 alterations significantly more frequent in the latter. The concept of two subtypes of leiomyomas with bizarre nuclei has also been recently described by Ubago et al39 in which their type I atypical leiomyomas were more likely to show ‘hereditary leiomyomatosis and renal cell carcinoma syndrome features’, whereas type II atypical leiomyomas had a higher rate of TP53 and MED12 mutations. It should be noted that one of the cases harboring a somatic FH missense mutation coupled with loss of the wild-type allele (SCU14T) retained FH expression while also being 2SC-positive. Interestingly, FH expression has been observed in hereditary leiomyomatosis and renal cell carcinoma syndrome cases harboring somatic/germline FH mutations,42 suggesting that the sensitivity of FH analysis by immunohistochemistry may be reduced and better used in combination with 2SC staining, although the latter antibody is currently not commercially available. In one case lacking FH expression (SCU12T), no FH genetic alterations could be identified. Loss of FH expression in renal cell carcinoma has been suggested to be due to epigenetic or miRNA regulation,43 and further studies assessing alternative mechanisms of FH loss in leiomyomas are warranted.
MED12 mutations have been reported in all types of uterine smooth muscle tumors, with 31–92% of leiomyomas44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62 and in up to 30% of leiomyosarcomas.47, 50, 52, 54, 55, 61, 63, 64, 65, 66 In five studies, leiomyomas with bizarre nuclei were specifically evaluated for MED12 mutations and were found in up to 25% of tumors.39, 47, 50, 53, 61 Two studies detected somatic MED12 mutations in a small subset of leiomyomas from patients with germline-proven hereditary leiomyomatosis and renal cell carcinoma syndrome; however, loss of heterozygosity for the FH allele was not identified in any of these tumors.53, 59 Similarly, Liegl-Atzwanger et al61 found MED12 and FH somatic mutations to be mutually exclusive in their series of leiomyomas, leiomyomas with bizarre nuclei, and leiomyosarcomas. In our series of leiomyomas with bizarre nuclei, the frequency of somatic MED12 exon 2 hotspot mutations was only 8% (2/24), suggesting that these tumors are distinct from conventional leiomyomas. MED12 mutations were found in both tumors with aberrant and normal FH/2SC profile (SCU9T, FH homozygous deletion; SCU22T, FH wild-type; Figure 3a), and in support of the findings by Liegl-Atzwanger et al,61 MED12 and FH somatic mutations were mutually exclusive.
The main limitation in our study was the inability to perform germline FH testing on all patients due to institutional review board restrictions. Although it is possible that this subset of leiomyomas with bizarre nuclei with FH somatic deletions/mutations could also harbor a germline FH mutation, we find it unlikely given the lack of family/personal history of hereditary leiomyomatosis and renal cell carcinoma syndrome in these patients, as well as the overall low incidence of hereditary leiomyomatosis and renal cell carcinoma syndrome in the population. Furthermore, neither germline FH mutations were identified in tumors with somatic mutations in the study by Harrison et al36 nor was a germline mutation detected in the one patient in the current study.
In summary, we have demonstrated that leiomyomas with bizarre nuclei are morphologically and genetically heterogeneous and that hereditary leiomyomatosis and renal cell carcinoma syndrome morphological features are not specific to hereditary leiomyomatosis and renal cell carcinoma syndrome-associated leiomyomas as they can also be observed in a subset of leiomyomas with bizarre nuclei. Those with an increasing number of hereditary leiomyomatosis and renal cell carcinoma syndrome features, as well as ‘alveolar pattern of edema’, appear more likely to show aberrant FH/2SC expression. Furthermore, we showed that there are two subgroups of sporadic leiomyomas with bizarre nuclei, one with an aberrant FH/2SC immunoprofile and somatic FH mutations/deletions and one with a normal FH/2SC immunoprofile and enrichment in TP53 and/or RB1 genetic alterations.
References
Launonen V, Vierimaa O, Kiuru M et al. Inherited susceptibility to uterine leiomyomas and renal cell cancer. Proc Natl Acad Sci USA 2001;98:3387–3392.
Alderson NL, Wang Y, Blatnik M et al. S-(2-succinyl)cysteine: a novel chemical modification of tissue proteins by a Krebs cycle intermediate. Arch Biochem Biophys 2006;450:1–8.
Nagai R, Brock JW, Blatnik M et al. Succination of protein thiols during adipocyte maturation: a biomarker of mitochondrial stress. J Biol Chem 2007;282:34219–34228.
Ternette N, Yang M, Laroyia M et al. Inhibition of mitochondrial aconitase by succination in fumarate hydratase deficiency. Cell Rep 2013;3:689–700.
Yang M, Ternette N, Su H et al. The succinated proteome of FH-mutant tumours. Metabolites 2014;4:640–654.
Garg K, Tickoo SK, Soslow RA et al. Morphologic features of uterine leiomyomas associated with hereditary leiomyomatosis and renal cell carcinoma syndrome: a case report. Am J Surg Pathol 2011;35:1235–1237.
Sanz-Ortega J, Vocke C, Stratton P et al. Morphologic and molecular characteristics of uterine leiomyomas in hereditary leiomyomatosis and renal cancer (HLRCC) syndrome. Am J Surg Pathol 2013;37:74–80.
Reyes C, Karamurzin Y, Frizzell N et al. Uterine smooth muscle tumors with features suggesting fumarate hydratase aberration: detailed morphologic analysis and correlation with S-(2-succino)-cysteine immunohistochemistry. Mod Pathol 2014;27:1020–1027.
Croce S, Young RH, Oliva E . Uterine leiomyomas with bizarre nuclei: a clinicopathologic study of 59 cases. Am J Surg Pathol 2014;38:1330–1339.
Bennett JA, Chiang S, Chen Y et al. Leiomyoma with bizarre nuclei: correlation between morphology and fumarate hydratase/S-(2-succino)-cysteine expression. Mod Pathol 2015;28 (Suppl 2):276 A.
Chen YB, Brannon AR, Toubaji A et al. Hereditary leiomyomatosis and renal cell carcinoma syndrome-associated renal cancer: recognition of the syndrome by pathologic features and the utility of detecting aberrant succination by immunohistochemistry. Am J Surg Pathol 2014;38:627–637.
Chen YB, Xu J, Skanderup AJ et al. Molecular analysis of aggressive renal cell carcinoma with unclassified histology reveals distinct subsets. Nat Commun 2016;7:13131.
Martelotto LG, De Filippo MR, Ng CK et al. Genomic landscape of adenoid cystic carcinoma of the breast. J Pathol 2015;237:179–189.
Schultheis AM, Ng CK, De Filippo MR et al. Massively parallel sequencing-based clonality analysis of synchronous endometrioid endometrial and ovarian carcinomas. J Natl Cancer Inst 2016;108:djv427.
Cheng DT, Mitchell TN, Zehir A et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J Mol Diagn 2015;17:251–264.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
DePristo MA, Banks E, Poplin R et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 2011;43:491–498.
Cibulskis K, Lawrence MS, Carter SL et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013;31:213–219.
Saunders CT, Wong WS, Swamy S et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 2012;28:1811–1817.
Koboldt DC, Zhang Q, Larson DE et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 2012;22:568–576.
De Mattos-Arruda L, Weigelt B, Cortes J et al. Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle. Ann Oncol 2014;25:1729–1735.
Shen R, Seshan VE . FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res 2016;44:e131.
Piscuoglio S, Ng CK, Murray MP et al. The genomic landscape of male breast cancers. Clin Cancer Res 2016;22:4045–4056.
Carter SL, Cibulskis K, Helman E et al. Absolute quantification of somatic DNA alterations in human cancer. Nat Biotechnol 2012;30:413–421.
Carter H, Chen S, Isik L et al. Cancer-specific high-throughput annotation of somatic mutations: computational prediction of driver missense mutations. Cancer Res 2009;69:6660–6667.
Schwarz JM, Rodelsperger C, Schuelke M et al. MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 2010;7:575–576.
Shihab HA, Gough J, Cooper DN et al. Predicting the functional consequences of cancer-associated amino acid substitutions. Bioinformatics 2013;29:1504–1510.
Martelotto LG, Ng CK, De Filippo MR et al. Benchmarking mutation effect prediction algorithms using functionally validated cancer-related missense mutations. Genome Biol 2014;15:484.
Choi Y, Sims GE, Murphy S et al. Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 2012;7:e46688.
Kandoth C, McLellan MD, Vandin F et al. Mutational landscape and significance across 12 major cancer types. Nature 2013;502:333–339.
Futreal PA, Coin L, Marshall M et al. A census of human cancer genes. Nat Rev Cancer 2004;4:177–183.
Lawrence MS, Stojanov P, Mermel CH et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 2014;505:495–501.
Chang MT, Asthana S, Gao SP et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat Biotechnol 2016;34:155–163.
Merino MJ, Torres-Cabala C, Pinto P et al. The morphologic spectrum of kidney tumors in hereditary leiomyomatosis and renal cell carcinoma (HLRCC) syndrome. Am J Surg Pathol 2007;31:1578–1585.
Alsolami S, El-Bahrawy M, Kalloger SE et al. Current morphologic criteria perform poorly in identifying hereditary leiomyomatosis and renal cell carcinoma syndrome-associated uterine leiomyomas. Int J Gynecol Pathol 2014;33:560–567.
Harrison WJ, Andrici J, Maclean F et al. Fumarate hydratase-deficient uterine leiomyomas occur in both the syndromic and sporadic settings. Am J Surg Pathol 2016;40:599–607.
Martinek P, Grossmann P, Hes O et al. Genetic testing of leiomyoma tissue in women younger than 30 years old might provide an effective screening approach for the hereditary leiomyomatosis and renal cell cancer syndrome (HLRCC). Virchows Arch 2015;467:185–191.
Miettinen M, Felisiak-Golabek A, Wasag B et al. Fumarase-deficient uterine leiomyomas: an immunohistochemical, molecular genetic, and clinicopathologic study of 86 cases. Am J Surg Pathol 2016;40:1661–1669.
Ubago JM, Zhang Q, Kim JJ et al. Two subtypes of atypical leiomyoma: clinical, histologic, and molecular analysis. Am J Surg Pathol 2016;40:923–933.
Joseph NM, Solomon DA, Frizzell N et al. Morphology and immunohistochemistry for 2SC and FH aid in detection of fumarate hydratase gene aberrations in uterine leiomyomas from young patients. Am J Surg Pathol 2015;39:1529–1539.
Bardella C, El-Bahrawy M, Frizzell N et al. Aberrant succination of proteins in fumarate hydratase-deficient mice and HLRCC patients is a robust biomarker of mutation status. J Pathol 2011;225:4–11.
Chen Y, Kong M, Bialik A et al. Hereditary leiomyomatosis and renal cell carcinoma (HLRCC)-associated renal cancer: a comparison of fumarate hydratase (FH) and S-(2-succino)-cysteine (2SC) immunohistochemistry as ancillary tools. Mod Pathol 2015;28 (Suppl 2):211A.
Sudarshan S, Shanmugasundaram K, Naylor SL et al. Reduced expression of fumarate hydratase in clear cell renal cancer mediates HIF-2alpha accumulation and promotes migration and invasion. PLoS ONE 2011;6:e21037.
Makinen N, Mehine M, Tolvanen J et al. MED12, the mediator complex subunit 12 gene, is mutated at high frequency in uterine leiomyomas. Science 2011;334:252–255.
Makinen N, Heinonen HR, Moore S et al. MED12 exon 2 mutations are common in uterine leiomyomas from South African patients. Oncotarget 2011;2:966–969.
McGuire MM, Yatsenko A, Hoffner L et al. Whole exome sequencing in a random sample of North American women with leiomyomas identifies MED12 mutations in majority of uterine leiomyomas. PLoS ONE 2012;7:e33251.
Perot G, Croce S, Ribeiro A et al. MED12 alterations in both human benign and malignant uterine soft tissue tumors. PLoS ONE 2012;7:e40015.
Je EM, Kim MR, Min KO et al. Mutational analysis of MED12 exon 2 in uterine leiomyoma and other common tumors. Int J Cancer 2012;131:E1044–E1047.
Markowski DN, Bartnitzke S, Loning T et al. MED12 mutations in uterine fibroids—their relationship to cytogenetic subgroups. Int J Cancer 2012;131:1528–1536.
Matsubara A, Sekine S, Yoshida M et al. Prevalence of MED12 mutations in uterine and extrauterine smooth muscle tumours. Histopathology 2013;62:657–661.
Rieker RJ, Agaimy A, Moskalev EA et al. Mutation status of the mediator complex subunit 12 (MED12) in uterine leiomyomas and concurrent/metachronous multifocal peritoneal smooth muscle nodules (leiomyomatosis peritonealis disseminata). Pathology 2013;45:388–392.
de Graaff MA, Cleton-Jansen AM, Szuhai K et al. Mediator complex subunit 12 exon 2 mutation analysis in different subtypes of smooth muscle tumors confirms genetic heterogeneity. Hum Pathol 2013;44:1597–1604.
Makinen N, Vahteristo P, Kampjarvi K et al. MED12 exon 2 mutations in histopathological uterine leiomyoma variants. Eur J Hum Genet 2013;21:1300–1303.
Schwetye KE, Pfeifer JD, Duncavage EJ . MED12 exon 2 mutations in uterine and extrauterine smooth muscle tumors. Hum Pathol 2014;45:65–70.
Bertsch E, Qiang W, Zhang Q et al. MED12 and HMGA2 mutations: two independent genetic events in uterine leiomyoma and leiomyosarcoma. Mod Pathol 2014;27:1144–1153.
Heinonen HR, Sarvilinna NS, Sjoberg J et al. MED12 mutation frequency in unselected sporadic uterine leiomyomas. Fertil Steril 2014;102:1137–1142.
Wang H, Ye J, Qian H et al. High-resolution melting analysis of MED12 mutations in uterine leiomyomas in Chinese patients. Genet Test Mol Biomarkers 2015;19:162–166.
Shahbazi S, Fatahi N, Amini-Moghaddam S . Somatic mutational analysis of MED12 exon 2 in uterine leiomyomas of Iranian women. Am J Cancer Res 2015;5:2441–2446.
Kampjarvi K, Makinen N, Mehine M et al. MED12 mutations and FH inactivation are mutually exclusive in uterine leiomyomas. Br J Cancer 2016;114:1405–1411.
Sadeghi S, Khorrami M, Amin-Beidokhti M et al. The study of MED12 gene mutations in uterine leiomyomas from Iranian patients. Tumour Biol 2016;37:1567–1571.
Liegl-Atzwanger B, Heitzer E, Flicker K et al. Exploring chromosomal abnormalities and genetic changes in uterine smooth muscle tumors. Mod Pathol 2016;29:1262–1277.
Osinovskaya NS, Malysheva OV, Shved NY et al. Frequency and spectrum of MED12 exon 2 mutations in multiple versus solitary uterine leiomyomas from russian patients. Int J Gynecol Pathol 2016;35:509–515.
Kampjarvi K, Makinen N, Kilpivaara O et al. Somatic MED12 mutations in uterine leiomyosarcoma and colorectal cancer. Br J Cancer 2012;107:1761–1765.
Markowski DN, Huhle S, Nimzyk R et al. MED12 mutations occurring in benign and malignant mammalian smooth muscle tumors. Genes Chromosomes Cancer 2013;52:297–304.
Ravegnini G, Marino-Enriquez A, Slater J et al. MED12 mutations in leiomyosarcoma and extrauterine leiomyoma. Mod Pathol 2013;26:743–749.
Makinen N, Aavikko M, Heikkinen T et al. Exome sequencing of uterine leiomyosarcomas identifies frequent mutations in TP53, ATRX, and MED12. PLoS Genet 2016;12:e1005850.
Acknowledgements
We thank Jorge S Reis-Filho (MSKCC) for his critical assessment of the manuscript. Research reported in this paper was supported in part by a Cancer Center Support Grant of the National Institutes of Health/National Cancer Institute (Grant No. P30CA008748). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work was presented in part as a platform presentation at the 104th United States and Canadian Academy of Pathology (USCAP) Annual Meeting, Boston, MA, USA.
This work was presented in part as a platform presentation at the 104th United States and Canadian Academy of Pathology (USCAP) Annual Meeting, Boston, MA, USA.
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Bennett, J., Weigelt, B., Chiang, S. et al. Leiomyoma with bizarre nuclei: a morphological, immunohistochemical and molecular analysis of 31 cases. Mod Pathol 30, 1476–1488 (2017). https://doi.org/10.1038/modpathol.2017.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.56
This article is cited by
-
Corded and hyalinized endometrioid endometrial carcinoma with high-grade features: a clinicopathological and TCGA-based molecular analysis
Virchows Archiv (2023)
-
Targeted sequencing of FH-deficient uterine leiomyomas reveals biallelic inactivating somatic fumarase variants and allows characterization of missense variants
Modern Pathology (2020)
-
Leiomyoma with Bizarre Nuclei: a Study of 108 Cases Focusing on Clinicopathological Features, Morphology, and Fumarate Hydratase Alterations
Pathology & Oncology Research (2020)