Abstract
During systemic inflammatory pathologies, mediators of inflammation circulate in the bloodstream and interact with endothelial cells (ECs), resulting in endothelial dysfunction that maintains and enhances the pathological condition. Inflammatory mediators change the protein expression profile of ECs, which become activated fibroblasts via endothelial-to-mesenchymal transition. This process is characterized by downregulated endothelial proteins and strongly upregulated fibrotic-specific genes and extracellular matrix-forming proteins. The main inductor of endothelial fibrosis is transforming growth factor-β1 (TGF-β1), which acts through the TGF-β1/activin receptor-like kinase 5 (ALK5)/Smads intracellular signaling pathway. The signal transducer and activator of transcription 3 (STAT3) is also involved in fibrosis in several tissues (e.g. heart and vascular system), where STAT3 signaling decreases TGF-β1-induced responses by directly interacting with Smad proteins, suggesting that decreased STAT3 could induce TGF-β1-mediated fibrosis. However, it is unknown if suppressed STAT3 expression induces EC fibrosis through a mechanism involving the TGF-β signaling pathway. The present study evaluated the fibrotic actions of STAT3 suppression in ECs and investigated TGF-β1/ALK5/Smad4 signaling pathway participation. Suppressed STAT3 expression stimulated fibrotic conversion in ECs, as mediated by protein expression reprograming that decreased endothelial marker expression and increased fibrotic and extracellular matrix protein levels. The potential mechanism underlying these changes was dependent on TGF-β1 secretion, the ALK5 activation pathway, and Smad4 translocation into the nucleus. We conclude that suppressed STAT3 expression converts ECs into activated fibroblasts via TGF-β1/ALK5/Smad4 signaling pathway involvement.
Similar content being viewed by others
Main
Endothelial cells (ECs), which compose the inner layer of blood vessels that form the endothelium, participate in several physiological functions, including vasorelaxation, hemostasis control, and vascular permeability, among others. However, several inflammatory conditions are associated with endothelial dysfunction, including hypertension, atherosclerosis, thrombosis, obesity, and diabetes. During inflammation, proinflammatory molecules interact with ECs on the inner walls of blood vessels, thereby generating several actions, including endothelial fibrosis.1, 2, 3, 4
Endothelial fibrosis is mediated by a widely reported process known as endothelial-to-mesenchymal transition (EndMT).5, 6, 7, 8, 9 There are several efficient inductors of EndMT, including cytokines and mediators of inflammation such as transforming growth factor-β1 (TGF-β1), TGF-β2, TNF-α, IL-1β, reactive oxygen species, and endotoxins, among various others.5, 6, 7, 9, 10, 11, 12, 13 Of these inductors, TGF-β types 1 and 2 participate in the main pathway for fibrotic intracellular signaling in ECs. Activation of the plasma membrane receptor of TGF-β, activin receptor-like kinase 5 (ALK5), stimulates intracellular signaling. Several proteins are involved in the TGF-β/ALK5 pathway, such as numerous Smad family proteins. Specifically, after activation of Smad2 and Smad3, the Smad4 protein is phosphorylated to promote translocation to the nucleus and the subsequent transcription of genes supporting profibrotic actions.14, 15
During endothelial fibrosis, ECs undergo protein expression reprograming and convert into activated fibroblasts. This process is characterized by the downregulation of endothelial proteins, such as vascular endothelial (VE)-cadherin and cluster of differentiation 31 (CD31)/PECAM, and the upregulation of fibrotic-specific markers, such as fibroblast-specific protein 1 (FSP-1), α-smooth muscle actin (α-SMA) and vimentin. Furthermore, the levels of extracellular matrix (ECM)-forming proteins, such as fibronectin and collagen types I and III, strongly increase during EndMT.5, 6, 7, 9
The signal transducer and activator of transcription 3 (STAT3) is involved in generating fibrosis in several tissues. Supporting this, tissue-specific, cardiomyocyte-restricted STAT3 knockout (STAT3-KO) mice exhibit enhanced cardiac fibrosis.16 Similarly, cardiomyocyte-restricted STAT3-KO mice show increased interstitial, ECM deposition-mediated fibrosis that affects cardiac function, impairs angiogenesis, and promotes dysfunctional vasculature maintenance, suggesting that the deleterious effects of STAT3 downregulation affect the heart and even extend to the vascular system.17, 18 In contrast, evidence in other tissues, such as the liver, indicates that STAT3 generates hepatic fibrosis through TGF-β1 production.19 Similarly, STAT3 activation in the kidney and lungs is associated with renal and pulmonary fibrosis.20, 21, 22 Therefore, both the inhibition and activation of STAT3 are related to fibrosis generation. Considering this context, we hypothesize that STAT3 exerts differential effects depending on the organ and tissue type. As a particular point of investigation within this subject area, we questioned whether the suppression of STAT3 expression would induce fibrotic-like conversion in ECs.
Several ligands can activate STAT3, including interleukins, growth factors, and toll-like receptor ligands. STAT3 is activated through phosphorylation, which promotes nuclear translocation. In the nucleus, STAT3 controls the transcriptional initiation and activation of several processes, including those associated with survival, proliferation, and fibrosis.23, 24 Interestingly, crosstalk might exist between STAT3 and TGF-β1 signaling. Reports in in vivo and in vitro models indicate that STAT3 signaling decreases TGF-β1-induced responses through a direct interaction between Smad3 and STAT3, resulting in a reduction of the Smad3–Smad4 complex. Therefore, decreased STAT3 promotes TGF-β-mediated transcriptional responses, including epithelial-to-mesenchymal transition.25 In another context, it has also been shown that increased STAT3 levels result in a cooperation between this transcription factor and TGF-β1 that exacerbates fibrosis.26, 27
Despite this existing knowledge, it remains unknown if the suppression of STAT3 expression induces EC fibrosis through a mechanism involving the TGF-β signaling pathway. Therefore, the aims of this study were to assess the fibrotic actions of STAT3 suppression on ECs and to investigate the participation of the TGF-β1/ALK5/Smad signaling pathway in this. The obtained results demonstrate that the expressional suppression of STAT3 stimulates a fibrotic-like phenotype in ECs. Furthermore, STAT3 downregulation-induced endothelial fibrosis was the result of protein expression reprograming that promoted a decreased expression of endothelial markers (VE-cadherin and CD31) and an increased expression of both fibrotic (FSP-1 and α-SMA) and ECM (fibronectin and collagen type III) proteins. The potential underlying mechanism for EC fibrosis appears dependent on TGF-β1 secretion, the ALK5 activation pathway, and Smad proteins signaling to mediate changes in protein expressions. These findings provide a better understanding of the mechanisms involved in fibrosis-induced endothelial dysfunction, information that can be applied to improve current treatments of vascular pathologies.
Materials and methods
Primary ECs and Vessel Cultures
Rat mesenteric ECs (RMECs) were isolated from Sprague–Dawley rats, ranging ~150–220 g in body weight. Animals were anesthetized and subjected to surgery. Primary ECs: the mesenteric artery was occluded close to the right kidney by tying it up, and then cannulated with a polyethylene tubing connected to a 21-gauge syringe. The mesentery was surgically removed and washed with sterile PBS. For the enzymatic isolation of RMECs, each mesenteric artery was slowly perfused in a culture hood for 5 min with 5 ml of M-199 medium supplemented with 40 μl Pen/Strept (10 000 U/ml/10 000 μg/ml), 20 μl Fungizone (250 μg/ml), and 12.5 mg collagenase type II. The cell suspension was centrifuged at 3000 r.p.m. for 7 min; the pellet was reconstituted on 3 ml of M-199 medium supplemented with 8 ml/l Pen/Strept (10 000 U/ml/10 000 μg/ml), 4 ml/l Fungizone (250 μg/ml), 10% FBS, and 10% CCS, and then plated on gelatin-coated wells in a six-well plate. Cells were allowed to attach to the gel matrix for 4 h, then rinsed once with sterile PBS, and finally cultured on M-199 medium supplemented with 8 ml/l Pen/Strept (10 000 U/ml/10 000 μg/ml), 4 ml/l Fungizone (250 μg/ml), 10% FBS, and 10% CCS. The primary culture was grown at 37 °C in a 5% CO2 incubator for 3 days. Aorta artery culture: intact aorta arteries from rats were cultured with the M-199 medium supplemented with 8 ml/l Pen/Strept (10 000 U/ml/10 000 μg/ml), 4 ml/l Fungizone (250 μg/ml), 10% FBS, and 10% CCS, as elsewhere reported by our group.9 Artery segments were treated for 24 h with culture medium containing the vehicle or the STAT3 inhibitor, S3I-201. The conducted investigation conformed with principles outlined in the Declaration of Helsinki. The Commission of Bioethics and Biosafety of Universidad Andres Bello also approved all experimental protocols.
Small Interfering RNA and Transfection
Small interfering RNA (siRNA) per STAT3 transcript (siRNA-STAT3) and non-targeting siRNA (siRNA-CTRL), used as a control, were purchased from Dharmacon (Lafayette, CO). Briefly, RMECs were transfected with 5 nM siRNA using the DharmaFECT 4 transfection reagent (Dharmacon) according to the manufacturer's protocols in a serum-free medium.
Quantitative Real-Time PCR and Western Blot Procedures
Equal amounts of RNA were used as templates in each reaction. qPCR analysis was performed using the SYBR Green PCR Master Mix (AB Applied Biosystems, Foster City, CA). Assays were run using an Eco Real-Time PCR System (Illumina). Data are presented as the relative mRNA levels for the gene of interest normalized to relative levels of 28S mRNA.
ECs transfected with siRNA-CTRL or siRNA-STAT3 were lysed in cold lysis buffer, and then proteins were extracted. Supernatants were collected and stored in the same lysis buffer. The protein extract and supernatant were subjected to SDS-PAGE, and resolved proteins were transferred to a nitrocellulose or PVDF membrane. The blocked membrane was incubated with the appropriate primary antibody, washed two times, and incubated with a secondary antibody. Bands were revealed using a peroxidase-conjugated IgG antibody. Peroxidase activity was detected through enhanced chemiluminescence (Bio-Rad, CA), and images were acquired using Fotodyne FOTO/Analyst Luminary Workstations Systems (Fotodyne, Hartland, WI). Protein content was determined by densitometric scanning of immunoreactive bands, and intensity values were obtained by densitometry of individual bands normalized against tubulin. For a detailed list of antibodies used in western blot procedures, see Supplementary Table S1.
Fluorescent Immunocytochemistry and Immunohistochemistry
Fluorescent immunocytochemistry: Cells were washed two times with PBS and fixed with 3.7% paraformaldehyde (PFA) for 30 min at room temperature (RT) before being permeabilized with 0.1% Triton X-100 in PBS for 30 min at RT, and blocked for 2 h at RT with 3% BSA in PBS. The cells were subsequently washed again and incubated with the primary antibodies. Then, cells were washed two times and incubated with the secondary antibodies. Samples were mounted with ProLong Gold antifade mounting medium with DAPI (Invitrogen). For a detailed list of antibodies used in fluorescent immunocytochemistry experiments, see Supplementary Table S2.
Fluorescent immunohistochemistry: Samples obtained from rat aorta artery were fixed with PFA 3.7 % for 1 h at RT, permeabilized with 0.1% Triton X-100 in PBS for 30 min at RT, and blocked for 3 h at RT with 3% BSA in PBS, and 50 mM NH4Cl in PBS for 20 min a RT. Samples were subsequently washed and incubated with the first primary antibodies. Then, cells were washed two times and incubated with the first secondary antibodies. Nuclei were stained with Hoechst (Sigma). For a detailed list of antibodies used, see Supplementary Table S2.
FITC-Dextran Transwell Assay
The RMEC and EA hy926 ECl monolayers were plated onto the insert of the transwell (Costar Transwell; Corning, New York, NY) and cultured until confluent. Then, cells were cultured in the presence of a vehicle or the STAT inhibitor S3I-201 for 72 h. At the end of the experiments, 0.5 mg/ml fluorescein isothiocyanate (FITC)-dextran 40 kDa (Sigma-Aldrich, St Louis, MO) was added to the upper chamber of the transwell. After 90 min, the samples were removed from the bottom compartment, and fluorescence was quantified (excitation 485 nm, emission 520 nm).
Reagents
Buffers and salts were purchased from Merck Biosciences. SB431542 (0.5 μM), S3I-201 (5–10 μM) and SIS3 (10 μM) were purchased from Tocris.
Data Analysis
All results are presented as the means±s.d.. Statistical differences were assessed by Student's t-test (Mann–Whitney) or one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. Differences were considered significant at P<0.05.
Results
Inhibiting STAT3 Expression in ECs Induces a Fibroblast-Like Phenotype
Non-transfected ECs showed a round and short-spindle morphology with a cobblestone appearance (Figure 1a). Similarly, ECs transfected with a non-targeting siRNA (siRNA-CTRL) used as a control exhibited a similar morphology to that observed in non-transfected ECs (Figure 1b). In contrast, ECs transfected with a siRNA against STAT3 (siRNA-STAT3) showed a spindle-shaped phenotype typical of fibroblasts (Figure 1c), suggesting the occurrence of an EndMT process. To investigate these phenotypic changes in more detail, cells were counted that were at least twice as long as wide (length/width >2). The lengths of ECs transfected with siRNA-STAT3 were ~6-fold longer than both non-transfected cells and siRNA-CTRL-transfected cells (Figure 1d). In addition to this, cell lengths were measured in each condition, with results showing a differential distribution of lengths. The mean cell length of ECs transfected with siRNA-STAT3 was ~3.5-fold longer than that of both non-transfected cells and siRNA-CTRL-transfected cells (Figure 1e).
Suppression of signal transducer and activator of transcription (STAT) expression generates a fibroblast-like morphology in endothelial cells (ECs). (a–c) Morphological changes in ECs resembling fibroblasts. Figures show representative phase-contrast images from at least three separates experiments of (a) non-transfected ECs and (b) in ECs transfected with a specific small interfering RNA (siRNA) against STAT3 (siRNA-STAT3) or (c) a non-targeting siRNA (siRNA-CTRL). Scale bar represents 50 μm. (d) EC length in which length/width >2, for non-transfected ECs and ECs transfected with a specific siRNA against STAT3 (siRNA-STAT3) or a non-targeting siRNA (siRNA-CTRL). (e) EC length distribution of non-transfected ECs (filled bars) and ECs transfected with a specific siRNA against STAT3 (siRNA-STAT3) (empty bars) or a non-targeting siRNA (siRNA-CTRL, gray bars). Statistical differences were assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. ***P<0.001 against the non-transfected condition. Graph bars show the mean±s.d. (N=4–7).
To assess the enrichment of endothelial cultures, a detailed examination was carried out using VE-cadherin as a specific endothelial marker. Over 99% of cells in the EC cultures were positive for VE-cadherin, demonstrating that the primary EC cultures were highly enriched in ECs, without potential contamination from fibroblasts or mesenchymal-like cells (Supplementary Figure S1). Transfection with siRNA-STAT3 was >90% efficient in downregulating STAT3 expression as compared with ECs transfected with siRNA-CTRL (Supplementary Figures S2a and b).
Suppression of STAT3 Expression Induces Decreased Endothelial Protein Expression and Increased Fibrotic Marker and ECM Protein Expression in ECs
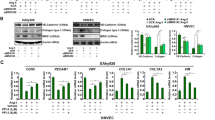
ECs transfected with siRNA-STAT3 showed a decreased expression of endothelial proteins as compared with both siRNA-CTRL-transfected cells and non-transfected cells. siRNA-STAT3-transfected ECs showed a decreased expression of the endothelial proteins VE-cadherin (Figures 2a and b) and CD31 (Figures 2c and d). Furthermore, ECs transfected with siRNA-STAT3 showed an increased expression of the fibrotic-specific markers α-SMA (Figures 2e and f) and FSP-1 (Figures 2g and h), as compared with both siRNA-CTRL-transfected and non-transfected cells. Considering that ECM protein accumulation is a crucial stage in fibrogenesis, it was relevant to also determine fibronectin and collagen protein levels in the supernatants of EC cultures. The expressions of fibronectin (Figures 2i and j) and collagen type III (Figures 2k and l) were significantly increased in cells transfected with siRNA-STAT3, whereas both non-transfected and siRNA-CTRL-transfected ECs did not exhibit any increases in fibronectin and collagen.
Suppression of signal transducer and activator of transcription 3 (STAT3) expression induces changes in endothelial, fibrotic, and extracellular matrix (ECM) proteins. Protein expression was analyzed in non-transfected (NT) endothelial cells (ECs) and in ECs transfected with a specific small interfering RNA (siRNA) against STAT3 (siRNA-STAT3) or a non-targeting siRNA (siRNA-CTRL). (a, c, e, g, i, and k) Representative images from western blot experiments performed to detect (a) VE-cadherin (VE-Cad), (c) cluster of differentiation 31 (CD31), (e) α-smooth muscle actin (α-SMA), (g) fibroblast-specific protein 1 (FSP-1), (i) fibronectin (FN), and (k) Col III. (b, d, f, h, j, and l) Densitometric analyses of the experiments shown in (a, c, e, g, i, and k, respectively). Protein levels were normalized against tubulin, and data are expressed relative to siRNA-CTRL-transfected cells (N=5). Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d.
As a compliment to siRNA technology, experiments were performed using the pharmacological inhibitor of STAT3, S3I-201. ECs exposed to S3I-201 showed decreased protein levels of the endothelial marker VE-cadherin (Supplementary Figures S3a and b) and an increased expression of the fibrotic markers α-SMA (Supplementary Figures S3c and d) and fibronectin (Supplementary Figures S3e and f). The efficiency of the STAT3 inhibitor was validated for the doses used (Supplementary Figure S4).
Of note, using the S3I-201 strategy, similar results were observed in the HUVEC-derived EC line EA hy926.28 EA cells were cultured in the presence or absence of the STAT3 inhibitor, and the expression of endothelial and fibrotic markers were analyzed. EA cells exposed to S3I-201 showed a decreased expression of the endothelial proteins VE-cadherin and CD31 (Supplementary Figures S5a–d, respectively), whereas expression of the fibrotic-specific markers α-SMA and FSP-1 (Supplementary Figures S5e–h, respectively) was severely increased, as compared with ECs in the absence of the inhibitor. Similarly, the expression of the ECM proteins fibronectin and collagen type III (Supplementary Figures S5i–l, respectively) were significantly increased in EA cells exposed to the STAT3 inhibitor, whereas vehicle-treated cells did not exhibit any increase in fibronectin and collagen.
Taken together, the obtained results suggest that endothelial fibrosis induced by suppressed STAT3 expression possibly occurs in several EC types.
Suppressing STAT3 Expression Induces Changes in Endothelial, Fibrotic and ECM Proteins Distribution in ECs
As a next step, the changes induced by STAT3 downregulation were studied in the context of cellular localization and distribution of endothelial and fibrotic proteins. Non-transfected ECs and cells transfected with siRNA-CTRL exhibited typical VE-cadherin labeling that was predominantly located in the plasma membrane, whereas FSP-1 expression was almost undetectable (Figures 3a and b). Additionally, CD31 labeling was detected at the plasma membrane, whereas α-SMA was weakly expressed (Figures 3d and e). In contrast, ECs transfected with siRNA-STAT3 displayed increased FSP-1 labeling and decreased VE-cadherin expression (Figure 3c). Furthermore, α-SMA labeling in fibrotic-like stress fibers was greatly increased in the intracellular compartment, whereas CD31 was virtually absent (Figure 3f).
Cellular distribution of endothelial and fibrotic markers involved in endothelial fibrosis induced by suppression of signal transducer and activator of transcription 3 (STAT3) expression. Representative images from experiments (a, d, g, and j) of non-transfected endothelial cells (ECs) and (b, e, h, and k) in ECs transfected with a non-targeting small interfering RNA (siRNA) (siRNA-CTRL) or (c, f, i, and l) siRNA against STAT3 (siRNA-STAT3). Endothelial markers cluster of differentiation 31 (CD31) or VE-cadherin (red), and the fibrotic markers α-smooth muscle actin (α-SMA), fibroblast-specific protein 1 (FSP-1), or fibronectin (FN) were detected (green). Boxes depicted in (b, c, e, f, h, i, k, and l) indicate the magnification shown in (b', c', e', f', h', i', k', and l'), respectively. Nuclei were stained using 4',6-diamidino-2-phenylindole (DAPI). Scale bar represents 30 μm.
To investigate the effect produced by STAT3 downregulation on the cellular localization and distribution of ECM proteins, these proteins were evaluated. ECs transfected with siRNA-STAT3 showed increased fibronectin labeling and decreased VE-cadherin (Figure 3i) and CD31 (Figure 3l) expressions. In contrast, non-transfected ECs and cells transfected with siRNA-CTRL showed typical VE-cadherin (Figures 3g and h) and CD31 (Figures 3j and k) labeling, which was restricted to the plasma membrane, whereas fibronectin (Figures 3g, h and j, k) was expressed at low levels.
Endothelial Fibrosis Induced by Suppression of STAT3 Expression in Endothelial Monolayers from Intact Blood Vessels
Next, we assessed if STAT3 suppression-induced endothelial fibrosis occurs in ECs in the native tridimensional environment of intact blood vessels. For this, experiments were performed in an intact aorta artery exposed to the vehicle or STAT3 inhibitor, S3I-201, for 24 and 48 h. ECs from the vehicle-treated aorta artery showed clear VE-cadherin labeling at the cell limits but low fibronectin staining (Figures 4a and c). In contrast, ECs from the S3I-201-treated aorta showed a significant downregulation of VE-cadherin staining at 48 h compared with the vehicle-treated aorta artery, whereas fibronectin labeling was greatly increased (Figures 4d and d′). Also, in 24 h treatment samples showed a weak VE-cadherin expression decrease and fibronectin labeling increase (Figures 4b and b′). These results indicate that STAT3 inhibition was able to initiate endothelial fibrosis in the endothelial monolayer of intact whole blood vessels. No significant changes were observed in tissue morphology between vessels at 0 vs 24 and 48 h of treatment (data not shown), suggesting that blood vessel culturing itself did not promote any alterations.
Cellular distribution of endothelial and fibrotic markers involved in endothelial fibrosis induced by signal transducer and activator of transcription 3 (STAT3) inhibition in intact whole aorta artery. Endothelial cells (ECs) from intact aorta artery treated with vehicle (a and c), or treated with 10 μM of STAT3 inhibitor (S3I-201) at 24 (b and b') or 48 h (d and d'). Endothelial marker VE-cadherin (red) and the fibrotic marker fibronectin (FN) were detected (green). Boxes depicted in (b and d) indicate the magnification shown in (b' and d'), respectively. Nuclei were stained using Hoechst. Scale bar represents 50 μm.
Endothelial Fibrosis Induced by Suppression of STAT3 Expression is Mediated through TGF-β1 Secretion and ALK5 Activation
It has been broadly reported that TGF-β1 is a main fibrotic inducer.5, 29 Furthermore, several profibrotic stimuli are supported by the production of TGF-β1.11, 12, 30 These previous findings prompted us to investigate if STAT3 downregulation induces the production of TGF-β1 as a mechanism to promote endothelial fibrosis.
ECs transfected with the siRNA-STAT3 exhibited an increased mRNA expression of TGF-β1 (Figure 5a), whereas TGF-β2 mRNA expression did not show change (Figure 5b). In concordance with this, siRNA-STAT3-transfected ECs showed increased TGF-β1 protein levels in the supernatant (Figures 5c and d) and protein extract (data not shown) as compared with both non-transfected cells and ECs transfected with siRNA-CTRL. These findings suggest that STAT3 expression suppression promotes an increased expression and secretion of TGF-β1 in ECs.
Suppression of signal transducer and activator of transcription 3 (STAT3) expression induces the expression and secretion of tumor growth factor-β1 (TGF-β1). (a and b) Endothelial cells (ECs) were transfected with small interfering RNA (siRNA)-CTRL, siRNA-STAT3 or non-transfected (NT), and mRNA expression of (a) TGF-β1 and (b) TGF-β2 was then measured by means of quantitative PCR (qPCR). Determinations were performed in at least triplicate, and the results are expressed normalized relative to 28S mRNA expression. (N=3–4). (c and d) ECs were transfected with small interfering RNA (siRNA)-CTRL, siRNA-STAT3, or NT, and the protein secretion of TGF-β1 was then measured in the supernatant. (c and d) Representative images of western blot experiments performed for detection of (c) TGF-β1 secretion. (d) Densitometric analyses of the experiments are shown in (c). Protein levels were normalized against tubulin, and data are expressed relative to the NT condition. Significant differences were assessed by a Student's t-test (Mann–Whitney). **P<0.01, ***P<0.001 against NT condition. Graph bars show the mean±s.d.
TGF-β actions are mediated by means of its receptor, ALK5. Therefore, assays were conducted to evaluate if ALK5 expression was modified by STAT3 expression suppression. ECs transfected with siRNA-STAT3 did not show change in ALK5 expression as compared with both siRNA-CTRL-transfected cells and non-transfected cells. This finding suggests that activation of the TGF-β1/ALK5/Smad4 signaling pathway by STAT3 suppression is dependent on increased TGF-β1 expression (Supplementary Figure S6).
During analysis of the above results, we questioned if the increased TGF-β1 protein levels were mediating the STAT3 suppression-induced endothelial fibrosis. The actions of TGF-β are carried out by this peptide, thus activating the ALK5 receptor to induce gene transcription and promote fibrosis.14, 15 Therefore, experiments were performed using SB431542, a specific ALK5 activation inhibitor. Frequently used SB431542 doses are toxic to EC cultures. For this reason, a non-toxic, but still effective, concentration of SB431542 for assay use was previously determined.9 ECs transfected with siRNA-STAT3 in the presence of SB431542 did not evidence decreased levels of the endothelial markers VE-cadherin (Figures 6a and b) and CD31 (Figures 6c and d). Furthermore, siRNA-STAT3-transfected cells exposed to SB431542 did not show any increase in expression of the fibrotic proteins α-SMA (Figures 6e and f) and FSP-1 (Figures 6g and h). Similarly, SB431542 treatment of siRNA-STAT3-transfected cells abolished the increased protein levels of fibronectin (Figures 6i and j) and collagen type III (Figures 6k and l). These results suggest that ALK5 activity is required for endothelial and fibrotic expression changes to occur during STAT3 supression-induced endothelial fibrosis.
Suppression of signal transducer and activator of transcription 3 (STAT3) expression induces changes in endothelial, fibrotic, and extracellular matrix (ECM) proteins dependent on ALK5 activity. Protein expression was analyzed in non-transfected (NT) endothelial cells (ECs) and in ECs transfected with a specific small interfering RNA (siRNA) against STAT3 (siRNA-STAT3) or a non-targeting siRNA (siRNA-CTRL) in the absence (–) or presence (+) of SB431542 (0.5 μM). (a, c, e, g, i, and k) Representative images from western blot experiments performed to detect (a) VE-cadherin (VE-Cad), (c) cluster of differentiation 31 (CD31), (e) α-smooth muscle actin (α-SMA), (g) fibroblast-specific protein 1 (FSP-1), (i) fibronectin (FN), and (k) Col III. (b, d, f, h, j and l) Densitometric analyses of the experiments shown in (a, c, e, g, i, and k, respectively). Protein levels were normalized against tubulin, and data are expressed relative to siRNA-CTRL-transfected cells (N=5). Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d.
Furthermore, the inhibition of ALK5 activation also abolished the cellular distribution of endothelial and fibrotic markers induced by STA3 suppression. In the absence of SB431542, non-transfected ECs and cells transfected with siRNA-CTRL displayed predominant VE-cadherin (Figures 7a, b and m, n) and CD31 (Figures 7g, h and s, t) staining close to the plasma membrane, whereas FSP-1 (Figures 7a and b), α-SMA (Figures 7g and h), and fibronectin (Figures 7m, n and s, t) expressions were almost undetectable. In contrast, ECs transfected with siRNA-STAT3 displayed increased FSP-1 and α-SMA labeling and decreased VE-cadherin and CD31 expression (Figures 7c and i), as well as strongly increased fibronectin expression (Figures 7o and u). However, in the presence of SB431542, ECs transfected with siRNA-STAT3 did not show any changes in protein localization (Figures 7f, l, r, and y) as compared with both non-transfected ECs (Figures 7d, j, p, and v) and cells transfected with siRNA-CTRL (Figures 7e, k, q, and x).
Cellular distribution of endothelial and fibrotic markers involved in endothelial fibrosis induced by suppression of signal transducer and activator of transcription 3 (STAT3) expression dependent on ALK5 activity. Representative images from experiments (a, d, g, j, m, p,s, and v) of non-transfected endothelial cells (ECs) and (b, e, h,k,n,q, t, and x) in ECs transfected with siRNA-CTRL or (c, f, i,l,o,u, and x) siRNA-STAT3 in the absence (–) or presence (+) of SB431542 (0.5 μM). Endothelial markers cluster of differentiation 31 (CD31) or VE-cadherin (red), and the fibrotic markers α-smooth muscle actin (α-SMA), fibroblast-specific protein 1 (FSP-1), or fibronectin (FN) were detected (green). Nuclei were stained using 4',6-diamidino-2-phenylindole (DAPI). Scale bar represents 30 μm.
Endothelial Fibrosis Induced by Suppression of STAT3 Expression is Mediated through Smad3 Activity and Nuclear Translocation of Smad4
Once ALK5 is activated by TGF-β, the Smad2 and Smad3 proteins are activated and consequently bind to Smad4, inducing the cytoplasm–nucleus migration of this protein complex. This migration elicits changes in protein expressions, leading to the generation of fibrosis.14, 15 Therefore, Smad2/3 activation and Smad4 translocation are decisive steps in triggering and maintaining fibrosis. Considering that the TGF-β1/ALK5 pathway was observed mediating endothelial fibrosis, we addressed the question of whether Smad3 activation participates in endothelial fibrosis induced by suppression of STAT3 expression. To this end, experiments were performed using SIS3, a specific Smad3 inhibitor.11
ECs transfected with siRNA-STAT3 in the presence of SIS3 did not show decreased levels of the endothelial marker VE-cadherin (Figure 8a). In line with this, siRNA-STAT3-transfected cells exposed to SIS3 did not show any increased expression of the fibrotic protein α-SMA (Figure 8b). Similarly, SIS3 treatment of siRNA-STAT3-transfected cells abolished the increased protein levels of fibronectin (Figure 8c). These results suggest that SIS3 activity is necessary for STAT3 supression-induced endothelial fibrosis.
Suppression of signal transducer and activator of transcription 3 (STAT3) expression induces changes in endothelial, fibrotic, and extracellular matrix (ECM) proteins dependent on Smad3 activity. Protein expression was analyzed in non-transfected (NT) ECs and in ECs transfected with a specific small interfering RNA (siRNA) against STAT3 (siRNA-STAT3) or a non-targeting siRNA (siRNA-CTRL) in the absence (–) or presence (+) of SIS3 (10 μM). Densitometric analyses from western blot experiments performed to detect (a) VE-cadherin (VE-Cad), (b) α-smooth muscle actin (α-SMA), and (c) fibronectin (FN). Protein levels were normalized against tubulin, and data are expressed relative to siRNA-CTRL-transfected cells (N=4). Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d.
Next, we questioned and sought to answer if Smad4 translocation is induced by STAT downregulation and requires ALK5 activity. In non-transfected and siRNA-CTRL-transfected ECs, Smad4 was principally detected in the cytoplasm (Figures 9a and b). Conversely, ECs transfected with siRNA-STAT3 showed a strong, significant translocation of Smad4 to the nucleus (Figure 9c). Densitometric analysis showed that the nucleus/cytosol ratio of Smad4 was highly increased when ECs were transfected with siRNA-STAT3, as compared with non-transfected and siRNA-CTRL-transfected ECs (Figure 9d). The Smad4 translocation observed in cells transfected with siRNA-STAT3 was similar to that observed in ECs exposed to exogenous TGF-β1 (Supplementary Figure S7).
Suppression of signal transducer and activator of transcription 3 (STAT3) induces the translocation of smad4 into the nucleus in endothelial cells (ECs). (a–c) Representative images showing the detection of Smad4 (green) in (a) non-transfected (NT) ECs and (b) ECs transfected with small interfering RNA (siRNA)-CTRL or (c) siRNA-STAT3. Boxes depicted in (d) indicate the magnification shown in (a'–c'), respectively. Nuclei were stained using 4',6-diamidino-2-phenylindole (DAPI). Scale bar represents 30 μm. (d) Densitometric analyses of the experiments shown in (a–c). Data were expressed as the normalized ratio of smad4 signal (pixels/μm2) in the nucleus/cytosol (N=5). The number of analyzed cells is depicted in bars. Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d.
In line with this observation, SB431542 was used to determine whether ALK5 participates in Smad4 translocation when STAT3 is suppressed. Non-transfected ECs and cells transfected with siRNA-CTRL did not show any changes in Smad4 translocation in the absence (Figures 10a and b) or presence (Figures 10d and e) of SB431542. Interestingly, ECs transfected with siRNA-STAT3 showed Smad4 translocation in the absence of SB431542 (Figure 10c), whereas in the presence of the ALK5 inhibitor (+SB431542), translocation was completely inhibited (Figure 10f). Densitometric analysis showed that the nucleus/cytosol ratio of Smad4 was severely inhibited when ECs were exposed to SB431542 (Figure 10h), as compared with cells without the ALK5 inhibitor (Figure 10g). Supporting these results, a nuclear-rich fraction extracted from ECs transfected with siRNA-STAT3 showed higher Smad4 levels compared with non-transfected ECs and cells transfected with siRNA-CTRL (Figures 10i and j). However, this translocation was inhibited using SB431542. In the presence of the inhibitor, the nuclear-rich fraction extracted from ECs transfected with siRNA-STAT3 did not show any change compared with non-transfected ECs and cells transfected with siRNA-CTRL (Figures 10m and n). The cytosol-rich fractions did not show significant changes in the absence (Figures 10k and l) and presence (Figures 10o and p) of SB431542.
Suppression of signal transducer and activator of transcription 3 (STAT3) induces the translocation of smad4 into the nucleus in endothelial cells (ECs) is dependent on ALK5 activity. (a–f) Representative images showing the detection of Smad4 (green) in (a and d) non-transfected (NT) ECs and (b and e) ECs transfected with siRNA-CTRL or (c and f) siRNA-STAT3 in the (a–c) absence or (d–f) presence of SB431542 (0.5 μM). Boxes depicted in (a–f) indicate the magnification shown in (a'–f'), respectively. Nuclei were stained using 4',6-diamidino-2-phenylindole (DAPI). Scale bar represents 30 μm. (g and h) Densitometric analyses of the experiments shown in (a–f). Data were expressed as the normalized ratio of smad4 signal (pixels/μm2) in the nucleus/cytosol (N=5). The number of analyzed cells is depicted in bars. Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis), followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d. (i–p) Nuclear- and cytosol-rich fractions were obtained from NT ECs and from ECs transfected with siRNA-CTRL or siRNA-TRPM4, and smad4 expression was analyzed. (i–o) Representative images from western blot experiments performed to detect smad4 in the nuclear- and cytosol-rich fractions from NT ECs and ECs transfected with siRNA-CTRL or siRNA-TRPM4. (j–p) Densitometric analyses of the experiments shown in (i–o), respectively. Protein levels were normalized against histone H3 or tubulin for nuclear- and cytosol-rich fractions, respectively, and the data are expressed relative to the NT condition (N=5). Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis) followed by Dunn's post hoc test. **P<0.01. Graph bars show the mean±s.d.
Suppressing STAT3 Expression Induces Endothelial Hyperpermeability
To assess if STAT3 suppression induced endothelial fibrosis produces functional alterations in the endothelium, we were prompted to evaluate the effect of STAT3 inhibition on endothelial barrier function. To this end, FITC-dextran endothelial leakage assays were performed by using a transwell system. RMECs were cultured in the absence or presence of the STAT3 inhibitor, S3I-201. RMECs exposed to S3I-201 showed increased FITC-dextran endothelial leakage (Figure 11). Similar results were obtained in EA cells exposed to S3I-201 (Supplementary Figure S8). These findings suggest that the inhibition of STAT3 activity participates in the increase of endothelial permeability, which is concordant with STAT3 suppression and inhibition inducing decreased VE-cadherin and CD31 expression.
Inhibition of STAT3 activity increased endothelial permeability. RMEC cells were cultured in the absence or presence of the STAT3 inhibitor, S3I-201, for 72 h, and then, Fluorescein isothiocyanateFITC-dextran endothelial leakage was measured by using a transwell system. Cells were incubated for 90 min with 0.5 mg/ml FITC-dextran before measurements. Data are expressed as relative barrier function (%) compared to the vehicle-treated group (100%) (N=5). Statistical significance was assessed by a one-way analysis of variance (ANOVA) (Kruskal–Wallis) followed by Dunn's post hoc test. *P<0.05. **P<0.01. Graph bars show the mean±s.d.
Discussion
The conversion of ECs into activated fibroblasts is a highly relevant process to consider due to involvement in endothelial dysfunction during human pathologies. Therefore, understanding the mechanisms underlying endothelial fibrosis is crucial for improving current therapies and developing novel drugs. This study focused on assessing the impact of inhibited STAT3 expression on endothelial fibrosis and the mediating mechanism of the observed effects. The results evidenced that suppressed STAT3 expression promoted a fibrotic-like phenotype in ECs by decreasing the expression of endothelial markers and the acquisition of both fibrotic and ECM proteins. The potential underlying mechanism was found dependent on TGF-β1 secretion, the ALK5 activation pathway, and Smad4 translocation into the nucleus.
STAT3 downregulation is involved in generating fibrosis in the heart and vascular tissue.16, 17, 18 Considering that TGF-β1 is the most studied fibrosis inducer in ECs, these prior findings are in line with the present results that recorded TGF-β secretion as induced by the expressional suppression of STAT3. Interestingly, STAT3 can inhibit TGF-β1 actions by interacting with smad3, thereby decreasing TGF-β-mediated transcriptional responses, including epithelial-to-mesenchymal transition.25 The present results support this, evidencing that STAT3 downregulation-induced changes in the expression of proteins concomitant with EndMT. Furthermore, TGF-β-induced epithelial-to-mesenchymal transition is enhanced through JAK/STAT3 pathway via the upregulation of phosphorylated Smad3 and Snail.31, 32 Snail is a zinc-finger-containing transcription factor that induces epithelial-to-mesenchymal transition and EndMT. The currently presented data showed that the downregulation of STAT3 induced phosphorylated Smad3 translocation, a process coinciding with the development of EndMT. Interestingly, medium extracted from cardiac cells derived from tissue-specific STAT3-KO mice evidenced decreased EC proliferation.17 The authors hypothesized that the presence of paracrine factors attenuated angiogenesis. This conjecture is concordant with the present results, which recorded decreased VE-cadherin and CD31 expressions in ECs when STAT3 was downregulated. EndMT is highly specific to ECs as this process is based on the expressional changes of endothelial-specific proteins, such as VE-cadherin, and endothelial progenitor markers.5, 9, 33, 34, 35, 36 In addition to this, epithelial-to-mesenchymal transition is widely reported as specific to epithelial cells. To our knowledge, no study has provided strong evidence for EndMT occurring in vascular smooth muscle cells. While skeletal muscle fibrosis has been well reported,37, 38, 39, 40, 41 fibrosis in vascular smooth muscle cells is virtually unreported.42
Studies support that STAT3 cooperates with TGF-β1 to enhance fibrosis.26, 27 In particular, STAT3 activation triggers/upregulates the fibrotic actions of TGF-β1 in the liver.26 Related to pathologies, IL-6 participates in STAT3-mediated TGF-β1 production, especially in the context of cancer.43, 44, 45 Considering the available literature and the presently obtained results, it appears that the down- or upregulation of STAT3 generates TGF-β1-induced fibrosis in a tissue-specific manner. Therefore, further experiments are needed to evaluate these contrasting results.
Inflammation mediators are also inducers of endothelial fibrosis, suggesting that these molecules can generate fibrotic conversion during inflammatory processes. Interestingly, STAT3 participates in the inflammatory process in several tissues, including in lung inflammation,32, 46 smoke-induced airway injury,47 colonic cells,48 the brain49 and vascular blood vessels,50 among other tissues. In this context, some researchers suggest that STAT3 is needed to suppress inflammation.16, 51 Notably, STAT3 inhibits nuclear factor-kappa B (NF-κB) transcription factor, which consequently results in the expressional inhibition of genes supporting the anti-inflammatory actions of STAT3.52, 53 The subsequent downregulation of STAT3 would translate into an unopposed, inflammatory-induced fibrotic process. Further regarding STAT3, signaling can function through the canonical Smad family pathway as well as through a noncanonical intracellular pathway that activates the NF-κB.54, 55 This suggests that STAT3-dependent NF-κB inhibition is determined by the use of the canonical or noncanonical intracellular pathways, with pathway determination probably dependent on both the type of ligand and stimulus intensity. However, more precise and complex experiments must be carried out to analyze this subject. Similarly, expression of the suppressor of cytokine signaling 3 (SOCS3), a well-known inducible negative regulator of the STAT3 signaling pathway, promotes inflammation while suppressor knockdown inhibits the inflammatory process.56 Interestingly, SOCS3 is associated with increased inflammation through the overexpression of inflammation mediators, such as TNF-β, and hormones, such as growth factors.46, 56
During moderate inflammation, proinflammatory cytokines trigger the phosphorylation of endothelial STAT3 in Y705 and S727. Specifically, S727 phosphorylation is slightly enhanced. This precise STAT3 activation state promotes a gene profile expression that stimulates actions in ECs. However, the inflammatory state (e.g. high vs low) conditionally generates either protective or detrimental inflammatory actions in ECs, as dependent on the relative levels of inflammation mediators and STAT3 activation.50, 57, 58, 59 Therefore, STAT3 evidently exercises either pro- or anti-inflammatory actions depending on its regulation, as well as on tissue and cell types. However, the fibrotic effects of STAT3 are not yet fully understood, and more studies are needed to further elucidate this issue.
In conclusion, the present results indicate that the suppression of STAT3 expression in ECs induces conversion into activated fibroblasts through the involvement of the TGF-β1/ALK5/Smad4 signaling pathway, thus describing a potential mechanism for endothelial dysfunction during systemic or local inflammatory diseases.
References
Mangge H, Hubmann H, Pilz S et al. Beyond cholesterol – inflammatory cytokines, the key mediators in atherosclerosis. Clin Chem Lab Med 2004;42:467–474.
Zhang C . The role of inflammatory cytokines in endothelial dysfunction. Basic Res Cardiol 2008;103:398–406.
Sprague AH, Khalil RA . Inflammatory cytokines in vascular dysfunction and vascular disease. Biochem Pharmacol 2009;78:539–552.
DeBoer MD . Obesity, systemic inflammation, and increased risk for cardiovascular disease and diabetes among adolescents: a need for screening tools to target interventions. Nutrition 2013;29:379–386.
Zeisberg EM, Potenta S, Xie L et al. Discovery of endothelial to mesenchymal transition as a source for carcinoma-associated fibroblasts. Cancer Res 2007;67:10123–10128.
Maleszewska M, Moonen J-RAJ, Huijkman N et al. IL-1β and TGFβ2 synergistically induce endothelial to mesenchymal transition in an NFκB-dependent manner. Immunobiology 2013;218:443–454.
Mahler GJ, Farrar EJ, Butcher JT . Inflammatory cytokines promote mesenchymal transformation in embryonic and adult valve endothelial cells. Arterioscler Thromb Vasc Biol 2012;33:121–130.
Pérez L, Muñoz-Durango N, Riedel CA et al. Endothelial-to-mesenchymal transition: cytokine-mediated pathways that determine endothelial fibrosis under inflammatory conditions. Cytokine Growth Factor Rev 2017;33:41–54.
Echeverría C, Montorfano I, Sarmiento D et al. Lipopolysaccharide induces a fibrotic-like phenotype in endothelial cells. J Cell Mol Med 2013;17:800–814.
Zeisberg EM, Tarnavski O, Zeisberg M et al. Endothelial-to-mesenchymal transition contributes to cardiac fibrosis. Nat Med 2007;13:952–961.
Montorfano I, Becerra A, Cerro R et al. Oxidative stress mediates the conversion of endothelial cells into myofibroblasts via a TGF-β1 and TGF-β2-dependent pathway. Lab Invest 2014;94:1068–1082.
Echeverría C, Montorfano I et al. Endotoxin-induced endothelial fibrosis is dependent on expression of transforming growth factors β1 and β2. Infect Immun 2014;82:3678–3686.
Echeverría C, Montorfano I, Hermosilla T et al. Endotoxin induces fibrosis in vascular endothelial cells through a mechanism dependent on transient receptor protein melastatin 7 activity. PLoS ONE 2014;9:e94146.
van Meeteren LA, ten Dijke P . Regulation of endothelial cell plasticity by TGF-β. Cell Tissue Res 2012;347:177–186.
Santibañez JF, Quintanilla M, Bernabeu C . TGF-β/TGF-β receptor system and its role in physiological and pathological conditions. Clin Sci 2011;121:233–251.
Jacoby JJ, Kalinowski A, Liu M-G et al. Cardiomyocyte-restricted knockout of STAT3 results in higher sensitivity to inflammation, cardiac fibrosis, and heart failure with advanced age. Proc Natl Acad Sci USA 2003;100:12929–12934.
Hilfiker-Kleiner D, Hilfiker A, Fuchs M et al. Signal transducer and activator of transcription 3 is required for myocardial capillary growth, control of interstitial matrix deposition, and heart protection from ischemic injury. Circ Res 2004;95:187–195.
Boengler K, Buechert A, Heinen Y et al. Cardioprotection by ischemic postconditioning is lost in aged and STAT3-deficient mice. Circ Res 2008;102:131–135.
Liu Y, Liu H, Meyer C et al. Transforming growth factor-beta (TGF-β)-mediated connective tissue growth factor (CTGF) expression in hepatic stellate cells requires Stat3 signaling activation. J Biol Chem 2013;288:30708–30719.
Yu Y, Wang Y, Niu Y et al. Leukemia inhibitory factor attenuates renal fibrosis through Stat3-miR-29c. Am J Physiol Renal Physiol 2015;309:F595–F603.
Tang J, Liu C-Y, Lu M-M et al. Fluorofenidone protects against renal fibrosis by inhibiting STAT3 tyrosine phosphorylation. Mol Cell Biochem 2015;407:77–87.
Prêle CM, Yao E, O'Donoghue RJJ et al. STAT3: a central mediator of pulmonary fibrosis? Proc Am Thorac Soc 2012;9:177–182.
Kim JE, Patel M, Ruzevick J et al. STAT3 Activation in glioblastoma: biochemical and therapeutic implications. Cancers (Basel) 2014;6:376–395.
Marco Demaria VP . Pro-malignant properties of STAT3 during chronic inflammation. Oncotarget 2012;3:359–360.
Wang G, Yu Y, Sun C et al. STAT3 selectively interacts with Smad3 to antagonize TGF-β. Oncogene 2015;35:4388–4398.
Xu M-Y, Hu J-J, Shen J et al. Stat3 signaling activation crosslinking of TGF-β1 in hepatic stellate cell exacerbates liver injury and fibrosis. Biochim Biophys Acta 2014;1842:2237–2245.
O'Reilly S, Ciechomska M, Cant R et al. Interleukin-6 (IL-6) trans signaling drives a STAT3-dependent pathway that leads to hyperactive transforming growth factor-beta (TGF-β) signaling promoting SMAD3 activation and fibrosis via gremlin protein. J Biol Chem 2014;289:9952–9960.
Edgell CJ, McDonald CC, Graham JB . Permanent cell line expressing human factor VIII-related antigen established by hybridization. Proc Natl Acad Sci USA 1983;80:3734–3737.
Potenta S, Zeisberg E, Kalluri R . The role of endothelial-to-mesenchymal transition in cancer progression. Br J Cancer 2008;99:1375–1379.
Echeverría C, Montorfano I, Cabello-Verrugio C et al. Suppression of transient receptor potential melastatin 4 expression promotes conversion of endothelial cells into fibroblasts via transforming growth factor/activin receptor-like kinase 5 pathway. J Hypertens 2015;33:981–992.
Saitoh M, Endo K, Furuya S et al. STAT3 integrates cooperative Ras and TGF-β signals that induce Snail expression. Oncogene 2016;35:1049–1057.
Liu R-Y, Zeng Y, Lei Z et al. JAK/STAT3 signaling is required for TGF-beta-induced epithelial-mesenchymal transition in lung cancer cells. Int J Oncol 2014;44:1643–1651.
Garcia J, Sandi MJ, Cordelier P et al. Tie1 deficiency induces endothelial–mesenchymal transition. EMBO Rep 2012;13:431–439.
Ghiabi P, Jiang J, Pasquier J et al. Breast cancer cells promote a notch-dependent mesenchymal phenotype in endothelial cells participating to a pro-tumoral niche. J Transl Med 2015;13:27.
Medici D, Kalluri R . Endothelial-mesenchymal transition and its contribution to the emergence of stem cell phenotype. Semin Cancer Biol 2012;22:379–384.
Medici D, Shore EM, Lounev VY et al. Conversion of vascular endothelial cells into multipotent stem-like cells. Nat Med 2010;16:1400–1406.
Chapman MA, Mukund K, Subramaniam S et al. Three distinct cell populations express extracellular matrix proteins and increase in number during skeletal muscle fibrosis. Am J Physiol Cell Physiol 2017;312:C131–C143.
Lieber RL, Ward SR . Cellular mechanisms of tissue fibrosis. 4. Structural and functional consequences of skeletal muscle fibrosis. Am J Physiol Cell Physiol 2013;305:C241–C252.
Li ZB, Kollias HD, Wagner KR . Myostatin directly regulates skeletal muscle fibrosis. J Biol Chem 2008;283:19371–19378.
Zhu J, Li Y, Shen W et al. Relationships between transforming growth factor-beta1, myostatin, and decorin: implications for skeletal muscle fibrosis. J Biol Chem 2007;282:25852–25863.
Grefte S, Kuijpers-Jagtman AM, Torensma R et al. Skeletal muscle fibrosis: the effect of stromal-derived factor-1α-loaded collagen scaffolds. Regen Med 2010;5:737–747.
Krishna CV, Singh J, Thangavel C et al. Role of microRNAs in gastrointestinal smooth muscle fibrosis and dysfunction: novel molecular perspectives on the pathophysiology and therapeutic targeting. Am J Physiol Gastrointest Liver Physiol 2016;310:G449–G459.
Zhu Q, Zhang X, Zhang L et al. The IL-6-STAT3 axis mediates a reciprocal crosstalk between cancer-derived mesenchymal stem cells and neutrophils to synergistically prompt gastric cancer progression. Cell Death Dis 2014;5:e1295.
Zhao G, Zhu G, Huang Y et al. IL-6 mediates the signal pathway of JAK-STAT3-VEGF-C promoting growth, invasion and lymphangiogenesis in gastric cancer. Oncol Rep 2016;35:1787–1795.
Cheng J-T, Deng Y-N, Yi H-M et al. Hepatic carcinoma-associated fibroblasts induce IDO-producing regulatory dendritic cells through IL-6-mediated STAT3 activation. Oncogenesis 2016;5:e198.
Zhang S, Hwaiz R, Luo L et al. STAT3-dependent CXC chemokine formation and neutrophil migration in streptococcal M1 protein-induced acute lung inflammation. Am J Physiol Lung Cell Mol Physiol 2015;308:L1159–L1167.
Geraghty P, Wyman AE, Garcia-Arcos I et al. STAT3 modulates cigarette smoke-induced inflammation and protease expression. Front Physiol 2013;4:267.
Nguyen AV, Wu Y-Y, Liu Q et al. STAT3 in epithelial cells regulates inflammation and tumor progression to malignant state in colon. Neoplasia 2013;15:998–1008.
Chen E, Xu D, Lan X et al. A novel role of the STAT3 pathway in brain inflammation-induced human neural progenitor cell differentiation. Curr Mol Med 2013;13:1474–1484.
Kurdi M, Booz GW . Deciphering STAT3 signaling in the heart: plasticity and vascular inflammation. Congest Heart Fail 2010;16:234–238.
Yang XO, Panopoulos AD, Nurieva R et al. STAT3 regulates cytokine-mediated generation of inflammatory helper T cells. J Biol Chem 2007;282:9358–9363.
Yu ZY, Zhang WZ, Kone BC . Signal transducers and activators of transcription 3 (STAT3) inhibits transcription of the inducible nitric oxide synthase gene by interacting with nuclear factor kappa B. Biochem J 2002;367:97–105.
He G, Karin M . NF-kB and STAT3 key players in liver inflammation and cancer. Cell Res 2011;21:159–168.
Freudlsperger C, Bian Y, Wise SC et al. TGF-β and NF-kB signal pathway cross-talk is mediated through TAK1 and SMAD7 in a subset of head and neck cancers. Oncogene 2012;32:1549–1559.
Ishinaga H, Jono H, Lim JH et al. Synergistic induction of nuclear factor-kappa B by transforming growth factor-beta and tumour necrosis factor-alpha is mediated by protein kinase A-dependent RelA acetylation. Biochem J 2009;417:583–591.
Liu Z, Gan L, Zhou Z et al. SOCS3 promotes inflammation and apoptosis via inhibiting JAK2/STAT3 signaling pathway in 3T3-L1 adipocyte. Immunobiology 2015;220:947–953.
Wang M, Zhang W, Crisostomo P et al. Endothelial STAT3 plays a critical role in generalized myocardial proinflammatory and proapoptotic signaling. Am J Physiol Heart Circ Physiol 2007;293:H2101–H2108.
Kurdi M, Booz GW . Can the protective actions of JAK-STAT in the heart be exploited therapeutically? Parsing the regulation of interleukin-6-type cytokine signaling. J Cardiovasc Pharmacol 2007;50:126–141.
Fan Y, Mao R, Yang J . NF-κB and STAT3 signaling pathways collaboratively link inflammation to cancer. Protein Cell 2013;4:176–185.
Acknowledgements
This work was supported by research grants from Fondo Nacional de Desarrollo Científico y Tecnológico—Fondecyt 1161288 (to FS) and 1161646 (to CCV), CONICYT grant for PhD students 21120399 (to AB) and 21130516 (to LP), Millennium Institute on Immunology and Immunotherapy P09-016-F (to FS, CCV), Association-Francaise Contre Les Myopathies AFM 16670 (to CCV), and UNAB DI-741-15/N (to FS, CCV).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Inflammatory mediators change endothelial cells (ECs) into activated fibroblasts via TGF-β1-meditaed epithelial-to-mesenchymal transition. Although decreased STAT3 can induce TGF-β1-mediated fibrosis, it is not known whether suppression of STAT3 expression induces EC fibrosis. This study shows that suppressed expression of STAT3 stimulates fibrotic conversion in ECs via the TGF-β1/ALK5/Smad4 signaling pathway.
Supplementary information
Rights and permissions
About this article
Cite this article
Becerra, A., Rojas, M., Vallejos, A. et al. Endothelial fibrosis induced by suppressed STAT3 expression mediated by signaling involving the TGF-β1/ALK5/Smad pathway. Lab Invest 97, 1033–1046 (2017). https://doi.org/10.1038/labinvest.2017.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2017.61
This article is cited by
-
Idiopathic Pulmonary Fibrosis Serum proteomic analysis before and after nintedanib therapy
Scientific Reports (2020)
-
OxHDL controls LOX-1 expression and plasma membrane localization through a mechanism dependent on NOX/ROS/NF-κB pathway on endothelial cells
Laboratory Investigation (2019)
-
Endotoxemia-induced endothelial fibrosis inhibition improves hypotension, tachycardia, multiple organ dysfunction syndrome, cytokine response, oxidative stress, and survival
Laboratory Investigation (2019)
-
Interleukin-6 determines protein stabilization of DNA methyltransferases and alters DNA promoter methylation of genes associated with insulin signaling and angiogenesis
Laboratory Investigation (2018)