Abstract
Many reports have elucidated the mechanisms and consequences of bacterial quorum sensing (QS), a molecular communication system by which bacterial cells enumerate their cell density and organize collective behavior. In few cases, however, the numbers of bacteria exhibiting this collective behavior have been reported, either as a number concentration or a fraction of the whole. Not all cells in the population, for example, take on the collective phenotype. Thus, the specific attribution of the postulated benefit can remain obscure. This is partly due to our inability to independently assemble a defined quorum, for natural and most artificial systems the quorum itself is a consequence of the biological context (niche and signaling mechanisms). Here, we describe the intentional assembly of quantized quorums. These are made possible by independently engineering the autoinducer signal transduction cascade of Escherichia coli (E. coli) and the sensitivity of detector cells so that upon encountering a particular autoinducer level, a discretized sub-population of cells emerges with the desired phenotype. In our case, the emergent cells all express an equivalent amount of marker protein, DsRed, as an indicator of a specific QS-mediated activity. The process is robust, as detector cells are engineered to target both large and small quorums. The process takes about 6 h, irrespective of quorum level. We demonstrate sensitive detection of autoinducer-2 (AI-2) as an application stemming from quantized quorums. We then demonstrate sub-population partitioning in that AI-2-secreting cells can ‘call’ groups neighboring cells that ‘travel’ and establish a QS-mediated phenotype upon reaching the new locale.
Similar content being viewed by others
Introduction
Bacteria, once thought to be the archetypal minimal autonomously operating living units, are now known to utilize an intricate communication system for sensing and interpreting self-generated molecular cues that coordinate far more expansive population-based behavior. Referred to as quorum sensing (QS), population ‘awareness’ and control are mediated by the production, release and detection of small chemical signaling molecules or ‘autoinducers’. As autoinducer concentration increases with the number of bacteria growing within a particular niche, a threshold is reached signaling and coordinating processes of the population (Ulitzur et al., 1976; Fuqua et al., 1994; Fuqua and Greenberg, 1998). Bacteria ‘self-report’ their number and modulate their behavior as a collective unit, although more recent analysis suggests they more appropriately report on the level of the signal molecule itself and this, in turn, is a surrogate for the number of cells in a quiescent niche (Hense et al., 2007). Interestingly, the exact fraction of the overall population that represents the quorum has not been elucidated and indeed should be a product of the particular niche (for example, local exposure to threshold autoinducer concentrations), as well as the QS signal transduction motif. The prevailing rationalization is that the bacteria benefit by operating collectively, although the emergence of parsed sub-populations that exhibit altruistic or even aberrant social behavior (that is, cheating on the collective good) has stimulated new hypotheses of sociomicrobiology including the suggestion that microbial QS serves as a surrogate for human sociobiology (Ben-Jacob et al., 2004; Parsek and Greenberg, 2005; Gardner et al., 2007; Davidson and Surette, 2008; Ben-Jacob and Schultz, 2010; Connell et al., 2012; Dandekar et al., 2012; Pradhan and Chatterjee, 2014; Smith et al., 2014). Certainly, among the beneficial phenotypes attributed to QS, which could also be referred to as the ‘currency’ of the public good, are bioluminescence (Bassler et al., 1994), virulence (Zhu et al., 2002), biofilm formation (Balestrino et al., 2005; Gonzalez Barrios et al., 2006; Li et al., 2007), cell division, motility and cooperativity (Dandekar et al., 2012). We note that intentional rewiring these QS-regulated systems also benefits biotechnological applications, if not the microbes themselves (Fernandes et al., 2010; Tsao et al., 2010; Wu et al., 2013; Swofford et al., 2015; Thompson et al., 2015). It is therefore intriguing to consider what is the particular quorum and why was it reached. That is, a particular subgroup may be needed that is not the collective whole. What is the benefit when say, 65% of the population elicits a coordinated response? Or is benefit maximized at another quantum unit, say 55%? Such questions are not easily answered using current methodologies as the quorums themselves are generated by the bacteria in response to their particular context. One cannot be separated from the other.
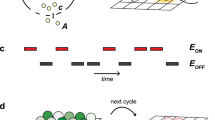
We sought to alter this scenario for the ‘programmed’ assembly of a bacterial quorum. In this work, we generate a stable quorum wherein a specific fraction of the whole exhibits the desired collective behavior. A particular sub-population may be ‘assigned’ a particular function that should be executed. For this, we developed an autoinducer signal amplification vector that overrides the native signal transduction process so that upon detection of autoinducer, the individual bacterial cell overproduces a fluorescent marker. Then, all QS responding cells rapidly synthesize similar levels of marker protein. The initial detection of the signal is directly transmitted to the population and the number of detecting cells is manifest into a controlled or quantized quorum (Figure 1a). In order to establish robustness of this methodology, cells were further engineered for sensitivity to autoinducer. A more sensitive cell is used to create a smaller quorum and a less sensitive cell is used to create a larger quorum. Thus, there are two elements to this strategy, the vector-based engineered signal transduction unit provides uniformity among quantized quorums and the cell-based detector system provides the magnitude of the quantized fractions. The combination of autoinducer concentration and genetic background enables tailored assembly of a bacterial quorum.
(a) Transition to quantized ‘quorum’ by the addition of varied levels of AI-2. Three levels (10%, 30% and 70%) are depicted. (b) Schematic of AI-2-mediated QS in E. coli. AI-2 is synthesized via two terminal synthases, Pfs and LuxS. It is secreted by YdgG (TqsA) and taken up by the Lsr transporter (LsrACDB). Following phosphorylation and binding to LsrR, phospho-AI-2 initiates the transcription of T7 polymerase (pCT6), which in turn, initiates the overexpression of phenotypic marker, DsRed. In addition to wild-type W3110ΔluxS, AI-2 hypersensitive strain CT104 (W3110ΔlsrFGΔluxS) was used. CT104 accumulates AI-2-P enabling detection of extracellular AI-2 at lower levels than the wild-type cells.
Although there are many species-specific autoinducers, one signal molecule, autoinducer-2 (AI-2), is produced by scores of bacteria and as such, mediates interspecies communication, indicating its potential as an indicator for the existence of a variety of species, including in planktonic conditions or as sequestered in biofilms (Federle and Bassler, 2003). Moreover, its signal transduction cascade is evolutionarily extensive and mechanistically diverse (Quan and Bentley, 2012). Hence, one’s ability to conceive of and engineer a system for a particular application is substantial when based on AI-2.
AI-2 is synthesized via LuxS and Pfs, the two terminal synthases yielding both AI-2 and homocysteine of the activated methyl cycle and exported from the cell via TqsA (Herzberg et al., 2006) (Figure 1b). AI-2 is imported (via LsrACDB), phosphorylated (via LsrK), detected (via LsrR) and processed (via LsrFG) by components of the Lsr operon (DeLisa et al., 2001c; Wang et al., 2005b; Xavier and Bassler, 2005). To establish this methodology, one must detect AI-2 and transduce its signaling function into a desired quantized phenotype. Further, in order to separate a self-induced quorum from an intentionally assembled quorum, the cell population must not endogenously synthesize AI-2. Hence, we constructed luxS mutants as hosts for this study. We note in wild-type cells, the synthesis of AI-2 is a direct consequence of carbon metabolism and is directly proportional to the growth rate of cells when grown either in batch or steady continuous cultures (DeLisa et al., 2001b). Hence, its concentration increases concomitantly with cell growth and this enables enumeration of bacterial density. We have also created a luxS- signal amplification methodology wherein the native AI-2 signal transduction process (lsr operon), which is a relatively weak initiator of gene expression, signals the expression of bacteriophage T7 polymerase, which when expressed, mediates expression from a second promoter (pT7). The addition of T7 polymerase, therefore, acts as an amplifier of the native lsr-mediated gene expression. Here, T7 polymerase is used to initiate the expression of DsRed, which, in turn, is used to define the quantized quorum. Note, although DsRed is a marker, its expression characteristics mimic those of other heterologous proteins produced from the same plasmid system (Tsao et al., 2010). That is, DsRed serves as both an indicator of a quorum and as a targeted phenotype of the assembled quorum. By analogy, therefore, any desired phenotype that is mediated by lsr gene expression is expected to follow similar trajectories. Perhaps too, QS-mediated phenotypes that have not been attributed to lsr-control but are a consequence of AI-2 signaling can be viewed in a similar light.
In order to modulate the size of the assembled quorum, we used two strains. The first is Escherichia coli (E. coli) W3110 ΔluxS providing the base case system that most closely reflects the natural system. Then, we engineered a strain that responds to lower levels of AI-2 increasing the sensitivity of the reporter system (Figure 1). To be specific, we deleted genes for the AI-2-P processing enzymes, LsrF and LsrG, yielding cells (E. coli CT104) that should accumulate AI-2-P the active AI-2 species providing derepression of the Lsr operon at lower external AI-2 concentrations (Hooshangi and Bentley, 2011; Ha et al., 2013).
Materials and methods
Bacterial growth conditions and plasmid construction.
Plasmids and strains used in this study are listed in Table 1. Strains were maintained and plasmids were manipulated according to standard procedures and previous reports (Maniatis et al., 1982). Oligonucleotides were from Integrated DNA Technologies (Coralville, IA, USA) and are listed in Table 2. DNA sequencing was performed by Genewiz (South Plainfield, NJ, USA).
Chromosomal deletions of lsrFG and luxS
The one-step replacement method described by Datsenko and Wanner (2000) was for gene deletion. We used the phage_Red recombination system to replace lsrFG with the lsrFG::cat PCR cat cassette. The cat cassette was produced by PCR amplifying pKD3 using Vent DNA polymerase (New England Biolabs, Ipswich, MA, USA) with primers lsrFGHP3 and lsrFGHP2 (Table 1). The blunt end PCR product was introduced by electroporation into E. coli W3110 containing plasmid pKD46, which expresses the Red recombinase, and was cured later by growth at 37 °C. Recombinants were selected on Luria-Bertani (LB) broth supplemented with chloramphenicol, resulting in the strain E. coli W3110 (ΔlsrFG::cat). In order to remove the antibiotic resistance gene, the plasmid 707FLPe (tetr) (Gene Bridges, Heidelberg, Germany) was chemically transferred into E. coli W3110 (ΔlsrFG::cat) using the Z-Competent E. coli Transformation Kit (Zymo Research, Irvine, CA, USA). The expression of the plasmid mediated the excision of the antibiotic resistance gene, and the product strain was named E. coli W3110ΔlsrFG. Deletion of luxS from W3110ΔlsrFG to make the lsrFG and luxS double knockout strain was conducted similarly by PCR amplification of pKD3 with primers luxSHP1 and luxSHP2 (Table 2) (Wang et al., 2005a), and excision of the antibiotic resistance gene was similar to that described above. The product strain, E. coli W3110 (lsrFG-, luxS-), was named E. coli W3110ΔlsrFGΔluxS. Deletion of luxS from E. coli W3110 was performed in the same manner, generating a strain named E. coli W3110ΔluxS. E.coli BL21 (luxS-) was made previously (Luo et al., 2012). All primers used for PCR verification of gene deletions here are listed in Table 2.
Transformation of cells
Electrocompetent cells were created from overnight cultures diluted 1:50 into fresh LB broth with appropriate antibiotics. Cells were incubated in a shaking incubator at 37 oC. Upon reaching mid-log phase, the cells were washed twice with ice-cold ultrapure water and twice with ice-cold 10% glycerol. In all, 1 μl of DNA and 35 μl of cells were combined in a 1 mm electroporation cuvette. Electroporation was performed using a BioRad MicroPulser (Hercules, CA, USA) on the EC-1 setting. The cells were recovered for 1 h before plating on LB agar with appropriate antibiotics (Maniatis et al., 1982).
AI-2 synthesis
In order to establish signaling exogenously, AI-2 was synthesized in vitro according to previous reports (Maniatis et al., 1982; Tavender et al., 2008; Tsao et al., 2010). Briefly, His6-Pfs and His6-LuxS were produced by induction of E. coli BL21 (pTrcHis-pfs) and BL21 (pTrcHis-luxS) with 1 mM isopropyl-beta-D-1-thiogalactopyranoside (IPTG) during mid-log growth at 37 °C. After 4–6 h of induction with IPTG, the cells were lysed using BugBuster solution (EMD Millipore, Billerica, MA, USA), and His6-Pfs and His6-LuxS were purified with Co2+ affinity resin (BD TALON, BD Biosciences, San Jose, CA, USA). The purified enzymes were eluted in 125 mM imidazole in phosphate buffer (pH=7.4) and combined to synthesize AI-2 by addition with 1 mM S-adenosylhomocysteine in 50 mM Tris–HCl (pH=7.8) at 37 °C for 4 h (Maniatis et al., 1982; Tavender et al., 2008; Tsao et al., 2010). The enzymatic reaction products were twice extracted by chloroform and recovered from the aqueous phase and used to make a 488 μM stock solution. AI-2 concentration was quantified using Ellman’s reagent, as has been described previously (Schauder et al., 2001; Winzer et al., 2002).
AI-2 detection and quorum quantification
Overnight cultures, A600~2.00, were diluted 1:50 into fresh LB broth with appropriate antibiotics and incubated for 1 h at 37 oC in a shaking incubator (New Brunswick Scientific Incubator Shaker Series I26, New Brunswick, NJ, USA) at 250 relative centrifugal force (RCF). In all, 374 μl of the diluted culture was transferred into a single well of a 24-well plate in triplicate for each dilution. Serial dilutions of AI-2 were made in triplicate. In total, 26 μl of each dilution was added to each well to achieve the specified concentrations. The plates were incubated at 37 °C in a shaking incubator (New Brunswick Scientific Incubator Shaker Series I26) at 250 RCF. In order to prevent evaporation of the samples, two 2 l beakers with 500 ml of water were placed in the shaker to act as humidifiers.
Flow cytometric analysis was performed to measure cell fluorescence, which correlates to production of DsRed fluorescent protein, using a FACSCanto II flow cytometer (BD Biosciences). Every hour, 10 μl of cells were pulled from each well and mixed with 250 μl of FACS Flow for analysis. Each culture was run in triplicate. The blue diode (488 nm) was used as the excitation source and DsRed fluorescence using the 585/40 nm filter. All raw data were further analyzed using BD FACSDiva 6.0 software (BD Biosciences). All cell samples were gated identically after initial baseline data were obtained (Supplementary Information,Supplementary Figure S1).
Development of a configuration for AI-2-mediated cell recruiting
Plasmid, pT5RT7G, was constructed for dual color expression. First, dsredexpress2 was placed in a T5-regulated cassette. The first lac operator in the promoter region was mutated to disable LacI repression at the mutated site (Betz et al., 1986); then the cassette was added to a pET vector containing egfp under T7-regulation. E. coli CT104+pCT6 was transformed with pT5RT7G to generate an AI-2-reporting strain with constitutive red fluorescence and AI-2-responsive green fluorescence.
Recruiter capsules were developed largely as previously shown (Gupta et al., 2013). Cells (BL21 or BL21ΔluxS) were grown to mid exponential phase and concentrated 30-fold by centrifugation and resuspension in LB broth. The resuspended cells were mixed with an equal volume of alginate (2% in 0.15 M NaCl), then added dropwise to a CaCl solution (0.5 M) using a 16-gauge needle. These ‘beads’ were allowed to ‘set’ for 10 min forming capsule cores containing entrapped cells. Between consecutive rinses in NaCl (0.15 M), the cores were transferred alternately into solutions of 0.5% chitosan and 0.5% alginate three times to create stabilizing outer shell layers. Three capsules were added to the bottom well of a transwell system and were confined to the center of each. A transwell membrane (1 μm pore size, BD Biosciences) was inserted. LB medium was diluted 10-fold in PBS (10 mM), supplemented with CaCl (0.5 mM, to maintain capsule integrity), and used as the incubation medium; three volumes were added to the bottom well per volume in the top of the transwell insert. Finally, E. coli CT104+pCT6+pT5RT7G, grown at room temperature to OD 0.1, were diluted 1000-fold into the top of the transwell apparatus. The system was sealed with Parafilm M (Bemis, Oshkosh, WI, USA) and incubated for 16 h or longer.
Results
Signal transduction—expression of DsRed on per cell basis
Plasmids pCT6 and pET200/DsRed (Yung et al., 2006; Tsao et al., 2010) were sequentially transformed into the E. coli W3110 luxS deletion mutant, W3110ΔluxS, and grown in LB media with appropriate antibiotics. Plasmid pCT6 senses and responds to the presence of phosphorylated AI-2 by the expression of T7 RNA polymerase, which, in turn, activates T7 polymerase-mediated overexpression of DsRed from plasmid pET200/DsRed. As this detector strain is luxS deficient, it produces DsRed only in response to exogenously added AI-2.
We also enhanced the detecting cell sensitivity by deletion of two enzymes which process/degrade phosphorylated AI-2. These cells putatively accumulate the QS autoinducer in its active phosphorylated form so that less autoinducer is needed in the environment to stimulate expression from the Lsr promoter. Specifically, lsrFG genes were deleted from the wild-type E. coli W3110, creating the strain W3110ΔlsrFG. The luxS gene from strain E. coli W3110ΔlsrFG was subsequently deleted, creating AI-2 hypersensitive strain E. coli W3110ΔlsrFGΔLuxS or E. coli CT104.
Our first objective was to test the signal transduction process from both the pCT6 amplifier vector and the pET200/DsRed reporter vector. Our intent was to rapidly express DsRed in response to the import of AI-2. Test cultures were monitored hourly and in triplicate commencing addition of exogenously added AI-2 at each of 10 concentrations (from 0 to 32 μM). Fractions of red fluorescing cells and per cell fluorescence levels were subsequently determined for each triplicate sample. The per cell fluorescence for three of the highest concentrations (2, 8 and 32 μM) are depicted over all times in Figure 2a. First, we found no significant differences in the average fluorescence levels among both strains and spanning all conditions. That is, in Figure 2a, we depict both the average and the standard deviation of the per cell fluorescence. Interestingly, the earliest time points exhibited the largest deviation from the mean and this deviation shrunk over time; in essence, distribution became more ‘focused’ as time ensued (in Figure 2a, the standard deviations are indicated by the dark red and blue bars and the magnitudes of these typically decreased over time). The variance in the fluorescence measurements decreased from the initial conditions to when the quorum emerged. By Student’s t-test (see Supplemental section) and replicate average of the means (depicted), there were no statistically significant differences among the per cell fluorescence data of cultures for all concentrations and over all times. That is, once the cells commenced expressing DsRed, their fluorescence level on a per cell basis was identical, especially after the first 4–5 h, as indicated by the variance data in Figure 2a.
(a) Specific fluorescence levels over time for 2, 8 and 32 μM AI-2 addition. The dotted horizontal line represents the average of all data sets, the band represents the overall average of standard deviation for the entire ensemble. Each fluorescence level is accompanied by an additional bar representing the standard deviation of the mean for that particular data set. (b) Induction of DsRed in response to varying concentrations of exogenously added AI-2 to the wild-type W3110ΔluxS strain. Response was measured via fluorescence-activated cell sorting (FACS). The results are displayed as percentage of population fluorescing above background. (c) Induction of DsRed in response to varying concentrations of AI-2 added to the enhanced CT104 strain (W3110ΔlsrFGΔluxS). As in (b), results are displayed as percentage of population fluorescing above background.
Quantized quorum system I
The ability of E. coli W3110 to respond as a population to exogenously added AI-2 was tested by via flow cytometry (Figure 2b). We have included the calculated limit of detection (LOD), which represents the average fluorescing population (% of total) for the zero AI-2 control added to three times its standard deviation (~1.4%). In this way, the LOD is the minimum quantized sub-population for which a fluorescence response can be differentiated from a background signal. Thus, this represents the smallest statistically significant quantized quorum for E. coli W3110ΔluxS in our study. Our results show that statistically significant quorums were achieved only when 2 μM AI-2 or more was added (the data from 2 μM were all above the LOD). We found the percentage of the population exhibiting red fluorescence increased in a dose-dependent manner (Figure 2b) particularly above 1 μM. Also, the fraction of fluorescing cells increased monotonically from time 0 until 6 h, when it remained steady for approximately the next 24 h. That is, transients in per cell fluorescence (Figure 2a) and fraction of fluorescing cells (Figure 2b) appeared to stabilize by 5–6 h, when they reached steady or ‘programmed’ values.
Quantized quorum system ii for enhanced sensitivity
Experiments with E. coli CT104 cells were executed identically. The culture dynamics were similar with a significant transient for all AI-2 concentrations over the first 4 h followed by a steady transition to quantized fractions at about 6 h (Figure 2c). These levels remained steady for over an hour and appeared to drop only for the highest concentrations as the cultures progressed to the 24-h experimental limit. As above, a LOD was calculated and is depicted (~1.2%). In this case, statistically valid quorums were obtained for AI-2 levels 0.25 μM and above. We note the LOD was similar to that of the E. coli W3110 cells, as expected. Also, we did observe increased AI-2 sensitivity because of the lsrFG deletion, and consequently, more sensitive quorum assembly at low levels of AI-2. That is, the E. coli CT104 cells appeared to be best to achieve discrete quorums comprising small sub-populations, specifically between 2% and 15%. By contrast, when larger quantized quorums were sought, the W3110 cells were superior. For example, for CT104 there were no appreciable differences between the 16 and 32 μM populations (64% for each at 7 h), whereas this difference in concentration led to a 9% population difference using W3110. For W3110 at 7 h, the assembled quorums were 61% and 70% for 16 and 32 μM, respectively. In sum, the W3110 cells appeared superior for reaching higher quorum levels (for reaching quantized quorums above 50%) and CT104 cells were superior for generating quorums below 20%. Discretization of quantized quorums, therefore, was a consequence of the engineered sensor cell—its sensitivity to the signal and its engineered signal response system. Although strain enhancement lowered the LOD slightly, its effect on the strain’s ability to guide population-wide quorum assembly was dramatic.
Applications and further insights—a call to arms?
By exposing sensing and actuating cells to different levels of AI-2, we found parsed sub-populations of cells could be assembled within 5 h that exhibit phenotypes specifically attributed to lsr-mediated gene expression. We wanted to consider two questions that emerge from these observations. First, are there biotechnological applications that may warrant further analysis? Second, can these concepts provide context to bacterial behaviors observed in their natural ecological niches?
To address the first question, we found the trajectories initiated by exogenous addition of AI-2 were robust—the eventual quorums reached were apparently established early on so that the end results were, in effect, predetermined by the strain and the initial level of AI-2. Thus, a potential application of this system is revealed by simply altering the lens through which the same data are visualized. In Figure 3a, we plot the percentage of fluorescing cells at 5- and 6-h time points for the different levels of added AI-2 (for example, a dose response). In Figure 3b, we transpose the data to illustrate that the percentage of DsRed-producing cells indicates quite accurately, the level of AI-2 added to the media. Hence, the E. coli CT104 cells constructed precisely report the AI-2 level in culture media; they serve as robust sensors of AI-2. Cell-based sensors have been widely used for reporting the activity of autoinducers, including AI-2, in fluids both from natural environments and from laboratory samples (Mok et al., 2003; Taga, 2005; Gantner et al., 2006; Vilchez et al., 2007; Dulla and Lindow, 2008; Zhu and Pei, 2008). Perhaps the most widely used is the Vibrio harveyi BB170 assay of Surette and Bassler (1998). Our work represents one additional method, here utilizing fluorescence-activated cell sorting rather than a luminometer, its use thus far limited to well-controlled environments such those in industrial biotechnology (DeLisa et al., 2001a).
(a) The fraction of fluorescing cells from Figure 2b is depicted for the 5 h (blue) and 6 h (red) time points. This dose-response curve reveals a linear region at or below 8 μM; the dotted line is a linearly regressed best fit correlation. (b) The data from a are transposed and lumped demonstrating cell-based sensing of AI-2. A full color version of this figure is available at the ISME Journal online.
To address the second question and to provide a more natural context, we considered the interplay between autoinducer concentration, the extent to which cells express DsRed and the time over which the phenotypes emerge. We also note that the E. coli CT104 cells took up AI-2 to develop quantized populations; they did not secrete AI-2 to the extracellular milieu. Native production of AI-2 would have confounded interpretation of the previous results. Thus, in a sense, the E. coli CT104 cells are independent bystanders that may ‘report’ on AI-2 presented to them from either the environment or neighboring cells and take on phenotypes (natural or synthetic) based on the prevailing level of AI-2 they encounter. It would be particularly interesting to evaluate the number of responding cells as a fraction of the whole. Recognizing that AI-2 is in fact a chemoattractant (Hegde et al., 2011), we configured a system where cells that are ‘guided’ to AI-2 could be discerned from cells that ‘report’ on AI-2 based on the local concentration and engineered gene expression. That is, we wanted to test a hypothesis that AI-2-producing cells or ‘sentinel’ cells could ‘call’ their neighbors, attracting them to a new locale. Could bacteria that secrete AI-2 recruit other AI-2-sensing cells to their locations and if so, could the recruited cells take on AI-2-mediated QS phenotypes? This scenario evokes consideration of the potentially disparate dynamics of AI-2 diffusion, AI-2-mediated cell motility and AI-2-mediated gene expression, as well as the different length scales involved. Similar concepts were reported previously using a synthetic system for AI-2 production (Wu et al., 2013). In that work, AI-2 signal molecules were generated in situ on the surface of cancer cells by the assembly of its synthases onto epidermal growth factor receptor (EGFR) surface receptors and with supplementation of the AI-2 precursor S-adenosylhomocysteine to the media. E. coli cells, engineered to express a red marker as a response to QS signaling, were placed in proximity to the cancer cells ladened with AI-2 synthases on their EGFR surface receptors. S-adenosylhomocysteine was added to the media and bacterial cells swam to the cancer cells, where, depending on the prevailing AI-2 concentration, they synthesized the red marker (surrogate for a cancer-killing drug). In the Supplemental section of that work, it was reported that the fraction of cells that swam increased with AI-2 concentration much like DsRed expression in Figures 2 and 3 here, and that (i) the response time for motility was quicker than the response time for lsr-mediated gene expression and (ii) that the AI-2 levels apparently needed to initiate motility were less than those needed for gene expression (see Supplementary Figure S2 here for details).
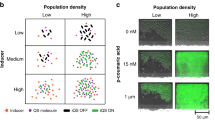
Shown in Figure 4a, we designed cell ‘recruits’ that express fluorescent proteins in response to different cues. CT104 cells were engineered to constitutively express DsRed so they can be detected in the presence of wild-type ‘sentinel’ cells. These ‘recruit’ cells were also engineered to produce green fluorescent protein (GFP) in response to the level of AI-2 encountered, exactly analogously to Figures 1,2,3 (we switched the fluorescent markers from the previous system). Thus, CT104 are tracked by red fluorescence and when encountering a ‘quorum’-producing level of AI-2, they fluoresce green. Non-fluorescing ‘sentinel’ cells were encapsulated within chitosan/alginate capsules (Gupta et al., 2013) in order to provide a centralized source of secreted AI-2 that diffuses into a quiescent buffer. We co-cultured bacteria as depicted in Figure 4a; ‘recruit’ cells were placed in the top of a transwell apparatus and ‘sentinel’ cells were contained within capsules in the bottom compartment. Cultures were maintained at room temperature for long incubation times (to completely encompass time domains of AI-2 diffusion, motility and gene expression over the imposed length scales). The transwell membrane was selected with a 1 μm pore size to restrict passive migration. Additional details of the experimental design are provided in Supplemental section. In Figure 4b(i), the bar graph shows the numbers of red and green cells counted from samples from the lower chamber at two time points for representative experiments. These are denoted the ‘recruited’ fraction. The experiments were carried out multiple times to catch the windows of motility and expression and detailed data analysis was carried out in quadruplicate; images (superimposed images from both red and green filters) are shown in Figure 4b(ii). The intensity profiles (gray value) for red and green fluorophores were plotted for the highlighted portions of the images from luxS+-treated red recruits. Colocalization is observed where strong red and green peaks are overlaid.
AI-2-mediated cell recruiting. (a) Recruitable cells are engineered to express two fluorescent proteins: DsRed is constitutively expressed and eGFP is initiated as in Figure 1, by phosphorylated AI-2, which, through LsrR derepression and amplification is expressed through T7 polymerase. A recruiting test is created by encapsulating AI-2-secreting ‘sentinel’ cells and co-incubating with recruitable (red) cells placed in the upper chamber of a transwell apparatus (porous membrane 1 μM). (b) (i) Bars indicate the recruited red cell count in the lower chamber at 16 h and 40 h. Both AI-2-positive test (luxS+) and AI-2-negative control (luxS-) counts are depicted. Cell counts were performed in quadruplicate by image analysis with standard deviations shown. The green cell count (lsr-mediated gene expression) is also depicted. (ii) Fluoresence images are overlayed (red and green filters) for representative lower chamber samples corresponding to bars in (i). In addition, we have scanned the bracketed regions of the luxS+ images for red and green intensity (plotted as gray value) as a function of horizontal distance (pixels). This depicts coincident red and green identification of cells. Although distinct peaks are revealed for red cells, all green cells were also identified in the red image. A full color version of this figure is available at the ISME Journal online.
The data show stratified AI-2-dependent phenotypic responses stemming from an initially homogeneous cell population over different time scales. Within a 16-h timeframe, AI-2 recruiting (indicated by the presence of AI-2-secreting sentinel cells in the lower chamber) led to 10-fold increase in cell migration across the membrane (Figure 4b(i)). Further, even after 40 h, we found the red recruited cells that had been co-incubated with luxS- sentinels never surpassed the recruited cell count from co-incubations with signal generating luxS+ cells. In addition, by 40 h, a significant sub-population of the recruited cells had fluoresced green (compared with none from cultures with the luxS- sentinels, Figure 4b(ii)). Plotting the fluorescence profiles found in the microscope fields of view for individual and cell clusters, quantitatively reveals the amplitude increase (10-fold in some cases) of green fluorescence, and importantly, the colocalization of both red and green fluorescence where strong peaks are overlaid.
That QS signaling can be utilized to select a sub-population of neighbors to migrate and later undergo a phenotypic shift, are demonstrated in this experiment. Exemplified is the construction of a dynamic, yet artificial, QS system that reveals the readily observable behaviors of chemotaxis and gene expression. The dynamics of the responses, where AI-2-induced chemotaxis preceded GFP expression, are consistent with our previous reports and are likely a determinant of the system as programmed by nature. That is, in Supplementary Figure S2 we show that the motility response is quicker and more sensitive to AI-2 than is the lsr-mediated gene expression. We found this to be both curious and compelling, when viewed in the context of optimization of the public good.
Discussion
Our results show that one can produce an intended or programmed quorum of cells that exhibit a particular trait from within an overall population. We have engineered the AI-2 signal transduction cascade of E. coli to that upon derepression through LsrR, all cells produce a significant and uniform level of DsRed. That is, the original signal, which in native cells would lead to upregulated expression of the transporter, kinase and repressor (Lsr ABDC, LsrK and LsrR, respectively), now provides an additional function in that it leads to upregulation of the T7 polymerase. This, in turn, initiates the overexpression of DsRed from a second plasmid, giving the overall system a significant boost in expression rate beyond that of the native signal (Tsao et al., 2010). We also used luxS mutant cells so that the assembled quorum was due solely to AI-2 exogenously added to the culture media. Finally, we engineered two host detector strains that provide altered sensitivity to the extracellular AI-2 concentration. The combination of the two-plasmid vector system and the particular host cell enables discretization of an emergent population. The best results for establishing a quorum with <15% of the total population used the E. coli CT104 strain (ΔlsrFG). Conversely, the best results for quorums above 50% of the whole were found using W3110.
Interestingly, we found the per cell fluorescence varied the most during the first 4–5 h after inoculation and AI-2 addition (Figure 2a). This time period corresponded to the same period over which the fraction of fluorescing cells increased to a plateau (Figures 2b and c). It is interesting that this time period roughly coincides with the time typically required for the wild-type AI-2-producing cells to reach a quorum in normal batch cultures (DeLisa et al., 2001a, b, c; Wang et al., 2005b). It is also interesting to compare with analogous systems where heterologous protein expression is induced by the addition of exogenous inducers. For example, in work by Siegele and Hu (1997), heterogeneity resulted from an analogous autocatalytic positive feedback inducer uptake system. In their study, arabinose was added to cells at levels yielding an intermediate level of induction of GFP; initially a strong and uniform distribution of GFP-producing cells was obtained but in a small fraction of cells that over time (roughly the same time as the current study) was seen to broaden as more cells became induced, owing in part to the loss of plasmid from sub-populations. Although similar time scales were observed, in the QS system here the distribution of per cell fluorescence was narrowed instead of broadened as in the araBAD system. Perhaps these elements shed light on competing factors at play in the native QS system: the dynamic accumulation of the autoinducer and the phenotypic coordination of the responding cells, both of which are transient and interconnected. For example, we show in Supplementary Figure S3, other AI-2-mediated genes are upregulated along with, but to a lesser extent than, the fluorescent marker. Our data, which suggest that sensor cells respond to AI-2 by initiating DsRed expression at slightly different times and at slightly different rates, point to a 4–5 h ‘focusing’ of the overall population while leading to the quantized quorum. That is, QS can function as a phenotypic focusing system (Wu et al., 2013), as well as a determinant of collective behavior.
We believe the overall process works here because (i) the amplification system is fairly strong and rapid, and (ii) the growing cells rapidly consume the exogenously added AI-2 present in the vicinity of each cell in a particular locale. The former is demonstrated here, whereas the latter is the subject of a comprehensive spatially organized mathematical model currently under development. With this methodology, it would be interesting to develop hypotheses surrounding phenotypes of interest. For example, does antibiotic resistance emerge as a consequence of the size of a quorum? Do bacteria migrate to new locales based on a fraction of seeker cells (Defoirdt, 2011)? We have provided the first suggestion that this may be the case. We developed an observable system in which sentinel cells are non-fluorescing and secrete AI-2. Receiver cells that are always red, chemotax toward AI-2 and eventually, given sufficient AI-2 in the new environs, turn on lsr gene expression. Interestingly, the data are all binned by sub-population. A fraction of the cells, not all, moved to the AI-2. A fraction of the cells, not all, initiated QS-regulated gene expression. Moreover, these fractions were different and the response dynamics were different. We note that a sub-population of cells moved first, then after reaching the new destination, responded by expressing GFP. Whether this occurs in natural niches remains to be seen, but our data suggest this is the case. Note that we have not explored whether stationary cells or those in biofilms may display a similar tendency to be organized into a quantized quorum. That is, does the rate of AI-2 uptake need to be spatially or temporally uniform for behaviors to emerge in a coordinated manner (Danino et al., 2010)? Also, from a biotechnological perspective, can host cells be engineered so that fully 100% of the population expresses recombinant proteins at high level? This is not clear to us for the AI-2 system where autoinducer signal is taken up in response to the QS switch. Even more provocatively, could one create divergent phenotypes for different purposes and could these, in turn, be coordinated? We believe these and other questions may be answered by further exploration of the methodology demonstrated here.
References
Balestrino D, Haagensen JAJ, Rich C, Forestier C . (2005). Characterization of type 2 quorum sensing in Klebsiella pneumoniae and relationship with biofilm formation. J Bacteriol 187: 2870–2880.
Bassler BL, Wright M, Silverman MR . (1994). Multiple signalling systems controlling expression of luminescence in Vibrio harveyi: sequence and function of genes encoding a second sensory pathway. Mol Microbiol 13: 273–286.
Ben-Jacob E, Becker I, Shapira Y, Levine H . (2004). Bacterial linguistic communication and social intelligence. Trends Microbiol 12: 366–372.
Ben-Jacob E, Schultz D . (2010). Bacteria determine fate by playing dice with controlled odds. Proc Natl Acad Sci USA 107: 13197–13198.
Betz JL, Sasmor HM, Buck F, Insley MY, Caruthers MH . (1986). Base substitution mutants of the lac operator: in vivo and in vitro affinities for lac repressor. Gene: Netherlands 50: 123–132.
Connell JL, Whiteley M, Shear JB . (2012). Sociomicrobiology in engineered landscapes. Nat Chem Biol 8: 10–13.
Dandekar AA, Chugani S, Greenberg EP . (2012). Bacterial quorum sensing and metabolic incentives to cooperate. Science 338: 264–266.
Danino T, Mondragon-Palomino O, Tsimring L, Hasty J . (2010). A synchronized quorum of genetic clocks. Nature 463: 326–330.
Datsenko KA, Wanner BL . (2000). One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97: 6640–6645.
Davidson CJ, Surette MG . (2008). Individuality in bacteria. Annu Rev Genet 42: 253–268.
Defoirdt T . (2011). Can bacteria actively search to join groups? ISME J 5: 569–570.
DeLisa MP, Valdes JJ, Bentley WE . (2001a). Quorum signaling via Al-2 communicates the ‘metabolic burden’ associated with heterologous protein production in Escherichia coli. Biotechnol Bioengineer 75: 439–450.
DeLisa MP, Valdes JJ, Bentley WE . (2001b). Mapping stress-induced changes in autoinducer AI-2 production in chemostat-cultivated Escherichia coli K-12. J Bacteriol 183: 2918–2928.
DeLisa MP, Wu CF, Wang L, Valdes JJ, Bentley WE . (2001c). DNA microarray-based identification of genes controlled by autoinducer 2-stimulated quorum sensing in Escherichia coli. J Bacteriol 183: 5239–5247.
Dulla G, Lindow SE . (2008). Quorum size of Pseudomonas syringae is small and dictated by water availability on the leaf surface. Proc Natl Acad Sci USA 105: 3082–3087.
Federle MJ, Bassler BL . (2003). Interspecies communication in bacteria. J Clin Invest 112: 1291–1299.
Fernandes R, Roy V, Wu HC, Bentley WE . (2010). Engineered biological nanofactories trigger quorum sensing response in targeted bacteria. Nat Nanotechnol 5: 213–217.
Fuqua C, Greenberg EP . (1998). Self perception in bacteria: quorum sensing with acylated homoserine lactones. Curr Opin Microbiol 1: 183–189.
Fuqua WC, Winans SC, Greenberg EP . (1994). Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J Bacteriol 176: 269–275.
Gantner S, Schmid M, Durr C, Schuhegger R, Steidle A, Hutzler P et al. (2006). In situ quantitation of the spatial scale of calling distances and population density-independent N-acylhomoserine lactone-mediated communication by rhizobacteria colonized on plant roots. FEMS Microbiol Ecol 56: 188–194.
Gardner A, West SA, Griffin AS . (2007). Is bacterial persistence a social trait? PLoS One 2: e752.
Gonzalez Barrios AF, Zuo R, Hashimoto Y, Yang L, Bentley WE, Wood TK . (2006). Autoinducer 2 controls biofilm formation in Escherichia coli through a novel motility quorum-sensing regulator (MqsR, B3022). J Bacteriol 188: 305–316.
Gupta A, Terrell JL, Fernandes R, Dowling MB, Payne GF, Raghavan SR et al. (2013). Encapsulated fusion protein confers ‘sense and respond’ activity to chitosan-alginate capsules to manipulate bacterial quorum sensing. Biotechnol Bioengineer 110: 552–562.
Ha JH, Eo Y, Grishaev A, Guo M, Smith JA, Sintim HO et al. (2013). Crystal structures of the LsrR proteins complexed with phospho-AI-2 and two signal-interrupting analogues reveal distinct mechanisms for ligand recognition. J Am Chem Soc 135: 15526–15535.
Hegde M, Englert DL, Schrock S, Cohn WB, Vogt C, Wood TK et al. (2011). Chemotaxis to the quorum-sensing signal AI-2 requires the Tsr chemoreceptor and the periplasmic LsrB AI-2-binding protein. J Bacteriol 193: 768–773.
Hense BA, Kuttler C, Muller J, Rothballer M, Hartmann A, Kreft JU . (2007). Does efficiency sensing unify diffusion and quorum sensing? Nat Rev Microbiol 5: 230–239.
Herzberg M, Kaye IK, Peti W, Wood TK . (2006). YdgG (TqsA) controls biofilm formation in Escherichia coli K-12 through autoinducer 2 transport. J Bacteriol 188: 587–598.
Hooshangi S, Bentley WE . (2011). LsrR quorum sensing ‘switch’ is revealed by a bottom-up approach. PLoS Comp Biol 7: e1002172.
Li J, Attila C, Wang L, Wood TK, Valdes JJ, Bentley WE . (2007). Quorum sensing in Escherichia coli is signaled by AI-2/LsrR: effects on small RNA and biofilm architecture. J Bacteriol 189: 6011–6020.
Luo X, Wu HC, Tsao CY, Cheng Y, Betz J, Payne GF et al. (2012). Biofabrication of stratified biofilm mimics for observation and control of bacterial signaling. Biomaterials 33: 5136–5143.
Maniatis T, Fritsch EF, Sambrook J . (1982) Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory: Cold Spring Harbor, : NY.
Mok KC, Wingreen NS, Bassler BL . (2003). Vibrio harveyi quorum sensing: a coincidence detector for two autoinducers controls gene expression. EMBO J 22: 870–881.
Parsek MR, Greenberg EP . (2005). Sociomicrobiology: the connections between quorum sensing and biofilms. Trends Microbiol 13: 27–33.
Pradhan BB, Chatterjee S . (2014). Reversible non-genetic phenotypic heterogeneity in bacterial quorum sensing. Mol Microbiol 92: 557–569.
Quan DN, Bentley WE . (2012). Gene network homology in prokaryotes using a similarity search approach: queries of quorum sensing signal transduction. PLoS Comp Biol 8: e1002637.
Schauder S, Shokat K, Surette MG, Bassler BL . (2001). The LuxS family of bacterial autoinducers: biosynthesis of a novel quorum-sensing signal molecule. Mol Microbiol 41: 463–476.
Siegele DA, Hu JC . (1997). Gene expression from plasmids containing the araBAD promoter at subsaturating inducer concentrations represents mixed populations. Proc Natl Acad Sci USA 94: 8168–8172.
Smith R, Tan C, Srimani JK, Pai A, Riccione KA, Song H et al. (2014). Programmed Allee effect in bacteria causes a tradeoff between population spread and survival. Proc Natl Acad Sci USA 111: 1969–1974.
Surette MG, Bassler BL . (1998). Quorum sensing in Escherichia coli and Salmonella typhimurium. Proc Natl Acad Sci USA 95: 7046–7050.
Swofford CA, Van Dessel N, Forbes NS . (2015). Quorum-sensing Salmonella selectively trigger protein expression within tumors. Proc Natl Acad Sci USA 112: 3457–3462.
Taga ME . (2005), Methods for analysis of bacterial autoinducer-2 production. Current protocols in microbiology Chapter 1: Unit 1C 1.
Tavender TJ, Halliday NM, Hardie KR, Winzer K . (2008). LuxS-independent formation of AI-2 from ribulose-5-phosphate. BMC Microbiol 8: 98.
Thompson JA, Oliveira RA, Djukovic A, Ubeda C, Xavier KB . (2015). Manipulation of the quorum sensing signal AI-2 affects the antibiotic-treated gut microbiota. Cell Rep 10: 1861–1871.
Tsao CY, Hooshangi S, Wu HC, Valdes JJ, Bentley WE . (2010). Autonomous induction of recombinant proteins by minimally rewiring native quorum sensing regulon of E. coli. Metabol Engineer 12: 291–297.
Ulitzur S, Yashphe J, Hastings JW . (1976). Inhibition and stimulation of the development of the bioluminescent system in Beneckea harveyi by cyclic GMP. Proc Natl Acad Sci USA 73: 4454–4456.
Vilchez R, Lemme A, Thiel V, Schulz S, Sztajer H, Wagner-Dobler I . (2007). Analysing traces of autoinducer-2 requires standardization of the Vibrio harveyi bioassay. Anal Bioanal Chem 387: 489–496.
Wang L, Hashimoto Y, Tsao CY, Valdes JJ, Bentley WE . (2005a). Cyclic AMP (cAMP) and cAMP receptor protein influence both synthesis and uptake of extracellular autoinducer 2 in Escherichia coli. J Bacteriol 187: 2066–2076.
Wang L, Li J, March JC, Valdes JJ, Bentley WE . (2005b). luxS-dependent gene regulation in Escherichia coli K-12 revealed by genomic expression profiling. J Bacteriol 187: 8350–8360.
Winzer K, Hardie KR, Burgess N, Doherty N, Kirke D, Holden MT et al. (2002). LuxS: its role in central metabolism and the in vitro synthesis of 4-hydroxy-5-methyl-3(2H)-furanone. Microbiology 148: 909–922.
Wu HC, Tsao CY, Quan DN, Cheng Y, Servinsky MD, Carter KK et al. (2013). Autonomous bacterial localization and gene expression based on nearby cell receptor density. Mol Syst Biol 9: 636.
Xavier KB, Bassler BL . (2005). Regulation of uptake and processing of the quorum-sensing autoinducer AI-2 in Escherichia coli. J Bacteriol 187: 238–248.
Yung CW, Barbari TA, Bentley WE . (2006). Integrated non-invasive system for quantifying secreted human therapeutic hIL2. Biotechnol Bioengineer 95: 938–945.
Zhu J, Miller MB, Vance RE, Dziejman M, Bassler BL, Mekalanos JJ . (2002). Quorum-sensing regulators control virulence gene expression in Vibrio cholerae. Proc Nati Acad Sci USA 99: 3129–3134.
Zhu J, Pei D . (2008). A LuxP-based fluorescent sensor for bacterial autoinducer II. ACS Chem Biol 3: 110–119.
Acknowledgements
We acknowledge support of the Defense Threat Reduction Agency (DTRA, HDTRA1-13-1-0037), the National Science Foundation (CBET 1264509 & CBET 1160005) and the RW Deutsch Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Servinsky, M., Terrell, J., Tsao, CY. et al. Directed assembly of a bacterial quorum. ISME J 10, 158–169 (2016). https://doi.org/10.1038/ismej.2015.89
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.89
This article is cited by
-
Bioelectronic control of a microbial community using surface-assembled electrogenetic cells to route signals
Nature Nanotechnology (2021)
-
Bacterial co-culture with cell signaling translator and growth controller modules for autonomously regulated culture composition
Nature Communications (2019)
-
Regulation of bacteria population behaviors by AI-2 “consumer cells” and “supplier cells”
BMC Microbiology (2017)
-
Electronic control of gene expression and cell behaviour in Escherichia coli through redox signalling
Nature Communications (2017)
-
Nano-guided cell networks as conveyors of molecular communication
Nature Communications (2015)