Abstract
The physical locations of 5S and 18S-25S rDNA sequences in 15 diploid Hordeum species with the I genome were examined by double-target in situ hybridization with pTa71 (18S-25S rDNA) and pTa794 (5S rDNA) clones as probes. All the three Asian species had a species-specific rDNA pattern. In 12 American species studied, eight different rDNA types were found. The type reported previously in H. chilense (the ‘chilense’ type) was observed in eight American species. The chilense type had double 5S rDNA sites — two sites on one chromosome arm separated by a short distance — and two pairs of major 18S-25S rDNA sites on two pairs of satellite chromosomes. The other seven types found in American species were similar to the chilense type and could be derived from the chilense type through deletion, reduction or addition of a rDNA site. Intraspecific polymorphisms were observed in three American species. The overall similarity in rDNA patterns among American species indicates the close relationships between North and South American species and their derivation from a single ancestral source. The differences in the distribution patterns of 5S and 18S-25S rDNA between Asian and American species suggest differentiation between the I genomes of Asian and American species. The 5S and 18S-25S rDNA sites are useful chromosome markers for delimiting Asian species, but have limited value as a taxonomic character in American species. On the basis of rDNA patterns, karyotype evolution and phylogeny of the I-genome diploid species are discussed.
Similar content being viewed by others
Introduction
The genus Hordeum (Triticeae) is classified into 31 species (in total 45 taxa) and 51 cytotypes at the diploid, tetraploid, and hexaploid levels with a basic chromosome number of x=7 (Baden & von Bothmer, 1994; von Bothmer et al., 1995). Based on the metaphase I (MI) chromosome pairing in hybrids (von Bothmer et al., 1986, 1995) the genus has been divided into four genome groups (H, I, X and Y). H. vulgare and H. bulbosum both carry the H genome, H. marinum carries the X genome, H. murinum has the Y genome, and the remaining species share variants of the I genome (genome designation according to Linde-Laursen et al., 1997). Recently, using genomic in situ hybridization, Taketa et al. (1999b) revealed that three polyploid species/cytotypes, H. secalinum, H. capense and H. brachyantherum 6x, were alloploids having a combination of the X and I genomes. Thus, the genus is divided into at least five groups according to their genome constitutions.
The I-genome group is the largest in the genus and includes 25 species. Fourteen species are diploid, five are tetraploid and four are hexaploid; the remaining two species include diploid, tetraploid and hexaploid cytotypes. The I-genome species are distributed from Central Asia to the Americas (von Bothmer et al., 1995). As a part of phylogenetic studies on the genus Hordeum, many I-genome species have been examined for C-banding patterns (Linde-Laursen et al., 1995), chloroplast DNA restriction patterns (Doebley et al., 1992), isoenzyme patterns (Jørgensen, 1986), rDNA restriction patterns (Molnar et al., 1989), repetitive DNA sequences (Svitashev et al., 1994), and RAPD marker profiles (Marillia & Scoles, 1996). Although these studies have confirmed the distinction of the four basic genomes in the genus, details on the relationships among the I-genome species are still largely unclear. Information on the phylogenetic relationship of the I-genome species is also indispensable for elucidating the phylogeny of the genus Elymus, the largest genus in Triticeae, because I is a component genome of Elymus (Dewey, 1984; Löve, 1984; Lu, 1993).
The 18S-5.8S-25S rRNA genes and intergenic spacer regions (18S-25S rDNA) exist as tandem repeats at the nucleolus organizer regions (NORs) in satellite (SAT) chromosomes and at other chromosomal sites where they may not be associated with a NOR (Mukai et al., 1991; Leitch & Heslop-Harrison, 1992; Pedersen & Linde-Laursen, 1994). The 5S rRNA genes and the intergenic spacers (5S rDNA) make up an independent multigene family and are not associated with a NOR (Mukai et al., 1990; Leitch & Heslop-Harrison, 1993). The physical mapping of these two multigene families by in situ hybridization has revealed species relationships in several genera (Aegilops: Bedaeva et al., 1996; Arachis: Raina & Mukai, 1999; Trifolium: Ansari et al., 1999). In the genus Hordeum, several species or cytotypes representing the four basic genomes were analysed and the usefulness of both rDNAs as phylogenetic markers was suggested (de Bustos et al., 1996; Taketa et al., 1999a). However, in the I-genome group, only one diploid species (H. chilense) and tetraploid cytotypes of two species (H. brevisubulatum and H. brachyantherum) were analysed (de Bustos et al., 1996; Taketa et al., 1999a) and most I-genome species remain to be tested. The physical mapping of both rDNAs would allow more precise estimation about the phylogenetic relationships of the I-genome species and may complement the estimation of karyotype evolution of the genus Hordeum by Linde-Laursen et al. (1995) using the SAT chromosome alone as an evolutionary marker. In the present study, we determined the physical locations of both 5S and 18S-25S rDNA in 15 diploid Hordeum species with the I genome by double-target fluorescence in situ hybridization. This molecular cytogenetic study provides new information on the karyotype evolution of I-genome diploid species of the genus Hordeum.
Materials and methods
Fifteen diploid species (20 taxa) of the I-genome group represented by 32 accessions were analysed in this study (Table 1). Actively growing root tips from germinating seeds or potted plants were treated in ice water at 0°C for 24 h to accumulate metaphases before fixation in 3:1 (v/v) 100% ethanol:acetic acid. The root tips were digested with an enzyme mixture containing cellulase and pectolyase (Fukui & Kakeda, 1990) and squashed in a drop of 45% acetic acid. Two DNA clones, pTa794 and pTa71 were used as probes. Clone pTa794 is a BamHI fragment of the 5S rDNA, that has a 120-bp coding sequence for the 5S rRNA gene and the intergenic spacers isolated from common wheat, Triticum aestivum L. (Gerlach & Dyer, 1980). Clone pTa71, the 18S-25S rDNA, is a 9-kb Eco RI fragment from common wheat, containing the coding sequences for the 18S, 5.8S, and 25S rRNA genes and the intergenic spacer sequences (Gerlach & Bedbrook, 1979). The in situ hybridization procedure described by Taketa et al. (1999a) was adopted. Slides were examined using an Olympus BX-50 epifluorescence microscope with appropriate filter sets (U-MWU for UV, U-MWIB for FITC, U-DM-Cy3 for Cy3, and U-DM-DA/FI/TX for simultaneous visualization of all fluorochromes). Photographs were taken on Fujicolor Super HG400 colour print film. Negatives were scanned to PhotoCD and printed from Adobe Photoshop using only cropping and processing functions that affect all pixels in the image equally. Idiograms of the chromosomes with hybridization sites were prepared on the basis of measurements of 10 homologous chromosomes.
Results
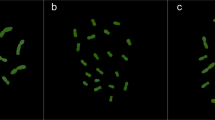
Figure 1(a–j) shows the somatic metaphase chromosomes of 10 representative species after in situ hybridization with 5S and 18S-25S rDNA probes and counterstaining with 4′,6-diamidino-2-phenylindole (DAPI). Idiograms of the chromosomes with rDNA sites are shown in Fig. 2. The present in situ hybridization experiment detected 5S and minor 18S-25S rDNA sites in addition to major 18S-25S rDNA sites. The number of major 18S-25S rDNA sites detected by in situ hybridization agrees with the number of SAT chromosomes reported for the respective species (Linde-Laursen et al., 1995). Here, we define SAT chromosomes as nucleolus organizing chromosomes with a secondary constriction. Depending on the species, either one or two pairs of major 18S-25S rDNA sites and up to two pairs of minor 18S-25S rDNA sites were observed on two to four pairs of chromosomes. The number of 5S rDNA sites varied from one to three pairs per species.
Double-target in situ hybridization to root tip metaphase cells from 10 representative diploid Hordeum species with the I genome. The micrographs were taken with a triple band filter allowing the simultaneous visualization of the DAPI-stained chromosomes (blue), the hybridization sites of the 18S-25S rDNA (red) and the 5S rDNA (green). Arrowheads indicate minor 18S-25S rDNA sites. Scale bar represents 10 μm in all figures. (a) H. brevisubulatum ssp. violaceum H315, (b) H. roshevitzii H7046, (c) H. bogdanii H240, (d) H. pubiflorum ssp. pubiflorum H1296, (e) H. euclaston H6036, (f) H. pusillum H2083, (g) H. brachyantherum ssp. californicum H3317, (h) H. muticum H6479, (i) H. flexuosum H1116, (j) H. cordobense H6429.
Asian species
The distribution patterns of 5S and 18S-25S rDNA sites in H. brevisubulatum and H. roshevitzii were similar except that H. brevisubulatum had a pair of additional metacentric SAT chromosomes with a major 18S-25S rDNA site (Fig. 1a, b). H. bogdanii had a unique metacentric chromosome pair with a proximal 5S rDNA site and a distal minor 18S-25S rDNA site on one chromosome arm (Fig. 1c). A submetacentric chromosome pair with 5S and 18S-25S rDNA sites and a metacentric chromosome pair with a minor 18S-25S rDNA site were common to all three Asian species. In H. bogdanii, the 18S-25S rDNA locus on the submetacentric chromosome pair was a major site with NOR-forming ability. In the other two species the locus consisted of a few blocks of weak 18S-25S rDNA signals that surrounded the 5S rDNA site without a visible secondary constriction. In H. brevisubulatum and H. roshevitzii, the major 18S-25S rDNA sites with NOR-forming ability were located on metacentric chromosome pair(s).
North and South American species
A total of eight rDNA types were found among the 12 American species (17 taxa) studied (Fig. 2). Eight species (13 taxa), namely the North American species H. brachyantherum (accession H1945) and H. intercedens, and the South American species H. chilense, H. erectifolium, H. euclaston (Fig. 1e), H. patagonicum (5 subspecies), H. pubiflorum (accessions H1348, H6046 and H6360) and H. stenostachys, shared the same rDNA pattern as that previously reported for a different accession of H. chilense (Taketa et al., 1999a). This rDNA pattern, named the chilense type, carried a submetacentric SAT-chromosome pair with double 5S rDNA sites — two sites on one chromosome arm separated by a short distance — plus a major and overlapping 18S-25S rDNA site, and a metacentric SAT-chromosome pair with a major 18S-25S rDNA site. On the basis of the studies using wheat — H. chilense chromosome addition lines (Miller et al., 1982; Cabrera et al., 1995), the submetacentric pair and the metacentric pair with rDNA sites are considered as wheat homoeologous group-5 and group-6 chromosomes, respectively. The remaining seven types were similar to the chilense type and are described briefly here. Two H. brachyantherum accessions (H2401 and H3317) had double 5S rDNA sites but no 18S-25S rDNA site on the submetacentric chromosome pair (Fig. 1g). One H. pubiflorum accession (H1296) had an additional 5S rDNA site in the long arm of the submetacentric SAT-chromosome pair (Fig. 1d). In H. pusillum, the submetacentric chromosome pair had only one 5S rDNA site overlapping with the 18S-25S rDNA site and the intensity of the 18S-25S rDNA signal varied among accessions; one accession (H1901) had a strong signal and two accessions (H722 and H2038) had a weaker signal (Fig. 1f). H. muticum (Fig. 1h) had a weaker 18S-25S rDNA signal on the metacentric chromosome pair. H. flexuosum (Fig. 1i) had only one 5S rDNA site distal to the major 18S-25S rDNA site on the submetacentric SAT-chromosome pair. H. cordobense (Fig. 1j) had a third chromosome pair with a minor 18S-25S rDNA site. Also in H. cordobense, the major 18S-25S rDNA site on the metacentric chromosome pair occupied a more distal location.
As summarized in Fig. 2, intraspecific polymorphism in rDNA pattern was observed in H. brachyantherum, H. pusillum and H. pubiflorum. Linde-Laursen et al. (1986) reported that H. brachyantherum and H. pusillum included populations with one and two pair(s) of SAT chromosomes. The present study revealed that accessions of these species with the metacentric SAT chromosome pair only had deletion or reduction of the major 18S-25S rDNA site on the submetacentric chromosome pair.
Discussion
Variation in rDNA pattern
The distribution patterns of 5S and 18S-25S rDNA sites differ somewhat between Asian species and American species. The number of rDNA-carrying chromosomes was higher in Asian species (three or four pairs) than in American species (usually two pairs). Asian species had species-specific rDNA patterns, while the American species shared the chilense type or its variants. Double 5S rDNA sites overlapping with a major 18S-25S rDNA site were observed in 10 of the 12 American species studied, but not in any Asian species. Thus, the present results suggest differentiation between the I genomes of Asian and American species.
The rDNA patterns in the I-genome species differ from those in the other three genomes (H, X and Y) of Hordeum (Leitch & Heslop-Harrison, 1993; Pedersen & Linde-Laursen, 1994; de Bustos et al., 1996; Taketa et al., 1999a), supporting the distinction of the I genome from the other genomes. In the I-genome species, 5S and 18S-25S rDNA sites on the short arm of the submetacentric chromosome pair occur in close proximity and their relative order cannot be resolved. Such close proximity of 5S and 18S-25S rDNA sites has not been reported in other Hordeum (de Bustos et al., 1996; Taketa et al., 1999a) and Aegilops (Bedaeva et al., 1996) species. Future studies employing prometaphase chromosomes or pachytene nuclei may reveal the relative order and organization of 5S and 18S-25S rDNA sites in the I-genome species.
Karyotype evolution of I-genome species
On the basis of the morphology of SAT chromosomes and other information, Linde-Laursen et al. (1995) proposed a hypothesis on karyotype evolution of I-genome species. The present in situ hybridization experiment detected minor 18S-25S rDNA sites and 5S rDNA sites in addition to the well-known major 18S-25S rDNA sites (Fig. 2). Taking advantage of such additional chromosome landmarks, we can critically evaluate the hypothesis on karyotype evolution of the I-genome species proposed by Linde-Laursen et al. (1995).
Linde-Laursen et al. (1995) estimated that an Asian outbreeding species, H. brevisubulatum was the most original form of I-genome diploid species and that two Asian inbreeding species, H. roshevitzii and H. bogdanii were derived from it, with a concomitant loss of two NOR loci. The present rDNA data support the view that H. roshevitzii was derived from H. brevisubulatum because of the close similarity in rDNA pattern. However, the marked differences in rDNA pattern between H. bogdanii and the other two Asian species do not support the direct derivation of H. bogdanii from H. brevisubulatum. Our rDNA data indicate that H. bogdanii occupies a unique position among the Asian species, which is also supported by, for example, morphological data (von Bothmer, 1979).
According to Rajhathy et al. (1964), I-genome species migrated from Asia to North America. Linde-Laursen et al. (1986, 1989) reported that most American species had a uniform karyotype with two SAT-chromosome pairs, one submetacentric pair with short and one metacentric pair with longer satellites (named ‘the common American diploid karyotype’). Linde-Laursen et al. (1995) hypothesized that the common American diploid karyotype was derived from an Asian species, H. brevisubulatum, with a concomitant loss of one NOR locus. However, in the rDNA pattern there are substantial differences between H. brevisubulatum and the American species. The detection of a minor 18S-25S rDNA site on a metacentric chromosome pair in H. roshevitzii also raises the possibility that the common American diploid karyotype could be derived from H. roshevitzii. In this case, we need to assume that the loss of the major 18S-25S rDNA site on the metacentric SAT-chromosome pair of H. roshevitzii resulted in amplification and activation of the other minor 18S-25S rDNA sites. From the present rDNA data, it is impossible to deduce which Asian species was the direct ancestor of American diploid species. Complete identification of the homoeology of the chromosomes of Asian species should help resolve the connection between Asian and American species.
Linde-Laursen et al. (1995) assumed that the common American diploid karyotype was the ancestral karyotype of American species and that other karyotypes found in American species were derived from it through deletion of one of the two NOR loci or a paracentric inversion. The minor 18S-25S rDNA sites detected in the present study have provided cytological evidence that deletions or reductions of major 18S-25S rDNA sites played an important role in the karyotype evolution of the I-genome species. The present data on 5S rDNA sites also revealed that the common American diploid karyotype includes four rDNA patterns (Fig. 2). Because the chilense type predominates in American species with the common American diploid karyotype, we propose that the ancestral rDNA pattern was the chilense type. Other explanations are more complicated and need more evolutionary steps. Of North American diploid species, H. brachyantherum ssp. californicum, with two SAT-chromosome pairs, is the best candidate for the chilense-type ancestor because this species has a perennial growth habit like most other American species. Other North American diploid species, H. pusillum and H. intercedens have an annual growth habit and are therefore unlikely to be the common ancestor of American species, which predominantly have a perennial growth habit.
A South American species, H. cordobense, has a unique karyotype with a smaller satellite on the metacentric SAT-chromosome pair than the species with the common American diploid karyotype (Linde-Laursen et al., 1989). However, the double 5S sites on the submetacentric SAT-chromosome pair in H. cordobense suggest its close relationship with other American species. Linde-Laursen et al. (1989) explained the smaller satellites of H. cordobense by assuming a paracentric inversion. However, a more likely explanation may be a transposition of the rDNA site because in Triticeae rDNA sites can change positions within a chromosome arm without disturbing the linkage groups (Dubcovsky & Dvorák, 1995). It is unknown how the minor 18S-25S rDNA sites in H. cordobense were originated.
From the present rDNA data, we infer that when the chilense type spread to the American Continents, variants of this type were derived, as shown in Fig. 2. This model suggests a monophyletic origin for American diploid species.
The phylogenetic relationships of I-genome species were estimated from other marker systems, such as chloroplast DNA restriction patterns (Doebley et al., 1992), isoenzyme patterns (Jørgensen, 1986), rDNA restriction patterns (Molnar et al., 1989), repetitive DNA sequences (Svitashev et al., 1994), and RAPD markers (Marillia & Scoles, 1996). Because each study differs in the kind and number of species analysed, comparisons with the present study are difficult. The present study did not test a South American species H. comosum. The study on chloroplast DNA restriction pattern by Doebley et al. (1992) showed differentiation of the two Asian species H. bogdanii and H. roshevitzii from other American I-genome species, but the study did not include H. brevisubulatum. Other studies did not clearly show differentiation between Asian and American I-genome species. For example, Southern hybridization using repetitive DNA sequences showed an association of two South American species, H. comosum and H. pubiflorum to the three Asian species (Svitashev et al., 1994), while H. chilense, a South American species and H. bogdanii, an Asian species, had an identical rDNA restriction pattern in separation from the other I-genome species (Molnar et al., 1989). Such associations between Asian and American species do not agree with the species relationships inferred from the present rDNA data.
In conclusion, the present study has shown that rDNA sites are useful chromosome markers for an investigation of karyotype evolution and phylogeny in the I-genome species of the genus Hordeum. Minor 18S-25S and 5S rDNA sites, which were undetectable in previous studies by Linde-Laursen et al. (1995), provided especially clear cytological evidence for modifications of SAT chromosomes that occurred during speciation. The present results are generally compatible with the hypothesis on karyotype evolution proposed by Linde-Laursen et al. (1995), but we also proposed a more likely model on karyotype evolution for several species. The distribution patterns of 5S and 18S-25S rDNA sites can be used for delimitation of the Asian I-genome species, but have a limited value as taxonomic characters in American I-genome species. The rDNA patterns of American I-genome species are characterized by the predominant occurrence of the chilense type and general lack of species-specificity. The low level of polymorphism in rDNA patterns among South American species contrasts sharply with the distinct morphological differences among species (von Bothmer et al., 1995). Such low levels of variations were also found in C-banding karyotypes (Linde-Laursen et al., 1989) and in rDNA restriction patterns (Molnar et al., 1989). Sequence analysis of the intergenic spacer and the internal transcribed sequences of rDNA (Rogers & Bendich, 1987; Baldwin, 1992) may reveal polymorphisms among South American species. The present rDNA data on I-genome diploid species would be useful for estimating not only the diploid ancestors of polyploid Hordeum species but also the I-genome donors of polyploid Elymus species.
References
Ansari, H. A., Ellison, N. W., Reader, S. M., Bedaeva, E. D., Friebe, B., Miller, T. E. et al (1999). Molecular cytogenetic organization of 5S and 18S–25S rDNA loci in white clover (Trifolium repens L.) and related species. Ann Bot, 83, 199–206.
Bedaeva, E. D., Friebe, B. and Gill, B. S. (1996). Genome differentiation in Aegilops. 2. Physical mapping of 5S and 18S–26S ribosomal RNA gene families in diploid species. Genome, 39, 1150–1158.
Baden, C. and von Bothmer, R. (1994). A taxonomic revision of Hordeum sect. Critesion. Nord J Bot, 14, 117–136.
Baldwin, B. G. (1992). Phylogenetic utility of the internal transcribed spacers of nuclear ribosomal DNA in plants: an example from the Compositae. Mol Phylogenet Evol, 1, 3–16.
von Bothmer, R. (1979). Revision of the Asiatic taxa of Hordeum sect. Stenostcahys. Bot Tidsskr, 74, 117–147.
von Bothmer, R., Fink, J. and Landström, T. (1986). Meiosis in interspecific hybrids. I. Diploid combinations. Can J Genet Cytol, 28, 525–535.
von Bothmer, R., Jacobsen, N., Baden, C., Jørgensen, R. B. and Linde-Laursen, I. (1995). An Ecogeographical Study of the Genus Hordeum, 2nd edn. Systematic and Ecogeographical Studies on Crop Genepools 7. IBPGR, Rome.
De Bustos, A., Cuadrado, A., Soler, C. and Jouve, N. (1996). Physical mapping of repetitive DNA sequences and 5S and 18S–26S rDNA in five wild species of the genus Hordeum. Chromosome Res, 4, 491–499.
Cabrera, A., Friebe, B., Jiang, J. and Gill, B. S. (1995). Characterization of Hordeum chilense chromosomes by C-banding and in situ hybridization using highly repeated DNA probes. Genome, 38, 435–442.
Dewey, D. R. (1984). The genomic system of classification as a guide to intergeneric hybridization with the perennial Triticeae. Stadler Genet Symp, 16, 209–279.
Doebley, J., von Bothmer, R. and Larson, S. (1992). Chloroplast DNA variation and the phylogeny of Hordeum (Poaceae). Am J Bot, 79, 576–584.
Dubcovsky, J. and Dvorák, J. (1995). Ribosomal RNA multigene loci: Nomads of the Triticeae genomes. Genetics, 140, 1367–1377.
Fukui, K. and Kakeda, K. (1990). Quantitative karyotyping of barley chromosomes by image analysis methods. Genome, 33, 450–458.
Gerlach, W. L. and Bedbrook, J. R. (1979). Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucl Acids Res, 7, 1869–1855.
Gerlach, W. L. and Dyer, T. A. (1980). Sequence organization of the repeated units in the nucleus of wheat which contains 5S-rDNA genes. Nucl Acids Res, 8, 4851–4865.
Jørgensen, R. B. (1986). Relationships in the barley genus (Hordeum): an electrophoretic examination of proteins. Hereditas, 104, 273–291.
Leitch, I. J. and Heslop-Harrison, J. S. (1992). Physical mapping of the 18S-5.8S–26S rRNA genes in barley by in situ hybridization. Genome, 35, 1013–1018.
Leitch, I. J. and Heslop-Harrison, J. S. (1993). Physical mapping of four sites of 5S rDNA sequences and one site of the LE-amylase-2 gene in barley (Hordeum vulgare). Genome, 36, 517–532.
Linde-Laursen, I., von Bothmer, R. and Jacobsen, N. (1986). Giemsa C-banded karyotypes of Hordeum taxa from North America. Can J Genet Cytol, 28, 42–62.
Linde-Laursen, I., von Bothmer, R. and Jacobsen, N. (1989). Giemsa C-banded karyotypes of South American Hordeum (Poaceae). I. 14 diploid taxa. Hereditas, 110, 289–305.
Linde-Laursen, I., von Bothmer, R. and Jacobsen, N. (1995). Karyotype differentiation and evolution in the genus Hordeum (Poaceae). In: Brandham, P. E. and Bennett, M. D. (eds) Kew Chromosome Conference IV, pp. 233–247. Royal Botanic Gardens, Kew.
Linde-Laursen, I., Heslop-Harrison, J. S., Shepherd, K. W. and Taketa, S. (1997). The barley genome and its relationship with the wheat genomes. A survey with an internationally agreed recommendation for barley chromosome nomenclature. Hereditas, 126, 1–16.
Löve, A. (1984). Conspectus of the Triticeae. Feddes Rep, 95, 425–521.
Lu, B. -R. (1993). Biosystematic investigations of Asiatic wheatgrasses – Elymus L. (Triticeae: Poaceae). Ph.D. Thesis, The Swedish Univeristy of Agricultural Sciences.
Marillia, E. F. and Scoles, G. J. (1996). The use of RAPD markers in Hordeum phylogeny. Genome, 39, 646–654.
Miller, T. E., Reader, S. M. and Chapman, V. (1982). The addition of Hordeum chilense chromosomes to wheat. In: Broertjes, C. (ed.) Induced Variability in Plant Breeding: Proceedings of the Eucarpia International Symposium, Wageningen, The Netherlands, pp. 79–81. Pudoc, Wageningen.
Molnar, S. J., Gupta, P. K., Fedak, G. and Wheatcroft, R. (1989). Ribosomal DNA repeat unit polymorphism in 25 Hordeum species. Theor Appl Genet, 78, 387–392.
Mukai, Y., Endo, T. R. and Gill, B. S. (1990). Physical mapping of the 5S rRNA multigene family in common wheat. J Hered, 81, 290–295.
Mukai, Y., Endo, T. R. and Gill, B. S. (1991). Physical mapping of the 18S.26S rRNA multigene family in common wheat: Identification of a new locus. Chromosoma, 100, 71–78.
Pedersen, C. and Linde-Laursen, I. (1994). Chromosomal locations of four minor rDNA loci and marker microsatellite sequence in barley. Chromosome Res, 2, 65–71.
Raina, S. N. and Mukai, Y. (1999). Detection of a variable number of 18S-5.8S–26S and 5S ribosomal DNA loci by fluorescent in situ hybridization in diploid and tetraploid Arachis species. Genome, 42, 52–59.
Rajhathy, T., Morrison, J. W. and Symko, S. (1964). Interspecific and intergeneric hybrids in Hordeum. In: Brookhuizen, I. S., Dantuma, G., Lamberts, H. and Lange, W. (eds) Barley Genetics, pp. 195–212. Pudoc, Wageningen.
Rogers, S. O. and Bendich, A. J. (1987). Ribosomal RNA genes in plants: variability in copy number and in the intergenic spacer. Plant Mol Biol, 9, 509–520.
Svitashev, S., Bryngelsson, T., Vershinin, A., Pedersen, C., Säll, T. and von Bothmer, R. (1994). Phylogenetic analysis of the genus Hordeum using repetitive DNA sequences. Theor Appl Genet, 89, 801–810.
Taketa, S., Harrison, G. E. and Heslop-Harrison, J. S. (1999a). Comparative physical mapping of the 5S and 18S–25S rDNA in nine wild Hordeum species and cytotypes. Theor Appl Genet, 98, 1–9.
Taketa, S., Ando, H., Takeda, K. and von Bothmer, R. (1999b). Detection of Hordeum marinum genome in three polyploid Hordeum species and cytotypes by genomic in situ hybridization. Hereditas, 130, 185–188.
Acknowledgements
The authors are particularly grateful to Dr J.S. Heslop-Harrison, Dr T. Schwarzacher and Ms G. Harrison (John Innes Centre, UK) for their kind guidance about in situ hybridization techniques, Dr A. Martin (Superior Council of Scientific Investigations, Spain) for the supply of H. chilense accessions, and Dr M. Ichii (Kagawa University, Japan) for encouragement. The work was supported in part by a grant from the Ministry of Education, Science, Sports and Culture, Japan (No. 09760006) and a grant from Kagawa University. R. von Bothmer acknowledges a visiting professorship supported by the Ministry of Education, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Taketa, S., Ando, H., Takeda, K. et al. Physical locations of 5S and 18S-25S rDNA in Asian and American diploid Hordeum species with the I genome. Heredity 86, 522–530 (2001). https://doi.org/10.1046/j.1365-2540.2001.00768.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00768.x
Keywords
This article is cited by
-
Oak ribosomal DNA: characterization by FISH and polymorphism assessed by IGS PCR–RFLP
Plant Systematics and Evolution (2016)
-
Use of the synthetic Oligo-pTa535 and Oligo-pAs1 probes for identification of Hordeum chilense-origin chromosomes in hexaploid tritordeum
Genetic Resources and Crop Evolution (2016)
-
Functional repetitive sequences and fragile sites in chromosomes of Lolium perenne L.
Protoplasma (2015)
-
Physical localization of rRNA genes by fluorescence in situ hybridization (FISH) and analysis of spacer length variants of 45S rRNA (slvs) genes in some species of genus Sesbania
Plant Systematics and Evolution (2014)
-
Analysis of chromosomal polymorphism in barley (Hordeum vulgare L. ssp. vulgare) and between H. vulgare and H. chilense using three-color fluorescence in situ hybridization (FISH)
Journal of Applied Genetics (2013)