Abstract
Geostatistics were used for describing and modelling spatial patterns of gene frequencies at eight allozyme loci in 93 populations of sessile oak well distributed over the natural range of the species. Spatial patterns were summarized by the variogram, a measure of gene frequency variance as a function of distance. They were of three kinds: stationary patterns, clines and random distributions of allele frequencies. The main directions of variation of gene frequencies over Europe were estimated using directional geostatistics. They were shown to reflect postglacial migration routes as well as a genetic divergence among populations from different glacial refugia. A method of spatial interpolation, called kriging, was used to draw synthetic maps for allele frequencies and for multilocus patterns of variation using two canonical variables. It appears from this study that geostatistics are useful tools for the description of complex patterns of genetic variation, and therefore also for the management of natural genetic resources in forest trees.
Similar content being viewed by others
Introduction
Long-lived woody plant species, particularly those with large geographical ranges and outcrossing breeding systems, maintain higher levels of genetic variation than most other plant species, but have generally very low levels of genetic differentiation among populations (Hamrick et al., 1992). The study of geographical variation in forest trees, which is particularly of interest for the identification and characterization of provenance regions, therefore necessitates large and range-wide sampling as well as accurate descriptive methods (Zanetto & Kremer, 1995)
Most studies conducted over the whole ranges of species have detected geographical patterns, which were most often gradual trends of variation. Among the descriptive methods used have been multivariate or clustering methods (Kinloch et al., 1986; Cheliak et al., 1988; Lagerkrantz & Ryman, 1990), the examination of the relationship between genetic and geographical distances (Li & Adams, 1989)or between allele frequencies and geographical variables such as latitude or longitude (Zanetto & Kremer, 1995), or the assessment of the variation within and between regional groups of populations (Li & Adams, 1989; Zanetto & Kremer, 1995). Implicit in all these studies is the assumption that there is a spatial autocorrelation among the data, that geographically close populations will share higher genetic similarity than populations located further apart. A stable autocorrelation of neutral gene frequencies may result from limited gene flow (‘isolation-by-distance’, Sokal & Wartenberg, 1983). Similarly, selection may lead to clines or to local genetic homogeneity (Sokal et al., 1989) Spatial autocorrrelation may also arise after range expansions (Sokal & Menozzi, 1982), and is particularly likely to persist in long-lived species like forest trees (Strauss et al., 1992)
However, the classical descriptive methods do not actually take into account spatial autocorrelation, because they all assume statistical independence among samples. Specific statistical tools have been developed that focus on the analysis of spatial dependence (see, e.g., Cressie, 1991, for a review). Among them are geostatistics and spatial autocorrelation analysis. The former approach was initially developed in the fields of geology and mining (Matheron, 1963), and has only recently been used in the field of ecology (Rossi et al., 1992), and also, but much more scarcely, in population genetics (Barbujani, 1988a; Monestiez et al., 1994). Conversely, autocorrelation analysis has been extensively used for the study of spatial genetic structures, after it was introduced into the field by Sokal & Oden (1978). The two approaches are closely related, but geostatistics has some advantages. First, it relies on a single, straightforward measure of spatial dependence called the variogram, whereas different expressions exist for the autocorrelation coefficient. Secondly, geostatistics allows the modelling of the spatial dependence, and, subsequently, the prediction (called ‘kriging’) of the variable everywhere in the studied space to obtain a map. Furthermore, like the correlogram, the variogram has a genetical interpretation within the frame of isolation-by-distance models (Piazza et al., 1981).
We will use geostatistics to describe spatial patterns of genetic variation in a range-wide set of European sessile oak (Quercus petraea (Matt.) Liebl.) populations analysed at eight polymorphic allozyme loci. Part of this sampling scheme was previously analysed using classical statistics (Zanetto & Kremer, 1995). In this study, geostatistics are used as a way to confirm and describe in more detail the geographical patterns previously detected. These patterns consisted mainly of the presence at most loci of a west–east cline, which was interpreted as a consequence of the northwards postglacial migration of oaks from separate refugia (located presumably in south Spain, south Italy and the Balkans).
We will describe the spatial patterns of 11 allozymic alleles and two canonical variables using the variogram. The main directions of geographical variation will be determined using directional geostatistics, and kriged maps will be drawn to illustrate patterns of genetic variation.
Materials and methods
Sampling of populations
The populations analysed in this study include 81 populations previously sampled and analysed electrophoretically by Zanetto & Kremer (1995), as well as an additional set of 12 populations from eastern Europe. All populations are of putative native origin and are well distributed over the natural range of the species, except that no population from Italy nor from eastern Germany could be included. Random samples of 120 seeds in each population were taken for allozyme analyses.
Allozyme analyses
Eight polymorphic loci where characterized in all the populations, either as part of the previous work by Zanetto & Kremer (1995) or for the present study. They are: Aap-A (alanine aminopeptidase, EC 3.4.11.1), Acp-C (acid phosphatase, EC 3.1.3.2), Dia-A (diaphorase, EC 1.8.1.4), Got-B (glutamic-oxaloacetic transaminase, EC 2.6.1.1), Lap-A (leucine aminopeptidase, EC 3.4.11.1), Mr-A (menadione reductase, EC 1.6.99.2), Pgi-B (phosphoglucoisomerase, EC 5.3.1.9) and Pgm (phosphoglucomutase, EC 5.4.2.2). Protein extraction and starch gel electrophoresis were performed as described in Zanetto et al. (1994)
For geostatistical analyses, only the most common allele was considered at each locus showing one or two major alleles and several minor alleles (Acp, Got, Lap, Mr and Pgm). For those loci showing three common alleles, the two most frequent were considered (Aap, Dia and Pgi). A total of 11 alleles was analysed. Their frequency distributions are summarized in Table 1. In addition to the analysis at individual loci, a factorial discriminant analysis (FDA) was carried out on individual multilocus genotypic data, in which the ratio of the variance among populations to the variance within populations was maximized. The first two canonical variables (FDA1 and FDA2), which accounted, respectively, for 20.6 and 13.6% of the total variation, were analysed using geostatistics.
Identification of outliers
Because geostatistics are based on measures of difference between pairs of locations, an unusually large or small value at a given location may strongly influence the results. Hawkins's test (Rossi et al., 1992)provides a way to detect outliers by comparing each value at location x, z(x), to its neighbouring values. Let n be the number of neighbouring values excluding z(x), M the arithmetic mean of the n- values, and σ2 the average variance measured over all equivalently sized neighbourhoods over the sampling space. Hawkins has shown the statistic n[ z(x)− M]2/(n−1)σ2 to be distributed as a χ2 with one degree of freedom. There is no objective criterion for the choice of the neighbourhood size. We used the smallest size for which most neighbourhoods contained more than five points, 450 km. Outliers were eliminated at the 5% significance level.
Construction of the variogram
The variogram measures the semivariance of the difference between points separated by a given distance h in space. The underlying assumption, called the ‘intrinsic hypothesis’ is that the difference between the values at any two points is a function of the distance between the points only. The empirical variogram is estimated from the data as a discrete function of given distance classes by:

where n is the number of points in the distance class h considered (Rossi et al., 1992)
The variogram may be calculated for all directions in space, or for specific directions to test for anisotropy. In the latter case, only the pairs of points connected along a given direction in space (±a tolerance angle) are taken into account in the calculations.
Geostatistical analyses consist of describing and modelling the variogram as a function of distance. The variogram usually increases, then reaches a constant value called the sill. The distance at which the sill is attained, called the range, represents the distance beyond which two values may be considered as independent. The variogram theoretically equals zero at the origin, but a discontinuity is often observed, called the ‘nugget effect’. This initial variance is caused either by fine-scale structures or by measurement errors. The empirical variogram may be adjusted to several theoretical models including the exponential, Gaussian and spherical models. The linear model is used for variograms that do not reach a plateau, and characterizes either clinal patterns, or patterns with ranges exceeding the maximum distance considered. Flat variograms, representing the ‘pure nugget’ effect, characterize random patterns of variation.
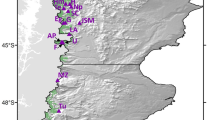
In this study, variograms were calculated separately for the whole sampling area (93 populations) and for a subarea corresponding to western Europe (67 populations, see Fig. 1). These separate analyses were conducted because the western and the eastern parts of the natural distribution are known to have been colonized from different glacial refugia (Ferris et al., 1993; Petit et al 1993). The objective was to test whether different migration routes could be detected by different directional variograms. However, the limited number of populations in the eastern part did not allow a separate analysis to be computed for that region.
A classical rule of thumb is to consider only pairs of points separated by less than half the maximum distance observed. Variograms were therefore calculated for 10 distance classes ranging from 0 to 1500 km for the whole area, and from 0 to 1000 km for the western subarea. More than 100 pairs of points were present in each class. Directional variograms were calculated for the following four sectors: 0±22.5 degrees (west–east), 45±22.5 degrees (southwest–northeast), 90±22.5 degrees (south–north), and 135±22.5 degrees (southeast–northwest), and within the same range of distances as the global variograms (but for eight classes only, to ensure that sufficient numbers of pairs were considered). Each variogram was standardized by dividing its values by the overall variance. This transformation will allow comparisons among different variables.
The variograms were adjusted either to the exponential model:

where c is the nugget, b is the difference between the sill and the nugget, and a is the distance at which 95% of the sill is asymptotically attained, or to the linear model:

A weighted least squares method was used to fit both models to the observed data (Cressie, 1991, p. 99). The model for which the sum of squared residuals was smaller was retained.
Genetical interpretation of the variogram
Variograms showing a sill can be interpreted within the frame of isolation-by-distance models. For a linear set of populations, Piazza et al. (1981) have shown that, at genetic equilibrium, the expression of the standardized variogram for gene frequencies is:

where σ is the standard deviation of parent-to-offspring dispersal distances and k is a ‘recall’ term representing mutation or selection.
This expression has been found to be empirically valid for a wide range of sampling schemes, including two-dimensional ones (Cavalli-Sforza & Feldman, 1990). In our study, it was used to interpret the ranges of the exponential variograms in terms of dispersal variances. For this purpose, we assumed k to be a mutation rate equal to 10−7.
Detecting mean directions of variation
The directional variograms were used to compute for each variable its mean direction of spatial variation following the method of Barbujani (1988b). Briefly, this method estimates the components of variation along each direction as the correlation coefficient (Kendal's τ) between each directional variogram value and the lag distance. The mean direction of variation is then computed as the average over the directional components. The significance of the mean direction may be assessed using Rayleigh's test (Greenwood & Durand, 1955), as described in Barbujani (1988b)
Kriging
Kriging uses the observed values of the variable in conjunction with the information provided by the variogram to predict values at unsampled locations. The estimates are linear combinations of the observed values taken within a given search neighbourhood. They are both optimal (the estimation variance is minimized) and unbiased (the mean of the estimation errors equals zero). Kriging was performed on a regular grid of 1517 equally spaced points (one point each 50 km) covering the sampling area. The search neighbourhoods were the ellipsoids defined by the ranges of the directional variograms for the variable studied (Wackernagel, 1995, p. 46).
Results
Between one and seven points (mean: 4.33) were eliminated as outliers for each variable, so that the variograms were calculated from 86 to 92 data points for the whole sampling area, and from 62 to 67 data points for the western subarea.
Variograms for the whole sampling area (Fig. 3 and Table 2)
Main types of experimental variograms and their adjusted models. Black circles represent estimated variogram values over the whole sampling area. Values estimated over the western subarea are represented by open circles. The variograms for the alleles Aap6, Dia2 and Pgi3 were similar to that for Aap4 (exponential model). The variogram for the allele Lap4 was similar to that for Acp2 (linear model with a small nugget effect). The variograms for Dia3, Got2, Mr2 and Pgi4 were similar to that for Pgm3 (predominance of the nugget effect).
Among the 11 alleles analysed, four (Aap4, Aap6, Pgi3 and Dia2), showed empirical variograms that fitted better to the exponential model. The adjusted ranges were 565 km, 623 km, 995 km and 776 km, respectively. The corresponding estimates of the standard deviation of dispersal distances, obtained by fitting an isolation-by-distance model, varied between 84 m (for Aap4) and 148 m (for Pgi3).
The other variograms fitted better to the linear model. The nugget effect accounted for more than 50% of the variation for Dia3, Got2, Mr2, Pgi4 and Pgm3. Although a clinal trend was present, the spatial variation for these five alleles was therefore predominantly random. Conversely, a clinal pattern was predominant for Acp2, and, to a lesser extent, for Lap4.
The empirical variogram for the first canonical variable, FDA1, showed two distinct spatial structures. From 0 to 700 km, the variogram fitted an exponential model with a range of 750 km. From 700 to 1500 km, the variogram showed a rapid linear increase, indicative of a strong clinal pattern. The variogram for the second canonical variable, FDA2, fitted a linear model with a high random component.
Variograms for the western subarea
The same four alleles as for the whole area showed exponential variograms. However, except for Pgi3, very long ranges (of the order of thousands of kilometres) were found, which indicates rather clinal patterns of variation, and nugget effects were higher than before. In particular, the patterns of variation at the alleles Dia3, Got2 and Pgm3 may be considered as purely random.
Mean directions of variation
Rayleigh's tests were all significant, indicating the presence of some anisotropy for all the variables, over the whole area as well as over the western subarea. Over the whole sampling area (Fig. 4a), the mean variation for most alleles is roughly along a west–east direction. The two exceptions are the alleles Pgi3 and Dia2, whose mean directions of variation are in the south–north direction. The first canonical variable, FDA1, varies mainly along a west–east direction, whereas the mean direction of variation of the second canonical variable, FDA2 is rather south–north. Over the western subarea, the mean directions of allelic variation showing a nonrandom pattern and the variation for FDA1 are approximately all distributed between the southwest– northeast and the south–north directions (Fig. 4b)
Kriged maps
An example of an allele frequency map is shown in Fig. 5, for the allele Aap4. There is a clear east–west trend of variation superimposed with some patches.
The kriged map for the first canonical variable, FDA1, was constructed using the exponential model fitted to its first component of variation. The resulting map (Fig. 6) clearly shows a gradual west–east cline, with a few patches of contrasting values. For FDA2, kriging was performed using a linear model. The map (Fig. 7)shows mainly the presence of a south–north cline throughout Europe. Although the two variables FDA1 and FDA2 follow different patterns, their kriged maps exhibit some similarities. For example, oak stands from France, Belgium, the Netherlands, northern Germany and Denmark are clearly separated from others.
Discussion
This study is the third report of range-wide variation at allozyme loci in sessile oak. In the first study (Zanetto & Kremer, 1995), it was shown that despite a very low level of genetic differentiation (mean FST-value over eight loci: 0.025), the allozyme variation was organized geographically. A majority of the allele frequencies was significantly correlated with longitude, whereas some were correlated with latitude. These trends were reinforced when multilocus and multivariate methods of analysis were applied (Kremer & Zanetto, 1997)
The enlargement of the sampling scheme for this study did not modify the observed level of genetic differentiation (mean FST-value over the same eight loci: 0.024). The use of geostatistics largely confirmed the geographical patterns previously observed by showing the presence of significant anisotropies at all allele frequencies, with a predominance of variations in the west–east direction. However, geostatistics provided a much more detailed insight into these geographical patterns. First, mean directions of variation were defined more precisely and were compared between the total sampling area and the western part of the species range. Secondly, the variograms indicated the presence of three kinds of spatial patterns: stationary patterns characterized by a finite range, clines and random patterns. Thirdly, the kriging method provided synthetic visual representations of the genetic variation of Q. petraea over its natural range.
These additional results offer new perspectives for the interpretation of the geographical patterns of genetic diversity. On account of the recent colonization history and present range-wide distribution of sessile oak, clinal trends of variation may result from three causes. First, the south-to-north postglacial range expansion may have caused a parallel cline in allele frequencies (Wijsman & Cavalli-Sforza, 1984). Secondly, clines may result from the contact between populations arising from different refugia (Zanetto & Kremer, 1995)Thirdly, selection acting along ecological gradients may also account for the presence of range-wide clines. Stationary patterns are known to result from isolation-by-distance among populations (Barbujani, 1988a), but they may also be caused by local selection (Sokal et al., 1989) Random patterns may be attributable to noise resulting from the sampling of populations or loci. Because of the high number of individuals analysed per population (120) the standard deviation of allele frequencies amounts in the worst case to 3.2%. Similarly, the high number of pairs of populations within each distance class ensures a good precision in the estimation of the variograms. The presence of a nugget effect would therefore indicate the presence of spatial structures occurring at a scale too small to be analysed with the present sampling scheme.
Heterogeneity of variograms among loci
Differences in the geographical patterns of variation at different unlinked loci are often taken as evidence of the action of natural selection (Barbujani, 1988b). However, Easteal (1989) also invoked the possible role of independent genetic drift events during range expansion in shaping different patterns of variation at different loci.
Among the eight allozyme loci analysed, only two pairs (Lap– Aap and Mr– Dia) exhibited significant linkage disequilibrium (Kremer & Zanetto, 1997). The different types of variogram observed (exponential, linear or random), as well as the different main directions of variation, were therefore shared by different independent loci. This suggests the combined action of a few systematic forces rather than the action of selection. The heterogeneity among variograms may result from the presence of different allele frequency patterns at the different loci before the postglacial expansion took place (Le Corre et al., 1997)
Directions of allelic variation and the past history of sessile oak populations
Palynological data (Huntley & Birks, 1983)provide evidence for the existence of three main glacial refugia for deciduous oaks: southern Spain, southern Italy and the Balkans. Moreover, the phylogeographical distribution of chloroplast DNA variation in deciduous oak species (Petit et al., 1993; Dumolin-Lapègue et al., 1997) attests to the distinct glacial origins of the present populations, and suggests that most populations in western Europe originated from Spain as a result of migration in a southwest to north-east direction. The main directions of allelic variation detected over the western part of the range are in agreement with this supposed direction of postglacial expansion. Clinal trends of variation were present for seven alleles among the 11 analysed. This indicates that founding events during the northwards expansion of sessile oak still have persistent effects on its present genetic structure at allozyme loci.
Over Europe, only two allele frequencies varied in a south–north direction. All other alleles showed a mean west–east direction of variation, which is in agreement with the hypothesis of a cline occurring after the populations that originated from south Spain and those that originated from Italy or the Balkans came into contact. The establishment of such a cline implies that allele frequencies in different refugia diverged in opposite directions during the glacial period, a situation which seems to have occurred more often than expected by chance. A bias may have been introduced by considering only alleles with intermediate frequencies, upon which genetic drift can act towards fixation or loss with equal probability. On the other hand, genetic divergence between the different refugia may have been reinforced by the action of divergent selection pressures at some loci, if environmental conditions differed among the different refugia during the glacial period. This hypothesis is in agreement with the fact that the patterns of variation for allozymes are more correlated with the glacial origin of populations than those for presumably neutral nuclear DNA markers (Le Corre et al., 1997)
Stationary patterns of variation
Stationary patterns of variation were found for only four alleles among the 11 analysed, but it is likely that such patterns were masked for other alleles by the presence of clinal trends of variation. The estimates of standard deviation of dispersal distances obtained by fitting an isolation-by-distance model to the stationary variograms have only an illustrative value, because genetic patterns are probably not at equilibrium, and also because of the uncertainty about the mutation rate used for estimation. The values obtained are, however, in agreement with the modes of dispersal of oaks (bird dispersal for the seed, wind dispersal for the pollen).
The ranges of the stationary variograms (found to vary roughly between 500 and 1000 km) can also be compared with the distances at which autocorrelation becomes negative in spatial autocorrelation analyses of natural populations. The two values are analogous and indicate the approximate size of homogeneous patches for the variable studied. In their study of 81 sessile oak populations, Zanetto & Kremer (1995) found distances that varied roughly between 400 and 650 km. Distances of about 700 km and 800 km were found, respectively, in Quercus ellipsoidallis (Jensen, 1986) and Quercus ilex (Michaud et al., 1995). These results and the present data suggest that the genetic structure of forest tree populations over their natural ranges is characterized by very large scales of variation.
Application of geostatistics to the management of natural genetic resources in forest trees
Geostatistics appear to be useful tools for the description of complex range-wide patterns of genetic variation in forest trees. The method of kriging applied to canonical variables provided synthetic and visual representations which summarized well the spatial genetic structures in sessile oak. More generally, geostatistics will probably be useful for the management of genetic resources of forest trees. Kriged maps of genetic diversity or the numbers of alleles within populations may be of direct interest for conservation purposes. Geostatistical analysis and mapping of adaptive traits may be used for the elaboration of collecting strategies for breeding, as proposed by Monestiez et al. (1994) in their study of perennial ryegrass populations. Kriged maps for sufficiently discriminant genetic markers may also be useful in forest management, particularly for the assessment of the native or introduced origin of populations.
References
Barbujani, G. (1988a). Diversity of some gene frequencies in European and Asian populations. IV. Genetic population structure assessed by the variogram. Ann Hum Genet, 52: 215–225.
Barbujani, G. (1988b). Detecting and comparing the direction of gene-frequency gradients. J Genet, 67: 129–140.
Cavalli-Sforza, L. L. and Feldman, M. W. (1990). Spatial subdivision of populations and estimates of genetic variation. Theor Pop Biol, 37: 3–25.
Cheliak, W. M., Wang, J. and Pitel, J. A. (1988). Population structure and genic diversity in tamarack, Larix laricina (Du Roi) K. Koch. Can J Forest Res, 18: 1318–1324.
Cressie, N. (1991). Statistics for Spatial Data. Wiley, New York.
Dumolin-Lapègue, S., Démesure, B., Fineschi, S., Lecorre, V. and Petit, R. J. (1997). Phylogeographic structure of white oaks throughout the European continent. Genetics, 146: 1475–1487.
Easteal, S. (1989). The effects of genetic drift during range expansion on geographical patterns of variation: a computer simulation of the colonization of Australia by Bufo marinus. Biol J Linn Soc, 37: 281–295.
Ferris, C., Olivier, R. P., Davy, A. J. and Hewitt, G. M. (1993). Native oak chloroplasts reveal an ancient divide across Europe. Mol Ecol, 2: 337–344.
Greenwood, J. A. and Durand, D. (1955). The distribution of length and components of the sum of n random units vectors. Ann Math Stat, 26: 233–246.
Hamrick, J. L., Godt, M. J. W. and Sherman-Broyles, S. L. (1992). Factors influencing levels of genetic diversity in woody plant species. New Forests, 6: 95–124.
Huntley, B. and Birks, H. J. B. (1983). An Atlas of Past and Present Pollen Maps for Europe: 0–13000 Years Ago. Cambridge University Press, Cambridge.
Jensen, R. J. (1986). Geographic spatial autocorrelation in Quercus ellipsoidalis. Bull Torrey Bot Club, 113: 431–439.
Kinloch, B. B., Westfall, R. D. and Forrest, G. I. (1986). Caledonian Scots pine: origins and genetic structure. New Phytol, 104: 703–729.
Kremer, A. and Zanetto, A. (1997). Geographical structure of gene diversity in Quercus petraea (Matt.) Liebl. II. Multilocus patterns of variation. Heredity, 78: 76–489.
Lagerkrantz, U. and Ryman, N. (1990). Genetic structure of Norway Spruce (Picea abies): concordance of morphological and allozymic variation. Evolution, 44: 38–53.
Lecorre, V., Dumolin-Lapègue, S. and Kremer, A. (1997). Genetic variation at allozyme and RAPD loci in sessile oak Quercus petraea (Matt.) Liebl.: the role of history and geography. Mol Ecol, 6: 519–529.
Li, P. and Adams, W. T. (1989). Range-wide patterns of allozyme variation in Douglas-fir (Pseudotsuga menziesii). Can J Forest Res, 19: 149–161.
Matheron, G. (1963). Principles of geostatistics. Econ Geol, 58: 1246–1266.
Michaud, H., Toumi, L., Lumaret, R., Li, T. X., Romane, F. and Anddigiusto, F. (1995). Effect of geographical discontinuity on genetic variation in Quercus ilex L. (holm oak). Evidence from enzyme polymorphism. Heredity, 74: 590–606.
Monestiez, P., Goulard, M. and Charmet, G. (1994). Geostatistics for spatial genetic structures: study of wild populations of perennial ryegrass. Theor Appl Genet, 88: 33–41.
Petit, R. J., Kremer, A. and Wagner, D. B. (1993). Geographic structure of chloroplast DNA polymorphisms in European oaks. Theor Appl Genet, 87: 122–128.
Piazza, A., Menozzi, P. and Cavalli-Sforza, L. (1981). The making and testing of geographic gene-frequency maps. Biometrics, 37: 635–659.
Rossi, R. E., Mulla, D. J., Journel, A. G. and Franz, E. H. (1992). Geostatistical tools for modeling and interpreting ecological spatial dependence. Ecol Monogr, 62: 277–314.
Sokal, R. R. and Menozzi, P. (1982). Spatial autocorrelations of HLA frequencies in Europe support demic diffusion of early farmers. Am Nat, 119: 1–17.
Sokal, R. R. and Oden, N. L. (1978). Spatial autocorrelation in biology. 1. Methodology. Biol J Linn Soc, 10: 199–228.
Sokal, R. R. and Wartenberg, D. E. (1983). A test of spatial autocorrelation analysis using an isolation-by-distance model. Genetics, 105: 219–237.
Sokal, R. R., Jacquez, G. M. and Wooten, M. C. (1989). Spatial autocorrelation analysis of migration and selection. Genetics, 121: 845–855.
Strauss, S. H., Bousquet, J., Hipkins, V. D. and Hong, Y. P. (1992). Biochemical and molecular genetic markers in biosystematic studies of forest trees. New Forests, 6: 125–158.
Wackernagel, H. (1995). Multivariate Geostatistics: An Introduction with Applications. Springer-Verlag, Berlin.
Wijsman, E. M. and Cavalli-Sforza, L. L. (1984). Migration and genetic structure with special reference to humans. Ann Rev Ecol Syst, 15: 279–301.
Zanetto, A. and Kremer, A. (1995). Geographical structure of gene diversity in Quercus petraea (Matt.) Liebl. I. Monolocus patterns of variation. Heredity, 75: 506–517.
Zanetto, A., Roussel, G. and Kremer, A. (1994). Geographic variation of inter-specific differentiation between Quercus robur L. and Quercus petraea (Matt.) Liebl. Forest Genet, 1: 111–123.
Acknowledgements
The research was supported by an ONF (Office National des Forêts) grant and by an EEC grant of DG12 (Programme Forest no. MA2B-CT91–0022) ‘Description of the genetic variation in oak populations by means of molecular markers and adaptive traits’. We thank Sandor Bordacz, Kornel Burg, Gerard Burzinski, Valeriu Enescu, Dusan Gomory, Jaroslav Koblizek, Armin König, Csaba Matyas, Ladislov Paule, Parel Radosta, Stelian Radu and the French National Forest Service (ONF) for providing the material. The authors also thank Alexis Ducousso and Laurent Salera for their assistance in collecting the material. Dr Rémy Petit provided useful comments about the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Le Corre, V., Roussel, G., Zanetto, A. et al. Geographical structure of gene diversity in Quercus petraea (Matt.) Liebl. III. Patterns of variation identified by geostatistical analyses. Heredity 80, 464–473 (1998). https://doi.org/10.1046/j.1365-2540.1998.00313.x
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00313.x
Keywords
This article is cited by
-
Performance and genetic diversity of 23 provenances of northern red oak (Quercus rubra L.) after 25 years of growth in South Korea
Journal of Forestry Research (2021)
-
Complex fine-scale spatial genetic structure in Epidendrum rhopalostele: an epiphytic orchid
Heredity (2019)
-
Effect of a red oak species gradient on genetic structure and diversity of Quercus castanea (Fagaceae) in Mexico
Tree Genetics & Genomes (2014)
-
Diversity of Ullucus tuberosus (Basellaceae) in the Colombian Andes and notes on ulluco domestication based on morphological and molecular data
Genetic Resources and Crop Evolution (2012)
-
Trend der Schwermetall-Bioakkumulation 1990 bis 2005: Qualitätssicherung bei Probenahme, Analytik, geostatistischer Auswertung
Environmental Sciences Europe (2009)