Abstract
Begonia is one of the ten largest plant genera, with over 1500 species. This high species richness may in part be explained by weak species cohesion, which has allowed speciation by divergence in allopatry. In this study, we investigate species cohesion in the widespread Central American Begonia heracleifolia and Begonia nelumbiifolia, by genotyping populations at microsatellite loci. We then test for post-zygotic reproductive barriers using experimental crosses, and assess whether sterility barriers are related to intraspecific changes in genome size, indicating major genome restructuring between isolated populations. Strong population substructure was found for B. heracleifolia (FST=0.364, F′ST=0.506) and B. nelumbiifolia (FST=0.277, F′ST=0.439), and Bayesian admixture analysis supports the division of most populations into discrete genetic clusters. Moderate levels of inferred selfing (B. heracleifolia s=0.40, B. nelumbiifolia s=0.62) and dispersal limitation are likely to have contributed to significant genetic differentiation (B. heracleifolia Jost’s D=0.274; B. nelumbiifolia D=0.294). Interpopulation crosses involving a divergent B. heracleifolia population with a genome size ∼10% larger than the species mean had a ∼20% reduction in pollen viability compared with other outcrosses, supporting reproductive isolation being polymorphic within the species. The population genetic data suggest that Begonia populations are only weakly connected by gene flow, allowing reproductive barriers to accumulate between the most isolated populations. This supports allopatric divergence in situ being the precursor of speciation in Begonia, and may also be a common speciation mechanism in other tropical herbaceous plant groups.
Similar content being viewed by others
Introduction
Species cohesion is maintained by gene flow (Slatkin, 1987; Morjan and Rieseberg, 2004). If gene flow between populations is disrupted, populations become genetically isolated from one another, and reproductive barriers may subsequently evolve leading to speciation (Coyne and Orr, 2004; Duminil et al., 2007; Martin and Willis, 2010). Studying this trajectory, from panmictic population to genetically differentiated species with strong reproductive barriers, provides insights into the processes of speciation (Coyne and Orr, 1989). Many factors will influence this trajectory, some promoting genetic differentiation between populations, others maintaining species cohesion in spite of reduced gene flow (Wright, 1943).
One major factor influencing cohesion and differentiation processes in plants is the rate of interpopulation gene flow (migration) mediated by pollen and seed dispersal. The extent of pollen- and seed-mediated gene flow in angiosperms can be estimated by comparing genetic structure at maternally inherited chloroplast loci and biparentally inherited nuclear loci (Ennos, 1994; Petit et al., 2005), which experience different levels of gene flow (Petit and Excoffier, 2009). Pollen is the main source for gene exchange between populations, as it is normally dispersed over greater distances than seeds (Ennos, 1994; Petit and Excoffier, 2005; Bacles et al., 2006, but see exceptions such as Anthoxanthum odoratum L., Freeland et al., 2011). Pollen dispersal may maintain species cohesion by homogenizing population differences at selectively neutral genes, or by transferring alleles for key traits that are under strong selection. By this means, genome-wide divergence may occur, with little differences at important quantitative trait loci (Morjan and Rieseberg, 2004; Strasburg et al., 2012). Although seed dispersal has a smaller role in gene flow, it is crucial in colonization and subsequent expansion of a species range (Petit et al., 2003). Therefore, studying genetic structure at both nuclear and chloroplast loci in related species allows us to identify common patterns of population dynamics and species cohesion within a group of interest.
Mating systems, through their influence on pollen flow, affect patterns of gene exchange between populations (Hamrick and Godt, 1996). Obligate outbreeding species, such as dioecious or self-incompatible species, maintain higher levels of genetic exchange among populations (Charlesworth, 2003), than populations of species that self-fertilize (Hamrick and Godt, 1996). Mating systems also influence the effective population size (Ne), such that Ne in a fully selfing population is half that in an equivalent outcrossing population. Thus, an increased rate of selfing, through its effects on both interpopulation gene exchange and effective population sizes, will enhance the geographic structuring of genetic diversity that is the precursor of reproductive isolation and speciation (Hamrick and Godt, 1996; Lasso et al., 2011).
While many factors, as outlined above, will promote genetic divergence between populations (Charlesworth, 2009), speciation will only result if gene flow among populations is limited by reproductive barriers (Rieseberg and Blackman, 2010). Although both pre- and post-zygotic barriers contribute to reproductive isolation, recent work has shown that post-zygotic barriers can evolve rapidly (Widmer et al., 2009; Jewell et al., 2012) and therefore will be polymorphic within a species (Scopece et al., 2010) before lineages split. These include sterility barriers that may evolve as a by-product of allopatric divergence, either through selection on few adaptive loci for important phenotypes (‘speciation genes’ see Wu, 2001) or by fixing allelic differences at neutral or adaptive loci that negatively interact between divergent lineages (Dobzhansky–Muller incompatibilities). Isolation between incipient species may also be caused by chromosomal changes, such as large inversions or deletions, or whole-genome duplications. Studies of incipient allopatric speciation will therefore be most informative if they jointly estimate the degree of genetic differentiation between populations, as well as whether genetic incompatibilities have evolved as a by-product of divergence (Scopece et al., 2010). Cytological or genome size data can then be used to test whether genome restructuring is correlated with reproductive isolation, or quantitative trait locus mapping used to test the genetic architecture of population-level differences.
Understanding modes of speciation is of particular importance in the tropics, given the extremely high levels of species diversity and endemism. Baker (1959) and Fedorov (1966) proposed a model of speciation for tropical trees, whereby high levels of self-fertilization occur due to low population densities, which restricts gene flow between populations, and results in allopatric divergence (Lasso et al., 2011). Although this model has since been largely rejected for trees (but see Dick et al., 2008) due to numerous molecular studies showing frequent long-distance gene flow and low levels of self-fertilization (for example, Chase et al., 1999), a recent study of the genus Piper (Lasso et al., 2011) has revisited this theory for shrubs and herbaceous plants. The authors found a pattern of fine-scale spatial genetic structure over short spatial scales, and strong population substructure over larger geographic areas, suggesting attributes of herbs and shrubs (such as limited pollen and seed dispersal and self-fertilization) may promote allopatric speciation.
A particularly suitable system to test modes of speciation in the tropics is the genus Begonia. Population genetic analyses have shown strong populations substructure in a number of African species (for example, Begonia sutherlandii, FST=0.485, F′ST=0.896; Hughes and Hollingsworth, 2008; Begonia dregei, FST=0.882; Begonia homonyma, FST=0.937; Matolweni et al., 2000). At the species level, phylogenetic work has shown geographically constrained monophyly of species radiations (Forrest and Hollingsworth, 2003; Thomas et al., 2011). This suggests that microevolutionary processes of high genetic differentiation, caused by limited dispersal and inbreeding, may be driving macroevolutionary patterns of recurrent speciation and frequent endemism (Hughes and Hollingsworth, 2008). These patterns can be studied in more detail using recently developed molecular resources (Brennan et al., 2012), and using conventional genetic approaches that exploit the cross-compatibility and relatively short life-cycle of Begonia species (Dewitte et al., 2011).
Here, we investigate whether there is evidence of incipient speciation between populations in two widespread Central American Begonia species. B. heracleifolia Cham. and Schltdl. and B. nelumbiifolia Cham. and Schltdl. are genotyped at nine nuclear microsatellite markers to estimate genetic differentiation. The data from nuclear markers are compared with plastid data (Twyford et al., 2013b), to infer the ratio of interpopulation pollen and seed dispersal. The co-dominant data are also used to infer the level of selfing, which may further contribute to genetic differentiation. We then test whether genetic incompatibilities have accumulated between differentiated populations, by looking at the fertility of crosses between populations. The hybrid sterility data are related to population differences in genome sizes, to test whether genome restructuring is associated with hybrid sterility. The joint genetic, crossing and genome size data are used to test whether allopatric divergence in situ within Begonia species could be the precursor of allopatric speciation.
Materials and methods
Study species
B. nelumbiifolia and B. heracleifolia were chosen because they are two of the most widespread Central American Begonia species in a genus of mostly narrowly distributed endemics (Hughes and Hollingsworth, 2008). They are both found throughout Mexico and their ranges extend south into Central America (B. heracleifolia to Honduras; B. nelumbiifolia to Columbia, Burt-Utley, 1985). Although they occur over similar geographic ranges, the species differ in the degree of morphological differentiation exhibited among populations. B. nelumbiifolia is relatively uniform in morphology across its range. In contrast, B. heracleifolia is highly variable in leaf shape and leaf colour, and many regional varieties have been recognized (Burt-Utley, 1985). Studying genetic differentiation and reproductive isolation in such a species may provide an insight into the early stages of incipient speciation. The species are easily distinguished from related Begonia taxa by their leaf and flower morphology (Burt-Utley, 1985). They also differ in their ecologies, with B. nelumbiifolia growing in moist shaded areas and B. heracleifolia in dry sun-exposed areas. These species typically occur in small isolated populations, although they can be locally abundant and form dense stands (Twyford, personal observation). Both species can be propagated by splitting rhizomes, allowing them to be easily transported, and grown in cultivation to be used in experimental crosses.
Sampling
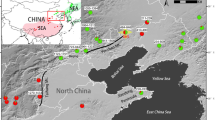
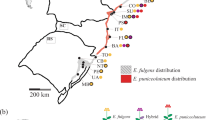
To test the patterns of genetic diversity and differentiation, an average of 25 individuals were sampled from each of 13 populations of B. heracleifolia, and 7 populations of B. nelumbiifolia (Table 1). Samples of B. nelumbiifolia were taken from the Mexican Gulf region, whereas samples of B. heracleifolia were made over a broader sampling distance from South Mexico to Guatemala (Figure 1). A representative specimen from each population was placed in the herbarium in Edinburgh (E) except for the sample from the Guatemalan population, which was sent to the University of San Guatemala (BIGU).
Genotyping
DNA extraction from silica dried material was performed using a modified protocol for the DNeasy 96-sample kit (Qiagen, Germantown, MD, USA), described in Twyford et al. (2013a). A preliminary test of nuclear microsatellite amplification was made with the 14 loci listed in Twyford et al. (2013a). The nine loci that amplified uniformly across species and populations were then used for the full genotyping of the population samples, with the same PCR protocol and amplification programme. Input files for genetic analysis were converted between formats with the programme CREATE (Coombs et al., 2008). To test whether data from microsatellite loci are independent of each other, linkage disequilibrium between markers was tested in FSTAT 1.2 (Goudet, 1995).
Begonia species can reproduce asexually, and clumps of clonal individuals may arise by rhizomatous growth or from vegetative material being broken off and rooting. The probability that individuals in a population shared identical genotypes at all nine loci through random mating was calculated using the approach of Parks and Werth (1993). Genotypes not considered a product of sexual mating (Psex) at P<0.01 when non-random mating was allowed (positive FIS value), were removed from analyses. Individuals from populations where putative hybrids were found were only included if they were confidently considered pure species (that is, assigned to a ‘pure’ parental class with at least 95% probability in New Hybrids, Anderson and Thompson, 2002; Twyford unpublished data). This approach was validated as removing populations where hybrids occurred did not affect values of F-statistics (results not shown).
Measures of genetic diversity
Diversity statistics were calculated per locus and per population in FSTAT v1.2 (Goudet, 1995). The statistics calculated were: number of alleles (A, calculated per locus only), gene diversity (hS, Nei, 1987), and allelic richness corrected for sample sizes (Ae, rarefaction method, Mousadik and Petit, 1996). Allelic richness was calculated only for populations where at least 10 individuals were scored for each locus, after considering missing data. The number of private alleles per population was scored manually and private allelic richness (pAe) calculated by rarefaction using HP-RARE (Kalinowsi, 2005).
Measures of inbreeding
FIS values were calculated per locus and per population, with significance values calculated by jackknifing (per locus) or randomization test (per population) in FSTAT v1.2. For each species, the inferred selfing rate was calculated from the inbreeding coefficient, using the formula of Allard et al. (1969):
S=2FIS/(1+FIS)
This approach does not take into account bi-parental inbreeding, which was not directly addressed in this study.
Self-compatibility was confirmed using greenhouse-grown plants. One cultivated accession of B. nelumbiifolia and five accessions of B. heracleifolia from different populations were self-fertilized. Seeds were germinated in 9-cm pots of finely sieved bark, which were kept in a propagator at 25 °C, and germination was recorded after 6 weeks.
Measures of population structure and genetic differentiation
FST statistics were calculated between sampling sites for each species, as well as between the distinct genetic clusters identified in the BAPS analysis (see below). FST statistics were also calculated for B. heracleifolia populations in the Mexican Gulf area (h2, h3, h5, h15, h21, h23, h26) to allow like-for-like comparisons between the two species over a similar geographic area. Weir and Cockerham’s (1984) estimator of FST, which is a measure of the genetic structure in the data, was calculated in FSTAT v1.2, along with 95% confidence intervals across loci. A standardized measure of population structure that takes into account sample sizes and allelic diversity (F ′ST) was calculated by using RecodeData v0.1 (Meirmans, 2006) and FSTAT. Absolute differentiation was measured with D (Jost, 2008) using SMOGD (Crawford, 2010), and confidence intervals calculated by bootstrapping with 1000 replicates. An FST analogue, which incorporates allele size length (RST) was calculated with SPAGeDI v1.3 (Hardy and Vekemans, 2002). We inferred whether there is phylogeographical signal in the data by testing for a significant difference between RST and FST by permutation test in SPAGeDI. The between-species FST was also calculated, which shows the similarities in allele frequencies between the species.
The ratio of interpopulation pollen–to-seed dispersal can be estimated using formula 5a of Ennos (1994), which relates the FST for biparentally inherited nuclear markers (FST(b)) and maternally inherited plastid markers (FST(m)), as well as the level of inbreeding (FIS). Maternal inheritance of plastids in Begonia have been confirmed by cytological observations (Corriveau and Coleman, 1988) and sequencing plastid DNA in experimental crosses (Peng and Chiang, 2000). No variation was found at seven plastid microsatellite markers in B. nelumbiifolia (Twyford et al., 2013b), so the pollen-to-seed ratio could not be calculated for this species. For B. heracleifolia, FST(m) was calculated from the plastid microsatellite data in Twyford et al. (2013b) using the same populations sampled for the nuclear microsatellites, and compared with the FST(b) and the FIS.
To test whether dispersal between populations fits a simple model of dispersal limitation between more distant populations (the stepping stone model of dispersal, Kimura and Weiss, 1964), the relationship between genetic similarity and geographic distance was tested using isolation-by-distance analysis. Pairwise comparisons of FST/(1-FST) for each population were plotted against the natural logarithm of geographic distance as suggested by Rousset (1997), and implemented in the Isolation by Distance Web Service v3.21 (Jensen et al., 2005).
We used the Bayesian clustering programme BAPS (Corander et al., 2008) to visualize the spatial structuring of populations. The ‘clustering of groups of individuals’ setting was used, and the number of genetic clusters (K) evaluated was K=1–13 for B. heracleifolia and K=1–7 for B. nelumbiifolia. Five replicates were made for each K value, and the results file was then used as the input for admixture analysis using the ‘mixture clustering option’. The minimum size of each population was set to three individuals, and runs were made of 10 000 iterations, and 5000 reference individuals were used. The optimum value of K is automatically calculated in BAPS using a greedy stochastic optimization algorithm (Corander et al., 2008). The BAPS admixture bar plots produced by the programme were used to display the results, with only significant admixture shown (P<0.05).
We confirmed our Bayesian clustering results from BAPS by re-analysing our data in InStruct (Gao et al., 2007), which accounts for deviations from Hardy–Weinberg caused by inbreeding (see Results), as well as STRUCTURE (Pritchard et al., 2000), using the following settings. Runs of 100 000 generations were performed following a burn-in of 100 000 generations. K values between 1 and 13 were evaluated for B. heracleifolia, and 1 and 7 for B. nelumbiifolia, with 10 independent replicates per K value. The ad hoc statistic ΔK was calculated across runs, and the greatest value inferred to be the optimal K value (Evanno et al., 2005). For analyses of a given K value, a consensus file correcting for equally optimal solutions (multi-modality) and label switching was produced in CLUMPP (Jakobsson and Rosenberg, 2007), and the results displayed in DISTRUCT (Rosenberg, 2004).
To visualize the relationship between populations, a neighbour-joining tree was constructed for each species. Allele frequencies from FSTAT were used as the input for POPTREE2 (Takezaki et al., 2010), where a neighbour-joining tree was constructed using the distance measure of Nei et al. (1983). The tree was edited in FigTree v1.2.2 (available from http://www.tree.bio.ed.ac.uk/software/figtree/).
Measures of reproductive isolation
To gain a first insight into reproductive isolation between differentiated populations of B. heracleifolia, pollen fertility was scored in artificial interpopulation crosses. We then tested whether reduced pollen fertility is associated with large-scale chromosome reorganization events, by comparing pollen viability results to a survey of genome sizes in representative individuals from a subset of populations.
Three groups of crosses were made (selfs, close outcrosses and wide outcrosses, herein), and pollen sterility compared between the groups. First, plants from South Oaxaca (population h28), which was found to be genetically divergent from populations in the Mexican Gulf (see Results), were used in wide outcrosses. Two individuals were selected, and used as pollen recipients from populations in the Mexican Gulf chosen at random. These crosses were compared with outcrosses between geographically proximal populations (close outcrosses) as well as selfed crosses, given that self-pollination may be common in the wild (see Results). Six maternal parents from different populations across the Gulf of Mexico were selected, and each used in self-pollinations, and outcrosses with plants from other populations. For each cross, the pollen donor was selected at random from the plants in flower. Crosses were performed by rubbing the dehiscing anthers against the stigma. Flowers were labelled and surrounding flowers removed to prevent cross-contamination of pollen. Seed capsules were harvested at maturity and stored at 4 °C. Seeds were germinated on finely sieved bark, and 6-week-old seedlings were transferred to 9-cm pots in sterilized potting mix (16 bark: 3 peat: 1 perlite plus finely sieved osmocote) and grown at 28 °C to flowering. Pollen sterility was measured by acetocarmine staining, which is a reliable method for viability assessment in Begonia (Dewitte et al., 2011), and corresponds well to artificial pollen germination and fluorescent staining (Twyford and Kidner, unpublished data). Dehiscing pollen from one flower per plant was stained with 1 M acetocarmine, visualized under a Leica Microscope, and the proportion of well-stained pollen recorded out of 200 pollen grains.
Fully expanded leaf material from four accessions of B. nelumbiifolia and five of B. heracleifolia were selected from different populations (Table 3) for preliminary evaluation of intraspecific nuclear DNA content (C-value) variation by flow cytometry, following the procedure of Brennan et al. (2012). Two to four technical replicates per accession were processed and the resulting fluorescence histograms were analysed with FlowMax software (Partec GmbH, Münster, Germany). The mean and standard error for each individual, and per species, were calculated.
Results
Genetic diversity
A total of 306 individuals from B. heracleifolia and 177 from B. nelumbiifolia were genotyped. Five individuals of B. heracleifolia were identical to another at nine polymorphic loci, and are likely to be clones (P<0.01), so were removed from the data set (Table 1).
Descriptive statistics are summarized per population in Table 1 and per locus in Table 2. All loci were polymorphic, except B5347, which was monomorphic in B. nelumbiifolia. Overall, the levels of genetic diversity were low for both B. heracleifolia (mean values across populations: A=6.6; Ae=2.274, hS=0.276) and B. nelumbiifolia (A=4.7; Ae=2.569, hS=0.417). Seventeen of the 54 alleles (29.8%) detected in B. heracleifolia were private alleles, as were nine of the 44 alleles (20.4%) in B. nelumbiifolia.
Inbreeding
The average FIS value across loci was 0.249 (s.e. 0.062) and 0.454 (s.e. 0.095) for B. heracleifolia and B. nelumbiifolia, respectively. FIS values varied from 0.01 in population h2 to 0.684 in population h13 (B. heracleifolia), and 0.306 in population n26 to 0.535 in population n18 (B. nelumbiifolia), see Table 2. The values for the inferred selfing rate (s) calculated from the inbreeding coefficient FIS, were 0.399 for B. heracleifolia, and 0.624 for B. nelumbiifolia. Seed set was high (>90%) in the self-pollination experiment with B. heracleifolia (n=5) and B. nelumbiifolia (n=1).
Population structure and genetic differentiation
Significant population substructure was found by FST analysis (average across loci, B. heracleifolia, FST=0.364; s.e. 0.028, 95% confidence interval 0.315–0.423; P<0.05; B. nelumbiifolia, FST=0.277; s.e. 0.055, 95% confidence interval 0.181–0.380; P<0.05), as well as when the FST was standardized for the maximum possible value for the loci sampled (B. heracleifolia, F ′ST=0.506, B. nelumbiifolia, F ′ST=0.439). Moderate levels of differentiation were found with Jost’s estimator of absolute differentiation D, with average values per locus for B. heracleifolia (D=0.274, P<0.05), and for B. nelumbiifolia (D=0.294, P<0.05). Recalculating measures of population substructure and genetic differentiation for the genetic clusters identified in the BAPS analysis for B. heracleifolia had little effect (results not shown). RST values, which use allele size lengths and a stepwise mutation model, were significantly different from 0 (B. heracleifolia RST=0.212; B. heracleifolia RST=0.257), but were not significantly larger than the GST values (not shown), suggesting the absence of phylogeographical structure. Isolation-by-distance accounted for a modest amount of the genetic variance (B. heracleifolia, R2=0.250, P=0.001; B. nelumbiifolia, R2=0.289, P=0.016). The FST value between species was 0.466 (P<0.01). When the B. heracleifolia populations over a similar spatial scale as B. nelumbiifolia were considered, values of genetic differentiation and population substructure were reduced (FST=0.233, s.e. 0.04; F ′ST=0.339; D=0.164, P<0.05).
When the FST value (FST(m)=0.728) from the highly polymorphic plastid microsatellites (total 39 haplotypes, mean 5.2 alleles per locus, hs=0.44, Twyford et al., 2013b) was related to the FST for nuclear microsatellites (FST(b) 0.364), while also considering inbreeding (FIS=0.249), the ratio of pollen-to-seed dispersal for B. heracleifolia was found to be 3.8. Transmission of genes through pollen is therefore ∼4 times more effective than through the seed.
The most likely number of genetic clusters for B. heracleifolia in the BAPS analysis was K=11. The genetic clusters were the same as the sampled populations, with two exceptions (Figure 2a). Populations h3 and h5, which are separated by less than 2 km, shared a common gene pool. BAPS assigned population h13, which contained only six individuals, as admixed between populations h3, h5, and h8 (270 km away). BAPS revealed seven main clusters corresponding to the seven populations genotyped for B. nelumbiifolia (Figure 2b). Levels of admixture were low, with nine individuals (5%) having admixed ancestry. Analyses in STRUCTURE and InStruct gave broadly consistent results to BAPS (that is, same optimum number of clusters, and similar population groupings), but showed higher admixture (Supplementary Figures S1–S4).
Bayesian assignment to genetic clusters and the relationships between populations. Bayesian admixture results in BAPS for B. heracleifolia (a) and B. nelumbiifolia (b). Each individual is represented by a vertical bar, and different colours represent the different genetic cluster. Asterisk indicates population h13. Neighbour-joining trees of Nei’s (1983) measure of population divergence for B. heracleifolia (c) and B. nelumbiifolia (d). Branches are coloured to correspond with the genetic clusters from the BAPS analyses.
The average value for the pairwise population distance measures (Nei et al., 1983), which were used to build the neighbour-joining trees, were 0.231 (0.011s.e.) for B. heracleifolia and 0.201 (0.013s.e.) for B. nelumbiifolia. The tree for B. heracleifolia included two long branches, connecting populations h28 (mean pairwise DNei=0.360) and h-g1 (mean pairwise DNei=0.313; Figure 2c). Neither of these divergent populations (h28 and h-g1) showed an aberrant genetic signature at microsatellite loci (for example, multiple bands). The tree for B. nelumbiifolia was roughly star-shaped (Figure 2d).
Reproductive isolation
The mean pollen stainability of selfed B. heracleifolia plants was 97.5% (s.e. 1.26, n=18), and only one plant had a value below 85%. Progeny from close outcrosses were similarly pollen-fertile, with a mean pollen stainability of 98.7% (s.e.=1.97, n=10). Progeny from wide outcrosses involving a B. heracleifolia individual from Oaxaca (h28) and populations from the Mexican Gulf (h8, h21, h24) had a mean viability of 78.3% (s.e.=1.70, n=3).
Our preliminary average 1C genome size estimates for B. heracleifolia and B. nelumbiifolia were 0.80 pg and 0.54 pg, respectively (Table 3). Genome size estimates were very consistent in replicates of each individual (results not shown), and across individuals within species, except for the B. heracleifolia individual from Oaxaca population h28 that had an estimated genome size of 0.88 pg, which is 10% higher than all other samples from the same taxon.
Discussion
Strong population substructure and genetic differentiation in widespread Begonia species
Dispersal limitation has left a clear genetic signature in populations of two widespread Central American Begonia species, B. heracleifolia and B. nelumbiifolia. This was evident from strong population substructure (B. heracleifolia FST=0.364; B. nelumbiifolia, FST=0.277), significant genetic differentiation (B. heracleifolia Jost’s D=0.274; B. nelumbiifolia D=0.294), and Bayesian structure analyses dividing most populations into discrete genetic clusters (Figure 2). These data suggest Begonia populations are isolated, with little homogenizing gene flow between them. These results are consistent with the high level of genetic differentiation at plastid loci for B. heracleifolia (Twyford et al., 2013b). Strong genetic differentiation is likely promoted by small census population sizes (∼10–500 individuals, AD Twyford personal observation), and significant selfing rates (described below), which contributes to differentiation by the joint effects on interpopulation gene flow and reduction in effective population size (Charlesworth, 2003).
In a broader context, genetically isolated Begonia populations have been inferred from population genetic analyses of three other Begonia species (B. sutherlandii, Hughes and Hollingsworth, 2008; B. dregei, B. homonyma, Matolweni et al., 2000). Each of these studies has inferred limited gene flow between populations, even over small spatial scales, which likely reflects the absence of mechanisms that promote dispersal. The joint use of plastid and nuclear markers in this study allowed us to infer an equilibrium ratio of pollen-to-seed flow for B. heracleifolia (r=3.8), which was at the lower end of the range seen in plants (for example, r=4–196, Ennos, 1994). Similarly, low pollen-to-seed ratios (r=4.0) have been found for other herbaceous plants such as Dysosma versipellis (Hance) M. Cheng ex T.S. Ying (Berberidaceae) (Guan et al., 2010), or other plant species that grow below the forest canopy, such as the epiphytic bromeliad Vriesea gigantea Lem. (Bromeliaceae) (r=3.3, Palma-Silva et al., 2009). Overall, results of population structure across Begonia species suggest a common set of population-level mechanisms (genetic drift in isolated populations) have a role in the evolution of intraspecific genetic diversity in the genus. This supports the prevailing view that the partitioning of genetic diversity between populations is often conserved among related species, due to shared ecological attributes and dispersal traits (Duminil et al., 2007).
The FST values among populations of B. heracleifolia and B. nelumbiifolia, as well as those for other Begonia species (Matolweni et al., 2000; Hughes and Hollingsworth, 2008), indicate a higher level of population structure than most other plant species (Gitzendanner and Soltis, 2000; Hey and Pinho, 2012). Particularly high pairwise FST values were found in this study between populations separated by major geographic barriers, such as the Southern Oaxacan population h28 (mean pairwise FST=0.463, range 0.342–0.638), isolated from other populations by the Sierra Madre del Sur. The geographic structuring of genetic diversity across this barrier has been found in other organisms, such as the bush-tanager (García-Moreno et al., 2004), the bark beetle Dendroctonus mexicanus Hopkins. (Anducho-Reyes et al., 2008) and the western lyre snake (Devitt, 2006). However, relatively high pairwise FST values were also found between populations in close proximity across semi-continuous habitats. Plant species with higher FST values distributed over similarly large geographic areas are typically much more patchy in their distribution, or are highly selfing. One example of this is the bromeliad Pitcairnia geyskesii L.B.Sm., where there is strong population structure (FST=0.533) between the large emergent rocky outcrops (inselbergs) where the species grows in French Guiana (Boisselier-Dubayle et al., 2010). Selfing species with higher FST values than Begonia species include Arabidopsis thaliana L. (FIS=0.969, FST=0.61, Bomblies et al., 2010); Bromus tectorum L. (FIS=1, FST=0.53; Ramakrishnan et al., 2006) and Medicago truncatula Gaertn. (FIS=0.978, FST=0.3–0.75, Siol et al., 2008).
Inbreeding promotes genetic differentiation
Both Begonia species were found to be fully self-compatible, with inferred equilibrium selfing rates of 40% for B. heracleifolia and 62% for B. nelumbiifolia. Marked population variation in inbreeding also occurs, as seen in B. heracleifolia (Table 1). The mechanisms underlying inbreeding in Begonia are not currently clear; it may either be autopollination or self-fertilization mediated by insect pollinators (geitonogamy). Most Begonia species are self-compatible (Ågren and Schemske, 1993; Dewitte et al., 2011; Wyatt and Sazima, 2011; Twyford and Kidner, unpublished data), with only two species studied to date not setting seed in a small number of experimental crosses (Brazilian Begonia integerrima Spreng. and Begonia itatinensis Irmsch. Ex Brade; Wyatt and Sazima, 2011). Levels of inbreeding in Begonia species varies, from close to panmictic (B. sutherlandii, mean FIS=0.158, seven microsatellites) to fully selfing (B. hirsuta single-locus outcrossing rate 0.03±0.01, 1 isozyme locus, Ågren and Schemske, 1993). Most Begonia species are monoecious and functionally protandrous, producing male flowers on an inflorescence before females (Forrest and Hollingsworth, 2003). However, as many inflorescences are borne over a flowering season, there are plenty of opportunities for self-pollination. This would particularly be the case for species such as B. nelumbiifolia, which produces many densely-packed inflorescences that can easily become intertwined. Self-pollination in such a way would assure mating success if pollinators are rare, which may be expected in the isolated populations in which Begonia species typically grow.
The inferred equilibrium selfing rates estimated here are upper estimates, and actual values may be lower due to technical artefacts of the loci sampled, and the population substructure. A positive FIS value can occur due to null alleles (Pemberton et al., 1995). This, however, seems unlikely given that few individuals could not be amplified for a given locus (null homozygotes), and considering primers were designed from transcriptome sequence data (Brennan et al., 2012) and have been shown to be conserved over a broad phylogenetic range (Twyford et al., 2013a), suggesting interpopulation polymorphisms in the primer regions are unlikely. Another factor contributing to the positive FIS may be the strong population substructure (discussed above). If sampling areas are large and span multiple populations that seldom interbreed, a positive FIS value may be obtained when the populations are ‘lumped’ together. This was considered a contributing factor to the positive FIS in a study of Begonia sutherlandii (Hughes and Hollingsworth, 2008), a species with very strong population substructure. We expect the contribution of population substructure to the FIS of the Begonia species we sampled here to be relatively small, as populations tended to be dense and continuous, and the FIS value was relatively consistent across loci and populations. Finally, the positive FIS value may actually be a product of bi-parental inbreeding, rather than self-fertilization. This would seem possible given the limited seed dispersal range, allowing many related individuals to grow in a particular patch. The presence of bi-parental inbreeding could be tested by comparing single-locus and multi-locus outcrossing rates in progeny arrays (Ritland, 2002, and references therein).
Reproductive isolation
In this study, we have identified intraspecific variation for post-zygotic reproductive isolation within a Begonia species. We showed an ∼20% reduction in pollen stainability in experimental crosses between divergent populations in B. heracleifolia. Moreover, we could relate this to cryptic genome size variation within the species, as the Oaxacan population had an ∼10% higher C-value relative to the species mean (Table 3). Although our C-value estimates are preliminary data based on few samples, the significantly higher value for population h28, low variance between other populations and the relationship to pollen sterility in crosses, is a striking result. These results support populations being isolated from homogenizing gene flow for sufficient time to allow diverge on a potential route towards speciation. The timing of which the Southern Oaxacan population has been isolated is currently uncertain, as this area has a complex geographic history, with mountain building occurring gradually since the late Cretaceous until the early Holocene (Anducho-Reyes et al., 2008, and references therein).
The evolution of reproductive isolation within species has only been studied to a limited extent, and has been identified as an important future research direction (Lexer and Widmer, 2008; Scopece et al., 2010). There are even fewer studies that have investigated intraspecific variation for reproductive isolation in species-rich tropical taxa. Pinheiro et al. (2013) reported polymorphic post-zygotic reproductive isolation between populations of Epidendrum denticulatum Barb.Rodr. (Orchidaceae) in South America, with greatly reduced seed set between individuals with different chloroplast haplotypes. They suggest variation in crossing results between populations may be a product of many loci of small effect. In contrast, the consistency between reduced pollen stainability in crosses and population differences in genome size suggests large-scale genome restructuring may be involved in this pattern of isolation in Begonia. Future studies should look at more pairwise comparisons of fertility and sterility, as well as estimate genome sizes in more populations (also sampling intrapopulation genome size variation), to better understand the relationship between divergence and reproductive isolation in B. heracleifolia.
Conclusion
The mean FST values for these species, and the other Begonia species studied to date, approach or exceed the threshold value of FST=0.35 suggested by Hey and Pinho (2012) for delimiting species. In addition to this strong geographic structure caused by dispersal limitation, inbreeding may further reduce the level of gene flow between populations and promote divergence. The joint role that genetic drift and inbreeding have on levels of genetic differentiation in species-rich tropical plant lineages is now beginning to be appreciated (the Baker–Fedorov hypothesis, Lasso et al., 2011). Moreover, we have shown the early stages of reproductive isolation between Begonia populations, supporting the hypothesis that differentiation leads to the accumulation of incompatibilities that may be involved in reproductive isolation, in a trajectory towards allopatric speciation.
Data archiving
Microsatellite genotype data and pollen viability scores have been submitted to Dryad: doi:10.5061/dryad.kf1td.
References
Ågren J, Schemske DW . (1993). Outcrossing rate and inbreeding depression in two annual monoecious herbs, Begonia hirsuta and B. semiovata. Evolution 47: 125–135.
Allard RW, Jain SK, Workman PL . (1969). Genetics of inbreeding populations. Adv Genet 14: 55–131.
Anderson EC, Thompson EA . (2002). A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160: 1217–1229.
Anducho-Reyes MA, Cognato AI, Hayes JL, Zúñiga G . (2008). Phylogeography of the bark beetle Dendroctonus mexicanus Hopkins (Coleoptera: Curculionidae: Scolytinae). Mol Phylogenet Evol 49: 930–940.
Bacles CFE, Lowe AJ, Ennos RA . (2006). Effective seed dispersal across a fragmented landscape. Science 311: 628.
Baker HG . (1959). Reproductive methods as factors in speciation in flowering plants. Cold Spring Harb Symp Quant Biol 24: 177–199.
Boisselier-Dubayle MC, Leblois R, Samadi S, Lambourdière J, Sarthou C . (2010). Genetic structure of the xerophilous bromeliad Pitcairnia geyskesii on inselbergs in French Guiana—a test of the forest refuge hypothesis. Ecography 33: 175–184.
Brennan AC, Bridgett S, Ali MS, Harrison N, Matthews A, Pellicer J et al. (2012). Genomic resources for evolutionary studies in the large, diverse, tropical genus. Begonia. Trop Plant Biol 5: 261–276.
Bomblies K, Yant L, Laitinen RA, Kim ST, Hollister JD, Warthmann N et al. (2010). Local-scale patterns of genetic variability, outcrossing, and spatial structure in natural stands of Arabidopsis thaliana. PLoS Genet 6: e1000890.
Burt-Utley K . (1985). A revision of Central American species of Begonia section Gireoudia (Begoniaceae). Tulane Studies in Zoology and Botany 25: 1–131.
Charlesworth B . (2009). Effective population size and patterns of molecular evolution and variation. Nat Rev Genet 10: 195–205.
Charlesworth D . (2003). Effects of inbreeding on the genetic diversity of populations. Philos Trans R Soc Lond B Biol Sci 358: 1051–1070.
Chase MR, Moller C, Kesseli R, Bawa KS . (1999). Distant gene flow in tropical trees. Nature 383: 398–399.
Coombs JA, Letcher BH, Nislow KH . (2008). Create: a software to create input files from diploid genotypic data for 52 genetic software programs. Mol Ecol Resources 8: 578–580.
Corander J, Marttinen P, Siren J, Tang J . (2008). Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinformatics 9: 539.
Corriveau JL, Coleman AW . (1988). Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 Angiosperm species. Am J Bot 75: 1443–1458.
Coyne JA, Orr AO . (1989). Patterns of speciation in Drosophila. Evolution 43: 362–381.
Coyne JA, Orr AO . (2004) Speciation. Sinauer associates: Sunderland.
Crawford NG . (2010). SMOGD: software for the measurement of genetic diversity. Mol Ecol Resources 10: 556–557.
Dewitte A, Twyford AD, Thomas DC, Kidner CA, Van Huylenbroeck J . (2011). The origin of diversity in Begonia: genome dynamism, population processes and phylogenetic patterns. Grillo, O, Venora, G (eds) The dynamical processes of biodiversity - case studies of evolution and spatial distribution. InTech: Rijeka, Croatia.
Devitt TJ . (2006). Phylogeography of the Western Lyre snake (Trimorphodon biscutatus): testing aridland biogeographical hypotheses across the Nearctic–Neotropical transition. Mol Ecol 15: 4387–4407.
Dick CW, Hardy OJ, Jones FA, Petit RJ . (2008). Spatial scales of pollen and seed-mediated gene flow in tropical rain forest trees. Trop Plant Biol 1: 20–33.
Duminil J, Fineschi S, Hampe A, Jordano P, Salvini D, Vendramin GG et al. (2007). Can population genetic structure be predicted from life-history traits? Am Nat 169: 662–672.
Ennos RA . (1994). Estimating the relative rates of pollen and seed migration among plant populations. Heredity 72: 250–259.
Evanno G, Regnaut S, Goudet J . (2005). Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14: 2611–2620.
Fedorov AA . (1966). The structure of the tropical rain forest and speciation in the humid tropics. J Ecol 54: 1–11.
Forrest LL, Hollingsworth PM . (2003). A recircumscription of Begonia based on nuclear ribosomal sequences. Plant Syst Evol 241: 193–211.
Freeland JR, Biss P, Silvertown J . (2011). Structure of sweet vernal grass (Anthoxanthum odoratum) populations. J Hered 103: 28–35.
Gao H, Williamson S, Bustamante CD . (2007). A Markov Chain Monte Carlo approach for joint inference of population structure and inbreeding rates from multilocus genotype data. Genetics 176: 1635–1651.
García-Moreno J, Navarro-Sigüenza AG, Peterson AT, Sánchez-González LA . (2004). Genetic variation coincides with geographic structure in the common bush-tanager (Chlorospingus ophthalmicus) complex from Mexico. Mol Phylogenet Evol 33: 186–196.
Gitzendanner MA, Soltis PS . (2000). Patterns of genetic variation in rare and widespread congeners. Am J Bot 87: 783–792.
Goudet J . (1995). FSTAT (version 1.2): A computer program to calculate F-statistics. J Hered 86: 485–486.
Guan B-C, Fu C-X, Qiu Y-X, Zhou S-L, Comes H . (2010). Genetic structure and breeding system of a rare understory herb, Dysosma versipellis (Berberidaceae), from temperate deciduous forests in China. Am J Bot 97: 111–122.
Hamrick J, Godt M . (1996). Effects of life history traits on genetic diversity in plants. Phil Trans R Soc Lond B Biol Sci 351: 1291–1298.
Hardy OJ, Vekemans X . (2002). SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2: 618–620.
Hey J, Pinho C . (2012). Population genetics and objectivity in species diagnosis. Evolution 66: 1313–1429.
Hughes M, Hollingsworth PM . (2008). Population genetic divergence corresponds with species-level biodviersity patterns in the large genus. Begonia. Mol Ecol 17: 2643–2651.
Jakobsson M, Rosenberg NA . (2007). CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23: 1801–1806.
Jensen J, Bohonak A, Kelley S . (2005). Isolation by distance, web service. BMC Genetics 6: 13.
Jewell C, Papineau AD, Freyre R, Moyle LC . (2012). Patterns of reproductive isolation in Nolana (Chilean Bellflower). Evolution 66: 2628–2636.
Jost L . (2008). GST and its relatives do not measure differentiation. Mol Ecol 17: 4015–4026.
Kalinowski ST . (2005). HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5: 187–189.
Kimura M, Weiss G . (1964). The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics 49: 561–576.
Lasso E, Dalling JW, Bermingham E . (2011). Strong spatial genetic structure in five tropical Piper species: should the Baker-Fedorov hypothesis be revived for tropical shrubs? Ecol Evol 1: 502–516.
Lexer C, Widmer A . (2008). The genic view of plant speciation: recent progress and emerging questions. Philos Trans R Soc London B Biol Sci 363: 3023–3036.
Martin NH, Willis JH . (2010). Geographical variation in post-zygotic isolation and its genetic basis within and between two Mimulus species. Philos Trans R Soc Lond B Biol Sci 365: 2469–2478.
Matolweni LO, Balkwill K, Mclellan T . (2000). Genetic diversity and gene flow in the morphologically variable, rare endemic Begonia dregei and Begonia homonyma (Begoniaceae). Am J Bot 87: 431–439.
Meirmans PG . (2006). Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 60: 2399–2402.
Morjan CL, Rieseberg LH . (2004). How species evolve collectively: implications of gene flow and selection for the spread of advantageous alleles. Mol Ecol 13: 1341–1356.
Mousadik A, Petit RJ . (1996). High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (l.) Skeels) endemic to Morocco. Theor Appl Genet 92: 832–839.
Nei M . (1987) Molecular evolutionary genetics. Columbia University Press: New York.
Nei M, Tajima F, Tateno Y . (1983). Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol 19: 153–170.
Palma-Silva C, Lexer C, Paggi G, Barbará T, Bered F, Bodanese-Zanettini M . (2009). Range-wide patterns of nuclear and chloroplast DNA diversity in Vriesea gigantea (Bromeliaceae), a Neotropical forest species. Heredity 103: 503–512.
Parks JC, Werth CR . (1993). A study of spatial features of clones in a population of bracken fern, Pteridium aquilinum (Dennstaedtiaceae). Am J Bot 80: 537–544.
Pemberton JM, Slate J, Bancroft DR, Barrett JA . (1995). Nonamplifying alleles at microsatellite loci: a caution for parentage and population studies. Mol Ecol 4: 249–252.
Peng C-I, Chiang T-Y . (2000). Molecular confirmation of unidirectional hybridization in Begonia x taipeiensis peng (Begoniaceae) from Taiwan. Ann Missouri Bot Gard 87: 273–285.
Petit RJ, Aguinagalde I, De Beaulieu J-L, Bittkau C, Brewer S, Cheddadi R et al. (2003). Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300: 1563–1565.
Petit RJ, Duminil J, Fineschi S, Hampe A, Salvini D, Vendramin GG . (2005). Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14: 689–701.
Petit RJ, Excoffier L . (2009). Gene flow and species delimitation. Trends Ecol Evol. 24: 386–393.
Pinheiro F, Cozzolino S, De Barros F, TMZM Gouveia, Suzuki RM, Fay MF et al. (2013). Phylogeographic structure and outbreeding depression reveal early stages of reproductive isolation in the Neotropical orchid Epidendrum denticulatum. Evolution 6: 2024–2039.
Pritchard JK, Stephens M, Donnelly P . (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Ramakrishnan AP, Meyer SE, Fairbanks DJ, Coleman CE . (2006). Ecological significance of microsatellite variation in western North American populations of Bromus tectorum. Plant Species Biol 21: 61–73.
Rieseberg LH, Blackman BK . (2010). Speciation genes in plants. Ann Bot 106: 439–455.
Ritland K . (2002). Extensions of models for the estimation of mating systems using n independent loci. Heredity 88: 221–228.
Rousset F . (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
Rosenberg NA . (2004). DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4: 137–138.
Scopece G, Lexer C, Widmer A, Cozzolino S . (2010). Polymorphism of postmating reproductive isolation within plant species. Taxon 59: 1367–1374.
Siol M, Prosperi JM, Bonnin I, Ronfort J . (2008). How multilocus genotypic pattern helps to understand the history of selfing populations: a case study in Medicago truncatula. Heredity 100: 517–525.
Slatkin M . (1987). Gene flow and the geographic structure of natural populations. Science 236: 787–792.
Strasburg JL, Sherman NA, Wright KM, Moyle LC, Willis JH, Rieseberg LH . (2012). What can patterns of differentiation across plant genomes tell us about adaptation and speciation? Phil Trans R Soc B 367: 364–373.
Takezaki N, Nei M, Tamura K . (2010). Poptree2: software for constructing population trees from allele frequency data and computing other population statistics with windows interface. Mol Biol Evol 27: 747–752.
Thomas DC, Hughes M, Phutthai T, Ardi WH, Rajbhandary S, Rubite R. et al. (2011). West to east dispersal and subsequent rapid diversification of the mega-diverse genus Begonia (Begoniaceae) in the Malesian Archipelago. J Biogeogr 39: 98–113.
Twyford AD, Ennos RA, Kidner CK . (2013a). Development and characterization of microsatellite markers for Central American Begonia section Gireoudia (Begoniaceae). Appl Plant Sci 1: 1200499.
Twyford AD, Kidner CK, Harrison N, Ennos RA . (2013b). Population history and seed dispersal in widespread Central American Begonia species (Begoniaceae) inferred from plastome-derived microsatellite markers. Bot J Linn Soc 171: 260–276.
Weir BS, Cockerham CC . (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Widmer A, Lexer C, Cozzolino S . (2009). Evolution of reproductive isolation in plants. Heredity 102: 31–38.
Wright S . (1943). Isolation by distance. Genetics 28: 114–138.
Wu C-I . (2001). The genic view of the process of speciation. J Evol Biol 14: 851–865.
Wyatt GE, Sazima M . (2011). Pollination and reproductive biology of thirteen species of Begonia in the Serra do Mar state park, São Paulo, Brazil. J Pollin Ecol 6: 95–107.
Acknowledgements
We thank Andrew Matthews, Jaume Pellicer and Ilia Leitch for providing Begonia genome size estimates; Mark Hughes, Héctor Domínguez, Arturo Sanchez, Keith Gardner, Hilda Flores, Miguel Angel Pérez-Farrera for help conducting fieldwork, Fredy Archila for proving DNA samples from Guatemala, Neil Watherston, Fiona Inches and Bruce Robertson for care of the living plants, Alex Clarke, Rhiannon Crichton, Michelle Hollingsworth, Ruth McGregor and Markus Ruhsam for help in the lab, and Peter Hollingsworth and Jon Ågren for comments on an early draft of the manuscript. ADT was supported by a BBSRC PhD scholarship, with additional fieldwork funding from The British Ecological Society, The Genetics Society, The MacIntyre Begonia Trust and The Merlin Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Twyford, A., Kidner, C. & Ennos, R. Genetic differentiation and species cohesion in two widespread Central American Begonia species. Heredity 112, 382–390 (2014). https://doi.org/10.1038/hdy.2013.116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2013.116
Keywords
This article is cited by
-
Past climatic fluctuations are associated with morphological differentiation in the cloud forest endemic tree Ocotea psychotrioides (Lauraceae)
Plant Systematics and Evolution (2018)
-
Three new species of Begonia endemic to the Puerto Princesa Subterranean River National Park, Palawan
Botanical Studies (2015)
-
Strong postzygotic isolation prevents introgression between two hybridizing Neotropical orchids, Epidendrum denticulatum and E. fulgens
Evolutionary Ecology (2015)