Abstract
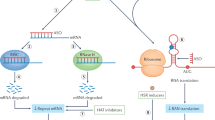
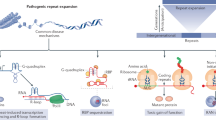
Genetic diseases that are accompanied by central nervous system involvement are often fatal. Among these are the autosomal dominant neurogenetic diseases caused by nucleotide repeat expansion. For example, Huntington's disease (HD) and spinal cerebellar ataxia are caused by expansion of a tract of CAGs encoding glutamine. In HD and the other CAG-repeat expansion diseases, the expansion is in the coding region. Myotonic dystrophy is caused by repeat expansions of CUG or CCTG in noncoding regions, and the mutant RNA is disease causing. Treatments for these disorders are limited to symptomatic intervention. RNA interference (RNAi), which is a method for inhibiting target gene expression, provides a unique tool for therapy by attacking the fundamental problem directly. In this review, we describe briefly several representative disorders and their respective molecular targets, and methods to accomplish therapeutic RNAi. Finally, we summarize studies performed to date.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Mathieu O, Bender J . RNA-directed DNA methylation. J Cell Sci 2004; 117: 4881–4888.
Matzke M, Aufsatz W, Kanno T, Daxinger L, Papp I, Mette MF et al. Genetic analysis of RNA-mediated transcriptional gene silencing. Biochim Biophys Acta 2004; 1677: 129–141.
Morris KV, Chan SW, Jacobsen SE, Looney DJ . Small interfering RNA-induced transcriptional gene silencing in human cells. Science 2004; 305: 1289–1292.
Kawasaki H, Taira K . Induction of DNA methylation and gene silencing by short interfering RNAs in human cells. Nature 2004; 431: 211–217.
Ting AH, Schuebel KE, Herman JG, Baylin SB . Short double-stranded RNA induces transcriptional gene silencing in human cancer cells in the absence of DNA methylation. Nat Genet 2005; 37: 906–910.
Tijsterman M, Plasterk RH . Dicers at RISC; the mechanism of RNAi. Cell 2004; 117: 1–3.
Kim VN . MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 2005; 6: 376–385.
Ma JB, Yuan YR, Meister G, Pei Y, Tuschl T, Patel DJ . Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 2005; 434: 666–670.
Parker JS, Roe SM, Barford D . Structural insights into mRNA recognition from a PIWI domain-siRNA guide complex. Nature 2005; 434: 663–666.
Schwarz DS, Hutvagner G, Du T, Xu Z, Aronin N, Zamore PD . Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003; 115: 199–208.
Khvorova A, Reynolds A, Jayasena SD . Functional siRNAs and miRNAs exhibit strand bias. Cell 2003; 115: 505.
Klockgether T, Evert B . Genes involved in hereditary ataxias. Trends Neurosci 1998; 21: 413–418.
Ranum LP, Day JW . Pathogenic RNA repeats: an expanding role in genetic disease. Trends Genet 2004; 20: 506–512.
Schols L, Bauer P, Schmidt T, Schulte T, Riess O . Autosomal dominant cerebellar ataxias: clinical features, genetics, and pathogenesis. Lancet Neurol 2004; 3: 291–304.
Gusella JF, MacDonald ME . Molecular genetics: unmasking polyglutamine triggers in neurodegenerative disease. Nat Rev Neurosci 2000; 1: 109–115.
Ross CA . When more is less: pathogenesis of glutamine repeat neurodegenerative diseases. Neuron 1995; 15: 493–496.
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 1993; 72: 971–983.
Snell RG, MacMillan JC, Cheadle JP, Fenton I, Lazarou LP, Davies P et al. Relationship between trinucleotide repeat expansion and phenotypic variation in Huntington's disease. Nat Genet 1993; 4: 393–397.
Sharp AH, Loev SJ, Schilling G, Li SH, Li XJ, Bao J et al. Widespread expression of Huntington's disease gene (IT15) protein product. Neuron 1995; 14: 1065–1074.
Landles C, Bates GP . Huntingtin and the molecular pathogenesis of Huntington's disease. EMBO Rep 2004; 5: 958–963.
Ona VO, Li M, Vonsattel JP, Andrews LJ, Khan SQ, Chung WM et al. Inhibition of caspase-1 slows disease progression in a mouse model of Huntington's disease. Nature 1999; 399: 263–267.
Ferrante RJ, Andreassen OA, Jenkins BG, Dedeoglu A, Kuemmerle S, Kubilus JK et al. Neuroprotective effects of creatine in a transgenic mouse model of Huntington's disease. J Neurosci 2000; 20: 4389–4397.
Ferrante RJ, Andreassen OA, Dedeoglu A, Ferrante KL, Jenkins BG, Hersch SM et al. Therapeutic effects of coenzyme Q10 and remacemide in transgenic mouse models of Huntington's disease. J Neurosci 2002; 22: 1592–1599.
Ferrante RJ, Kubilus JK, Lee J, Ryu H, Beesen A, Zucker B et al. Histone deacetylase inhibition by sodium butyrate chemotherapy ameliorates the neurodegenerative phenotype in Huntington's disease mice. J Neurosci 2003; 23: 9418–9427.
Ferrante RJ, Ryu H, Kubilus JK, D’Mello S, Sugars KL, Lee J et al. Chemotherapy for the brain: the antitumor antibiotic mithramycin prolongs survival in a mouse model of Huntington's disease. J Neurosci 2004; 24: 10335–10342.
Dedeoglu A, Kubilus JK, Jeitner TM, Matson SA, Bogdanov M, Kowall NW et al. Therapeutic effects of cystamine in a murine model of Huntington's disease. J Neurosci 2002; 22: 8942–8950.
Sanchez I, Mahlke C, Yuan J . Pivotal role of oligomerization in expanded polyglutamine neurodegenerative disorders. Nature 2003; 421: 373–379.
Chung MY, Ranum LP, Duvick LA, Servadio A, Zoghbi HY, Orr HT . Evidence for a mechanism predisposing to intergenerational CAG repeat instability in spinocerebellar ataxia type I. Nat Genet 1993; 5: 254–258.
Banfi S, Servadio A, Chung MY, Kwiatkowski Jr TJ, McCall AE, Duvick LA et al. Identification and characterization of the gene causing type 1 spinocerebellar ataxia. Nat Genet 1994; 7: 513–520.
Klement IA, Skinner PJ, Kaytor MD, Yi H, Hersch SM, Clark HB et al. Ataxin-1 nuclear localization and aggregation: role in polyglutamine-induced disease in SCA1 transgenic mice. Cell 1998; 95: 41–53.
Cummings CJ, Reinstein E, Sun Y, Antalffy B, Jian Y-H, Ciechanover A et al. Mutation of the E6-AP ubiquitin ligase reduces nuclear inclusion frequency while accelerating polyglutamine-induced pathology in SCA1 mice. Neuron 1999; 24: 879–892.
Arrasate M, Mitra S, Schweitzer ES, Segal MR, Finkbeiner S . Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 2004; 431: 805–810.
Cornett J, Cao F, Wang CE, Ross CA, Bates GP, Li SH et al. Polyglutamine expansion of huntingtin impairs its nuclear export. Nat Genet 2005; 37: 198–204.
Saudou F, Finkbeiner S, Devys D, Greenberg ME . Huntingtin acts in the nucleus to induce apoptosis but death does not correlate with the formation of intranuclear inclusions. Cell 1998; 95: 55–66.
Sugars KL, Rubinsztein DC . Transcriptional abnormalities in Huntington disease. Trends Genet 2003; 19: 233–238.
Mankodi A, Logigian E, Callahan L, McClain C, White R, Henderson D et al. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science 2000; 289: 1769–1773.
Liquori CL, Ricker K, Moseley ML, Jacobsen JF, Kress W, Naylor SL et al. Myotonic dystrophy type 2 caused by a CCTG expansion in intron 1 of ZNF9. Science 2001; 293: 864–867.
Robb GB, Brown KM, Khurana J, Rana TM . Specific and potent RNAi in the nucleus of human cells. Nat Struct Mol Biol 2005; 12: 133–137.
Rodriguez-Lebron E, Paulson HL . Gene Therapy Review. In Press 2005.
Matilla A, Roberson ED, Banfi S, Morales J, Armstrong DL, Burright EN et al. Mice lacking ataxin-1 display learning deficits and decreased hippocampal paired-pulse facilitation. J Neurosci 1998; 18: 5508–5516.
Zeitlin S, Liu JP, Chapman DL, Papaioannou VE, Efstratiadis A . Increased apoptosis and early embryonic lethality in mice nullizygous for the Huntington's disease gene homologue. Nat Genet 1995; 11: 155–163.
Dragatsis I, Levine MS, Zeitlin S . Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat Genet 2000; 26: 300–306.
Harper SQ, Staber PD, He X, Eliason SL, Martins I, Mao Q et al. RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model. Proc Natl Acad Sci USA 2005; 102: 5820–5825.
Wheeler VC, Gutekunst CA, Vrbanac V, Lebel LA, Schilling G, Hersch S et al. Early phenotypes that presage late-onset neurodegenerative disease allow testing of modifiers in Hdh CAG knock-in mice. Hum Mol Genet 2002; 11: 633–640.
Menalled LB, Sison JD, Wu Y, Olivieri M, Li XJ, Li H et al. Early motor dysfunction and striosomal distribution of huntingtin microaggregates in Huntington's disease knock-in mice. J Neurosci 2002; 22: 8266–8276.
Menalled LB, Sison JD, Dragatsis I, Zeitlin S, Chesselet MF . Time course of early motor and neuropathological anomalies in a knock-in mouse model of Huntington's disease with 140 CAG repeats. J Comp Neurol 2003; 465: 11–26.
Hemmings-Mieszczak M, Dorn G, Natt FJ, Hall J, Wishart WL . Independent combinatorial effect of antisense oligonucleotides and RNAi-mediated specific inhibition of the recombinant rat P2X3 receptor. Nucleic Acids Res 2003; 31: 2117–2126.
Thakker DR, Natt F, Husken D, Maier R, Muller M, van der Putten H et al. Neurochemical and behavioral consequences of widespread gene knockdown in the adult mouse brain by using nonviral RNA interference. Proc Natl Acad Sci USA 2004; 101: 17270–17275.
Xia H, Mao Q, Paulson HL, Davidson BL . siRNA-mediated gene silencing in vitro and in vivo. Nat Biotechnol 2002; 20: 1006–1010.
Hommel JD, Sears RM, Georgescu D, Simmons DL, DiLeone RJ . Local gene knockdown in the brain using viral-mediated RNA interference. Nat Med 2003; 9: 1539–1544.
Xia H, Mao Q, Eliason SL, Harper SQ, Martins IH, Orr HT et al. RNAi suppresses polyglutamine-induced neurodegeneration in a mouse model of SCA1. Nat Med 2004; 10: 816–820.
Raoul C, Abbas-Terki T, Bensadoun JC, Guillot S, Haase G, Szulc J et al. Lentiviral-mediated silencing of SOD1 through RNA interference retards disease onset and progression in a mouse model of ALS. Nat Med 2005; 11: 423–428.
Ralph GS, Radcliffe PA, Day DM, Carthy JM, Leroux MA, Lee DC et al. Silencing mutant SOD1 using RNAi protects against neurodegeneration and extends survival in an ALS model. Nat Med 2005; 11: 429–433.
Miller TM, Kaspar BK, Kops GJ, Yamanaka K, Christian LJ, Gage FH et al. Virus-delivered small RNA silencing sustains strength in amyotrophic lateral sclerosis. Ann Neurol 2005; 57: 773–776.
Rodriguez-Lebron E, Denovan-Wright EM, Nash K, Lewin AS, Mandel RJ . Intrastriatal rAAV-mediated delivery of anti-huntingtin shRNAs induces partial reversal of disease progression in R6/1 Huntington's disease transgenic mice. Mol Ther 2005, July E-pub.
Davidson BL, Stein CS, Heth JA, Martins I, Kotin RM, Derksen TA et al. Recombinant adeno-associated type 2, 4 and 5 vectors: transduction of variant cell types and regions in the mammalian CNS. Proc Natl Acad Sci USA 2000; 97: 3428–3432.
Passini MA, Watson DJ, Vite CH, Landsburg DJ, Feigenbaum AL, Wolfe JH . Intraventricular brain injection of adeno-associated virus type 1 (AAV1) in neonatal mice results in complementary patterns of neuronal transduction to AAV2 and total long-term correction of storage lesions in the brains of beta-glucuronidase-deficient mice. J Virol 2003; 77: 7034–7040.
Liu G, Martins IH, Chiorini JA, Davidson BL . Adeno-associated virus type 4 (AAV4) targets ependyma and astrocytes in the subventricular zone and RMS. Gene Therapy 2005; 12 (20): 1503–1508.
Caplen NJ, Taylor JP, Statham VS, Tanaka F, Fire A, Morgan RA . Rescue of polyglutamine-mediated cytotoxicity by double-stranded RNA-mediated RNA interference. Hum Mol Genet 2002; 11 (2): 175–184.
Xia H, Mao Q, Davidson BL . The HIV tat protein transduction domain improves the biodistribution of β-glucuronidase expressed from recombinant viral vectors. Nat Biotechnol 2001; 19: 640–644.
Burright EN, Clark HB, Servadio A, Matilla T, Feddersen RM, Yunis WS et al. SCA1 transgenic mice: a model for neurodegeneration caused by an expanded CAG trinucleotide repeat. Cell 1995; 82: 937–948.
Schilling G, Becher MW, Sharp AH, Jinnah HA, Duan K, Kotzuk JA et al. Intranuclear inclusions and neuritic aggregates in transgenic mice expressing a mutant N-terminal fragment of huntingtin. Hum Mol Genet 1999; 8 (3): 397–407.
Mangiarini L, Sathasivam K, Seller M, Cozens B, Harper A, Hetherington C et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 1996; 87 (3): 493–506.
Zu T, Duvick LA, Kaytor MD, Berlinger M, Zoghbi H, Clark HB et al. Recovery from polyglutamine-induced neurodegeneration in conditional SCA1 transgenic mice. J Neurosci 2004; 24: 8853–8861.
Zuccato C, Ciammola A, Rigamonti D, Leavitt BR, Goffredo D, Conti L et al. Loss of huntingtin-mediated BDNF gene transcription in Huntington's disease. Science 2001; 293: 493–498.
Zuccato C, Tartari M, Crotti A, Goffredo D, Valenza M, Conti L et al. Huntingtin interacts with REST/NRSF to modulate the transcription of NRSE-controlled neuronal genes. Nat Genet 2003; 35: 76–83.
Zuccato C, Liber D, Ramos C, Tarditi A, Rigamonti D, Tartari M et al. Progressive loss of BDNF in a mouse model of Huntington's disease and rescue by BDNF delivery. Pharmacol Res 2005; 52: 133–139.
Yamamoto A, Lucas JJ, Hen R . Reversal of neuropathology and motor dysfunction in a conditional model of Huntington's disease. Cell 2000; 101: 57–66.
Acknowledgements
We thank Christine McLennan for manuscript assistance, and Ines Martins and Song-Lin Ding for help with in vivo studies. The work was supported by NS050210 and HD44093.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Denovan-Wright, E., Davidson, B. RNAi: a potential therapy for the dominantly inherited nucleotide repeat diseases. Gene Ther 13, 525–531 (2006). https://doi.org/10.1038/sj.gt.3302664
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gt.3302664
Keywords
This article is cited by
-
Repeat-associated non-AUG (RAN) translation: insights from pathology
Laboratory Investigation (2019)
-
Experimental Models for Identifying Modifiers of Polyglutamine-Induced Aggregation and Neurodegeneration
Neurotherapeutics (2013)
-
Using non-coding small RNAs to develop therapies for Huntington's disease
Gene Therapy (2011)
-
RNAi: a potential new class of therapeutic for human genetic disease
Human Genetics (2011)
-
Allele-specific silencing of mutant huntingtin and ataxin-3 genes by targeting expanded CAG repeats in mRNAs
Nature Biotechnology (2009)