Abstract
One important piece of information about the human Mendelian disorders is the mode of inheritance. Recent studies of human genetic diseases on a large scale have provided many novel insights into the underlying molecular mechanisms. However, most successful analyses ignored the mode of inheritance of diseases, which severely limits our understanding of human disease mechanisms relating to the mode of inheritance at the large scale. Therefore, we here conducted a systematic large-scale study of the inheritance mode of Mendelian disorders, to bring new insight into human diseases. Our analyses include the comparison between dominant and recessive disease genes on both genomic and proteomic characteristics, Mendelian mutations, protein network properties and disease connections on both the genetic and the population levels. We found that dominant disease genes are more functionally central, topological central and more sensitive to disease outcome. On the basis of these findings, we suggested that dominant diseases should have higher genetic heterogeneity and should have more comprehensive connections with each other compared with recessive diseases, a prediction we confirm by disease network and disease comorbidity.
Similar content being viewed by others
Introduction
Massive efforts and resources have been devoted to mapping human disease loci genetically and later physically over the past decades.1, 2 With the advance of genome-wide association studies (GWAS), detection of molecular interactions and especially the ongoing cancer genome sequencing projects, an impressive list of disorder–gene associations and their mutations has been generated.3, 4 Researchers have begun to explore human diseases on a large scale by genome-wide analysis using complex cellular networks on the basis that disease genes have distinct biochemical characteristics and function by interacting with other genes.1, 5
One important piece of information about human Mendelian diseases is the mode of inheritance, which is usually the first step toward understanding the molecular mechanism of inherited human diseases.6, 7 The mode of inheritance of a disease gene has been shown to have a close relationship with its molecular function.8 However, most large-scale studies of diseases have ignored the mode of inheritance of disease subtypes and their constraints on disease interrelationships, which severely limits our understanding of the relation between human diseases and the mode of inheritance. This study is a large-scale investigation of the genomic, biochemical and functional characteristics, associated mutations and topological properties in a protein network of disease genes with different modes of inheritance. Different analytical approaches yielded consistent results for disease genes with different modes of inheritance, prompting us to examine the interrelationship of dominant versus recessive disease pairs that share genes or show significant comorbidity.
Materials and methods
Inheritance mode and disease–gene association
We retrieved manually curated inheritance and disorder–gene associations from the Online Mendelian Inheritance in Man (OMIM) database.9 In total, we obtained 918 protein-coding genes for which a single mutated allele is associated with autosomal disease (AD genes), and 1065 protein-coding genes of which both mutated alleles are necessary to cause an autosomal disease (AR genes). There are relatively few sex chromosome-linked diseases and associated genes, so these were not included in this study.
Data and calculation of genomic characteristics
The information of gene length, 3′-untranslated region (UTR) and 5′-UTR was retrieved from the Ensembl database using BioMart tool. Human miRNAs were obtained from miRBase database (release 19.0),10 and miRNA targets were retrieved from starBase database, which provides a comprehensive integrated miRNA-target map from CLIP-Seq (HITS-CLIP, PAR-CLIP) and degradome sequencing (Degradome-Seq, PARE) data.11
Data and calculation of biochemical and functional features
Information on genes encoding enzymes was obtained from two sources: 2774 human enzyme genes from KEGG (www.genome.jp/kegg/) and 5326 human genes annotated ‘catalytic activity’ from Gene Ontology (www.geneontology.org/). Also, 1003 transcription factor (TF) genes annotated ‘transcription regulator activity’ and 1731 structural protein-coding genes annotated ‘cytoskeleton’ were obtained from Gene Ontology.
A total of 4357 genes (2064 house-keeping and 2293 tissue-specific) were obtained by a microarray meta-analysis of 1431 samples in 43 normal human tissues from 104 microarray data sets.12 Phenotype data of the mouse orthologs were used to predict the lethality of the corresponding human genes. The information on human–mouse orthologous gene pairs and the phenotype data of mouse knockout genes were retrieved from Mouse Genome Informatics (MGI, http://www.informatics.jax.org/). A gene was defined as essential if its knockout resulted in a lethal phenotype. We identified 2510 human genes with a mouse-lethal ortholog, of which 937 were disease genes according to OMIM.
Phosphorylation sites of genes were extracted from Phospho.ELM. The domain architecture of disease genes was extracted by searching the SMART and Pfam databases of Hidden Markov Models. Hand annotated lists of domains of the SMART database involved in cell signaling domains and nuclear domains (as defined by the SMART database) were used to identify differences between AD and AR genes. Evolutionarily promiscuous domains containing both SMART and Pfam domains were obtained from Butte’s study.13
High quality, comprehensive protein network and protein complex
The following resources were used to compile a high-quality protein network: MINT;14 BioGRID;15 IntAct (version of 1 May 2012);16 DIP;17 BIND Translation;18 HPRD release 9;19 iRefWeb 4.1 to integrate the interactions from innatedb, matrixdb and MIPS MPPI;20 and large-scale Y2H data sets from the literature.4, 21, 22, 23 We manually checked the evidence code of the experiments that were used to detect the interactions in order to filter out low-confidence and non-binary interactions. Any interaction detected by experiments that cannot generate binary interactions was removed. A comparison with an independently collected PPI data set (STRING) confirmed the reliability of our network (Supplementary Figure 1). The comprehensive sources of mammalian protein complexes (Corum)24 and HPRD release 9 were used to compile a comprehensive list of protein complexes. In total, 77 192 binary interactions and 3347 protein complexes were obtained.
Comorbidity disease pairs and the relative risk of comorbidity
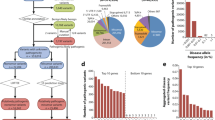
The US Medicare database documenting the diagnoses of 13 039 018 patients from 1990–1993 was used to study the comorbidity related to mode of inheritance (http://barabasilab.neu.edu/projects/hudine/).25, 26 The mapping between ICD-9-CM codes and OMIM disease ID was provided by the Unified Medical Language System (UMLS). Significant disease pairs were selected that met the following criteria: (1) both diseases had >10 hospitalized patients; (2) the randomly expected co-occurrence of the two diseases was ≥1; and (3) the relative risk (RR) value (see below) of the disease pair was significant and >2, which is the benchmark level of all disease pairs. The disease pairs that met criteria (1) and (2) included 2675 disease pairs mapped to AD disease and 340 mapped to AR disease (Figure 5d).
RR was used to quantify the degree that the disease pairs co-occur in patients compared with random expectation. RR is defined as:

where Ii is the incidence of disease i, Cij is the number of patients who were affected simultaneously by diseases i and j, and N=13 039 018. RR>1 means that a disease pair co-occurs more frequently than expected by chance alone.26, 27 The Pearson correlation for binary variables was used as a quantitative measure of comorbidity to check the robustness of the results.
Results and Discussion
Genomic characteristics of AD and AR genes
Genome-wide analysis was used to explore the genomic characteristics of disease genes as a first step toward novel insight into the difference between modes of inheritance. We found AD genes were significantly longer compared with AR genes (86 922 bp versus 68 914 bp; P=5e–04, Wilcoxon rank sum test; Figure 1a). Significant differences were observed for 3′ and 5′-UTRs, whose average lengths in AD genes are 571 bp versus 419 bp (P=1.2e–012, Wilcoxon rank sum test) and 159 bp versus 141 bp (P=3e–013) compared with AR genes (Figure 1a). A growing body of evidence has revealed the important role of 3′ and 5′-UTRs of mRNAs in human diseases.28 Previous studies have also suggested that longer gene structures are most likely associated with gene regulation.29, 30, 31 Hu29 found that miRNA targets have longer 3'-UTR compared with non-miRNA targets. We further analyzed the miRNA regulation for AD and AR genes and found a significantly greater proportion of AD genes regulated at the post-transcriptional level by miRNAs compared with AR genes (29.3 versus 21.2%; P=3.9e–05, Fisher’s exact test). More importantly, AD genes are regulated by, on average, 23 miRNAs, which is more than twice the average 10 miRNA regulators of AR genes (Figure 1b, P=9.7e–014). These results indicate that AD genes are more functionally central than AR genes and suggest AD genes are under stronger selective pressure during evolution compared with AR genes. This is supported by the significantly more conserved coding sequence of AD genes compared with AR genes (Figure 1c, the measurements of evolutionary conservation using phyloP method across 46 vertebrate from UCSC database. P=6.0e–05, Wilcoxon rank sum test). The results of an earlier study of the evolutionary history of dominant and recessive disease genes suggested that dominant disease genes are more conserved than recessive disease genes.32
Comparison of genomic characteristics between AD and AR genes. (a) Comparison of the structure parameters of AD genes with those of AR genes. (b) Comparison of average number of miRNA regulators from the 3′-UTR sequences of AD and AR genes. (c) Evolutionary conservation using phyloP method across 46 vertebrate between AD and AR genes.
Biochemical and functional features of AD and AR genes
We asked whether AD and AR genes display characteristic biochemical and functional features. A preliminary analysis revealed significant differences between the sizes (Figure 2a; 857 versus 702 AA, P=0.003), domain numbers (Figure 2b; 2.6 versus 3.5, P=1.1e–07) and phosphorylation sites (Figure 2c; 2.4 versus 1.0, P=1.3e–08) of the proteins encoded by AD and AR genes. Further analysis of the domain distribution revealed significant differences of domain types between AD and AR genes. First, cell signaling domains were found preferentially in AD genes (Figure 2d; P<0.001, Sign test). For example, 33 signaling domains were found to have a greater distribution frequency in AD genes compared with AR genes. In particular, nine signaling domains (ARM, FH2, GS, G_alpha, PTB, SPRY, S_TK_X, UBQ and ZnF_RBZ) occurred exclusively in AD genes. The same domain distribution difference in AD and AR genes was observed for nuclear domains and promiscuous domains, but not for extracellular or other domains. The domain distribution of disease genes indicates an inherent property of inheritance mode. For instance, the enrichment of nuclear domains in AD genes confirms the particular role of TFs in causing dominant phenotypes, whereas the enrichment of promiscuous domains suggests AD genes are involved more frequently in protein interactions compared with AR genes.13
Comparison of biochemical and functional features between AD and AR genes. (a) Protein sequence lengths of AD genes and AR genes. (b) Number of different domains/protein between AD and AR genes. (c) Mean number of experimentally validated phosphosites from Phospho-ELM database between AD and AR genes. (d) Domain distribution between AD and AR genes. The frequency of a particular domain in AD genes minus its frequency in AR genes is plotted for each of the domains, with a frequency of 1 meaning domains found exclusively in AD genes and –1 meaning domains found exclusively in AR genes. (e) Functional features and essentiality of AD and AR genes.
We investigated biological roles of disease genes associated with different inheritance modes. In accord with earlier work,8 we found AD genes are significantly more likely to encode TFs and less likely to encode enzymes compared with AR genes (Figure 2e). We found also that AD genes are more likely than AR genes to encode structural proteins, consistent with the expectation that the assembly of abnormal proteins into a structural complex disrupts the integrity and function of the complex (Figure 2e). These results fit well with our understanding of how proteins of various functions are associated with phenotypes. Human diseases are generally associated with specific tissue types corresponding to the physiological systems affected,33 therefore, we analyzed 2064 house-keeping genes and 2293 tissue-specific genes obtained from analysis of extensive gene expression data sets,12 and found both AD and AR genes are more likely to be tissue-specific genes (Figure 2e). However, no significant difference of the predisposition of tissue-specific genes between AD and AR genes was found. To assess their relative overall biological importance, we asked whether there was a difference between AD and AR genes in being essential in early development. We considered human orthologs of mouse knockout genes that result in lethality. The classes of embryonic lethality, postnatal lethality, prenatal lethality and perinatal lethality were considered as lethal phenotypes, whereas other phenotypes (including weaning or preweaning lethality) were considered non-lethal. This gave us 2510 human genes having mouse-lethal orthologs, of which 420 were AD genes and 301 were AR genes (Figure 2e; P=10–5, χ2-test). This result further confirms that AD genes are more functionally central compared with AR genes.
Disease mutations of AD and AR genes
To explore AD and AR genes in detail, we analyzed a comprehensive list of Mendelian mutations compiled from both OMIM and the Human Gene Mutation Database.1, 9, 32 In all, 43 625 mutations were mapped into AD and AR genes. We found AD genes typically harbored more disease mutations compared with AR genes, whether in missense mutations, nonsense mutations or indels (Figure 3a; P=0.01, P=4.1e–14 and P=0.02, respectively, two-sample Kolmogorov–Smirnov test). This difference in the number of disease mutations is significant even after controlling for gene length (Supplementary Figure 2, P=0.05, P=2.6e–14 and P=0.02, respectively). We divided the mutations into two categories: 26 980 in-frame mutations, including missense mutations and in-frame indels, and 16 645 out-frame mutations, including nonsense mutations and frameshift indels. It has been suggested that in-frame mutations are likely to give rise to proteins with local defects, whereas out-frame mutations are likely to result in complete loss-of-function of genes.1 Most disease genes harbor both types of mutations; however, we extracted 444 genes showing significantly different frequencies of mutation types. We found that genes harboring primarily in-frame mutations are significantly more likely to be AD genes, whereas genes harboring primarily out-frame mutations are significantly more likely to be AR genes (Figure 3b). This result may suggest that mutations causing local defects in AD genes are more frequently associated with disease outcome than mutations causing local defects in AR genes.
Distribution of Mendelian mutations between AD and AR genes. (a) Distribution of different types of mutations between AD and AR genes. (b) Distribution of AD and AR genes that show significant difference in mutation types, plotted for genes with primarily out-frame disease mutations, with primarily in-frame disease mutations, and with both types of mutations, respectively. (c) The ratio of number of AD genes/number of AR genes is plotted as a function of the number of associated diseases.
Given the functional role of a disease gene from different modes of inheritance might affect its sensitivity to disease outcome, we suspect it could influence the pleiotropy of disease outcome. To test this hypothesis, we analyzed the number of phenotypically different diseases associated with AD and AR genes. AD genes displayed a higher level of pleiotropy compared with AR genes. Among genes associated with six different diseases, the number of AD genes is ca fivefold higher compared with AR genes, whereas among genes associated with at most three different diseases, the number of AD and AR genes is similar (Figure 3c).
Large-scale properties of AD genes and AR genes in the human protein interaction network (PIN)
We undertook a large-scale analysis of a comprehensive and reliable human protein PIN, consisting of literature-curated binary interactions from multiple resources, to provide a system-level understanding of the mechanisms underlying different inheritance modes of human disease. In total, 1667 AD and AR genes were mapped into the PIN, forming a disease gene interaction network with 3094 edges between disease genes (Figure 4a).
AD and AR genes in the human PIN. (a) The procedure to map inheritance information of disease genes into the PIN and the corresponding disease gene interaction network. (b) The closeness centrality of AD and AR genes in PIN, with high closeness values indicating central positions of PIN. (c) Average connectivity in PIN of AD and AR genes. (d) Proportion of AD and AR genes as a member of network module and protein complex. (e) Proportion of AD and AR genes as a member of K-core, plotted for increasing K values. High K value represents densely connected center of network. (f) The innermost core of PIN enriched for AD genes, which is detected using k-core decomposition method by setting k=19.
An apparent characteristic of this disease gene network (GN) is that AD genes tend to occupy central positions and AR genes tend to segregate at the network periphery. We assessed this using a topological measure known as closeness.34 Closeness is the inverse of average lengths through networks, with high closeness reflecting central positions in the network. The closeness values of AD genes in PIN were significantly higher compared with AR genes (Figure 4b; P<10−21, Kolmogorov–Smirnov test). The centrality of AD genes was confirmed by node connectivity. We found the connectivity of AD genes is >2-fold higher compared with AR genes (Figure 4c). Mutations affecting hubs are expected to perturb the network severely, whereas those affecting the peripheral genes have less effect.2 Thus, the topological features of AD and AR genes are consistent with their biological functional importance. Moreover, the level of pleiotropy and the larger number of disease mutations of AD genes might be a consequence of diversiform network perturbations introduced by highly connected genes.
Specific cellular functions are believed to be carried out by modules, usually the aggregation of nodes in a network neighborhood, whose disruption results in a particular phenotype.35, 36, 37 According to the hypothesis of haploinsufficiency, AD disease should be associated more frequently with the disruption of modularity, as the haploinsufficiency of a core component might cause haploinsufficiency of a particular functional module. To address this, we used a probabilistic modeling algorithm to define and to detect network modules.38 This revealed 3123 modular genes, for which the AD genes were significantly more enriched compared with the AR genes (Figure 4d; 41.4% versus 25.6%; P<10−10, Fisher’s exact test). More interestingly, we found a positive correlation between the size of modules and the odds ratio of AD/AR genes, indicating the greater enrichment of AD genes in important functional modules. To confirm our conclusion, we examined a comprehensive list of human protein complexes, a typical representative of modules extracted from widely used databases, which confirmed that AD genes are significantly more enriched in modules compared with AR genes (Figure 4d). We used a core decomposition method to find the centers of network’s intrinsic modules, with higher k values representing more densely connected centers of modules.39 We observed an obvious enrichment of AD genes in the k-cores of PIN, and this trend was more dramatic for large values of k (Figure 4e), confirming AD diseases are associated with the disruption of modules, especially with the disruption of the center of modules. By setting k at 19, we were able to extract the innermost core of human interactome, which consists of 313 genes involved primarily in intracellular signaling cascade and regulation of transcription (Figure 4f). Interestingly, only nine of these genes in the innermost core were AR genes, whereas 56 were associated with AD diseases, including dominant types of deafness, diabetes, Alzheimer’s and Parkinson’s disease.
These results are in accordance with those of an earlier study focused on haploinsufficient (HI) genes in the human genome,40 which found HI genes have a more conserved coding sequence, longer transcripts, longer 3′-UTRs and more interaction partners than haplosufficient (HS) genes. Although the definition of HI and HS genes is very different for AD and AR genes, the findings suggest that one copy loss of functional central genes is more likely to cause human diseases than that of functional peripheral genes. It has been brought to our attention that AD genes resemble Mendelian and complex diseases (MC) genes more than Mendelian but not complex disease (MNC) genes in many properties.41 For example, both MC genes and AD genes are involved in more protein interactions, have greater protein lengths and are more conserved than other genes. Therefore, we investigated the overlap between AD versus AR genes and complex disease genes shown in the genetic association database and found AD genes are significantly over-represented by 1.64-fold compared with AR genes (52% versus 33%; P<0.0001, χ2-test). Finally, we found highly significant differences in genomic, proteomic, disease mutations and network topologies between AD and AR genes are predictive of the inheritance mode of human diseases (Supplementary Table 1 and Supplementary Figure 3), presenting a potential to develop automatic tools for determining the inheritance mode of diseases.
Disease connections of different modes of inheritance
The dramatic difference of functional centrality between AD and AR genes could have immense influence on connections between diseases manifesting different inheritance modes. As an instrument for the large-scale analysis of disease connections, we constructed a disease GN and a disease network (DN) for AD genes/diseases and for AR genes/diseases. First, ignoring the inheritance mode, we grouped disease subtypes into diseases according to their given names. Then, according to the disease–gene associations, we generated two classes of biologically relevant networks as described.2 In the GN class, nodes represent genes and two nodes are connected if they are associated with the same disease. In the DN class, nodes represent diseases and two nodes are connected if they share AD/AR genes. However, unlike the earlier study, this procedure was used for AD and AR genes, respectively, resulting in two networks for each class (Figures 5a and b).
GN, DN and disease comorbidity network. (a) Disease GNs constructed by AD and AR genes, respectively. Nodes represent genes and two nodes are connected if they are associated with the same disease, colored based on the disorder class to which a node belongs. The size of each node is relatively proportional to the number of disorders that the gene is associated. (b) DNs constructed by sharing AD and AR genes respectively. The sizes of nodes are proportional to the number of associated genes, and edge width is proportional to the number of shared genes. (c) Comorbidity networks of ICD-codes pairs mapped to AD diseases and of ICD-codes pairs mapped to AR diseases. Only significant comorbidity links are shown. (d) Average comorbidity value between AD and AR disease pairs, plotted for RR and ϕ-correlation respectively.
The GN class of AD genes contains a giant component connecting most nodes, whereas the GN class of AR genes is segmented into many small clusters (Figure 5a). Genes associated with phenotypically similar diseases tend to form clusters in the GNs. Some AD genes (eg, PTEN and PAX6) are involved in various biological functions and are associated with many phenotypically different diseases, resulting in major hubs in the network and connecting different clusters to the giant component. In contrast, several AR genes (eg, RLBP1 and GDAP1) are associated with multiple diseases that are phenotypically similar, resulting in segmented clusters that represent disease modules of a specific disorder class.
The DN class would be segmented into isolated clusters if the human disorder tended to have distinct and unique genetic origin, else the DN class would be a connected network. In the DN class of sharing AD genes, most disorders form a giant component, suggesting a high level of genetic heterogeneity for most dominant disorders (Figure 5b). Even so, the network is naturally clustered according to the disorder classes, supporting the predisposition of common genetic origins of phenotypically similar disorders. Yet, the dominant subtype of diabetes mellitus, Alzheimer’s disease and deafness do not appear to be located in clusters of the same disorder class, representing an extremely high level of locus heterogeneity and complexity of disease phenotype. In stark contrast, the DN class of sharing AR genes is grouped into many small clusters of a few closely related diseases and contains many single nodes (data not shown), revealing that recessive disorder tend to have a distinct and unique genetic origin. Unlike in the DN of sharing AD genes, diabetes mellitus and deafness are linked to only a few other phenotypes, representing the low level of genetic heterogeneity of their recessive subtypes. Metabolic disorders appear mainly in this network, which is consistent with the tendency for genes encoding enzymes to cause recessive diseases (Figure 2e), and explains the earlier finding that metabolic disorders are under-represented in the giant component of the human diseasome.2
It was shown recently that genetic connections between disorders occur at the population level as well: disease pairs sharing common genetic origins tend to show significant comorbidity.27 We hypothesize that the different disease connections suggested by DN classes of sharing AD and AR genes also have an influence on disease comorbidity. A total of 396 significant comorbidity links were found between AD disorders, connecting in a network covering 46.1% of the mapped AD diseases, where phenotypically similar diseases are adjacent to each other. In contrast, only 65 significant comorbidity links were found between AR disorders, covering only 12.2% of the mapped AR diseases, of which 22 links are provided by metabolic disorders. This result is consistent with the distinction in DN classes, reflecting an association between disease connections at the genetic level and disease comorbidity at the population level. Although the number of significant AR disease comorbidity links is highly under-represented, we find AR disease pairs show significantly stronger comorbidity level than AD disease pairs (Figure 5d). Possible explanations for the low comorbidity level of AD disease pairs are the high level of genetic heterogeneity of AD diseases and the high level of pleiotropy of the shared AD genes, as suggested by this study.
It was found recently that protein–protein interaction can be used to understand disease connections on both genetic and population levels.27, 42 Our study indicates that AD genes are located in the center of the protein network and have a high level of connectivity, resulting in a large number of interactions between AD genes, whereas AR genes are segregated at the periphery of the protein network and have a low level of connectivity, resulting in large network distances between them. Therefore, we believe the disease connections constructed by protein–protein interactions of AD and AR genes should have similar results.
References
Wang X, Wei X, Thijssen B, Das J, Lipkin SM, Yu H : Three-dimensional reconstruction of protein networks provides insight into human genetic disease. Nat Biotechnol 2012; 30: 159–164.
Goh KI, Cusick ME, Valle D, Childs B, Vidal M, Barabasi AL : The human disease network. Proc Natl Acad Sci USA 2007; 104: 8685–8690.
Broeckel U, Schork NJ : Identifying genes and genetic variation underlying human diseases and complex phenotypes via recombination mapping. J Physiol 2004; 554: 40–45.
Yu H, Tardivo L, Tam S et al: Next-generation sequencing to generate interactome datasets. Nat Methods 2011; 8: 478–480.
Dezso Z, Nikolsky Y, Nikolskaya T et al: Identifying disease-specific genes based on their topological significance in protein networks. BMC Syst Biol 2009; 3: 36.
Kennedy MA : Mendelian Genetic Disorders. eLS; e-pub ahead of print 23 September 2005; doi:10.1038/npg.els.0003934.
Guo Y, Wei X, Das J et al: Dissecting disease inheritance modes in a three-dimensional protein network challenges the ‘guilt-by-association’ principle. Am J Hum Genet 2013; 93: 78–89.
Jimenez-Sanchez G, Childs B, Valle D : Human disease genes. Nature 2001; 409: 853–855.
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA : Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 2005; 33: D514–D517.
Kozomara A, Griffiths-Jones S : miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011; 39: D152–D157.
Yang JH, Li JH, Shao P, Zhou H, Chen YQ, Qu LH : starBase: a database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res 2011; 39: D202–D209.
Chang CW, Cheng WC, Chen CR et al: Identification of human housekeeping genes and tissue-selective genes by microarray meta-analysis. PLoS One 2011; 6: e22859.
Butte AJ, Kohane IS : Creation and implications of a phenome-genome network. Nat Biotechnol 2006; 24: 55–62.
Licata L, Briganti L, Peluso D et al: MINT, the molecular interaction database: 2012 update. Nucleic Acids Res 2012; 40: D857–D861.
Chatr-Aryamontri A, Breitkreutz BJ, Heinicke S et al: The BioGRID interaction database: 2013 update. Nucleic Acids Res 2013; 41: D816–D823.
Kerrien S, Aranda B, Breuza L et al: The IntAct molecular interaction database in 2012. Nucleic Acids Res 2012 40: D841–D846.
Salwinski L, Miller CS, Smith AJ, Pettit FK, Bowie JU, Eisenberg D : The database of interacting proteins: 2004 update. Nucleic Acids Res 2004; 32: D449–D451.
Isserlin R, El-Badrawi RA, Bader GD : The biomolecular interaction network database in PSI-MI 2.5. Database (Oxford) 2011; 2011: baq037.
Keshava Prasad TS, Goel R, Kandasamy K et al: Human protein reference database—2009 update. Nucleic Acids Res 2009; 37: D767–D772.
Turner B, Razick S, Turinsky AL et al: iRefWeb: interactive analysis of consolidated protein interaction data and their supporting evidence. Database (Oxford) 2010; 2010: baq023.
Rual JF, Venkatesan K, Hao T et al: Towards a proteome-scale map of the human protein-protein interaction network. Nature 2005; 437: 1173–1178.
Stelzl U, Worm U, Lalowski M et al: A human protein-protein interaction network: a resource for annotating the proteome. Cell 2005; 122: 957–968.
Venkatesan K, Rual JF, Vazquez A et al: An empirical framework for binary interactome mapping. Nat Methods 2009; 6: 83–90.
Ruepp A, Waegele B, Lechner M et al: CORUM: the comprehensive resource of mammalian protein complexes—2009. Nucleic Acids Res 2010; 38: D497–D501.
Lee DS, Park J, Kay KA, Christakis NA, Oltvai ZN, Barabasi AL : The implications of human metabolic network topology for disease comorbidity. Proc Natl Acad Sci USA 2008; 105: 9880–9885.
Hidalgo CA, Blumm N, Barabasi AL, Christakis NA : A dynamic network approach for the study of human phenotypes. PLoS Comput Biol 2009; 5: e1000353.
Park J, Lee DS, Christakis NA, Barabasi AL : The impact of cellular networks on disease comorbidity. Mol Syst Biol 2009; 5: 262.
Chatterjee S, Pal JK : Role of 5'- and 3'-untranslated regions of mRNAs in human diseases. Biol Cell 2009; 101: 251–262.
Hu Z : Insight into microRNA regulation by analyzing the characteristics of their targets in humans. BMC Genomics 2009; 10: 594.
Eisenberg E, Levanon EY : Human housekeeping genes are compact. Trends Genet 2003; 19: 362–365.
Chiaromonte F, Miller W, Bouhassira EE : Gene length and proximity to neighbors affect genome-wide expression levels. Genome Res 2003; 13: 2602–2608.
Furney SJ, Alba MM, Lopez-Bigas N : Differences in the evolutionary history of disease genes affected by dominant or recessive mutations. BMC Genomics 2006; 7: 165.
Jiang X, Liu B, Jiang J et al: Modularity in the genetic disease-phenotype network. FEBS Lett 2008; 582: 2549–2554.
Koschützki D, Schreiber F : Centrality analysis methods for biological networks and their application to gene regulatory networks. Gene Regulation Systems Biol 2008; 2: 193.
Vidal M, Cusick ME, Barabasi A-L : Interactome networks and human disease. Cell 2011; 144: 986–998.
Furlong LI : Human diseases through the lens of network biology. Trends Genet 2013; 29: 150–159.
Barabasi AL, Gulbahce N, Loscalzo J : Network medicine: a network-based approach to human disease. Nat Rev Genet 2011; 12: 56–68.
Finn RD, Marshall M, Bateman A : iPfam: visualization of protein-protein interactions in PDB at domain and amino acid resolutions. Bioinformatics 2005; 21: 410–412.
Wuchty S, Almaas E : Peeling the yeast protein network. Proteomics 2005; 5: 444–449.
Huang N, Lee I, Marcotte EM, Hurles ME : Characterising and predicting haploinsufficiency in the human genome. PLoS Genet 2010; 6: e1001154.
Jin W, Qin P, Lou H, Jin L, Xu S : A systematic characterization of genes underlying both complex and Mendelian diseases. Hum Mol Genet 2012; 21: 1611–1624.
Zhang X, Zhang R, Jiang Y et al: The expanded human disease network combining protein-protein interaction information. Eur J Hum Genet 2011; 19: 783–788.
Acknowledgements
This work was supported by the Foundation of Harbin Medical University (WLD-QN1107).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Hao, D., Wang, G., Yin, Z. et al. Systematic large-scale study of the inheritance mode of Mendelian disorders provides new insight into human diseasome. Eur J Hum Genet 22, 1260–1267 (2014). https://doi.org/10.1038/ejhg.2013.309
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2013.309