Abstract
Sole-source business models for genetic testing can create private databases containing information vital to interpreting the clinical significance of human genetic variations. But incomplete access to those databases threatens to impede the clinical interpretation of genomic medicine. National health systems and insurers, regulators, researchers, providers and patients all have a strong interest in ensuring broad access to information about the clinical significance of variants discovered through genetic testing. They can create incentives for sharing data and interpretive algorithms in several ways, including: promoting voluntary sharing; requiring laboratories to share as a condition of payment for or regulatory approval of laboratory services; establishing – and compelling participation in – resources that capture the information needed to interpret the data independent of company policies; and paying for sharing and interpretation in addition to paying for the test itself. US policies have failed to address the data-sharing issue. The entry of new and established firms into the European genetic testing market presents an opportunity to correct this failure.
Similar content being viewed by others
Background
Interpreting the clinical significance of genomic information depends on broad access to DNA sequence variants and clinical information about those tested. Some proprietary genetic test providers have developed privately controlled databases containing information essential to interpreting the results of their tests. This is exemplified by BRCA1/2 testing by Myriad Genetics (Salt Lake City, UT, USA) in the United States.
As the provider of BRACAnalysis, the sole BRCA1/2 diagnostic test commercially available in the United States and one of the most commercially successful genetic tests worldwide, Myriad Genetics serves as an excellent case study of the importance of collecting clinical data and the implications of keeping those data private. Myriad Genetics has enjoyed market success with BRACAnalysis – Myriad notes that nearly one million patients have had BRCA testing, and it has payment agreements with 2500 insurers or payers.1 Its status as the sole commercial provider of BRCA testing in the United States is a consequence of its exclusive US patent rights. In 1994, scientists, some of whom were affiliated with Myriad, discovered BRCA1, which when mutated results in pronounced predisposition to breast, ovarian and certain other cancers.2, 3, 4 Myriad-associated scientists co-discovered the BRCA2 gene the following year. Myriad patented its discoveries and acquired BRCA patent rights from others. It became the sole commercial testing service for BRCA1/2 in the United States by asserting its patents and clearing the market of US competitors,5 generating over $105 million from its BRACAnalysis test in the second quarter for calendar year 2012.6
Myriad has several competitive advantages based on its long experience in BRCA testing. It runs a highly efficient laboratory, has developed a network of health professionals who use its services, has secured agreements with hundreds of payers, has brand recognition based in part on direct-to-consumer advertising, and has a trained sales force. Although those advantages should abide any change in patent status or data access policies, Myriad’s entry into Europe, projected for later this year,6 presents an opportunity to implement policies on access to BRCA mutation data that can set a salutary precedent not only for BRCA but for genetic testing in general, including whole-genome analysis.7, 8
Interpreting variants of unknown significance
Most patients who get BRCA testing have results that can be interpreted in a relatively straightforward manner – either no variations from ‘wild type’, harmless sequence variations or a clearly deleterious mutation. Such results are valuable to those tested and to their providers, influencing decisions about treatment options, including prophylactic surgery or close monitoring and medical management. Mutations that clearly disrupt protein function (e.g., through a small insertion or deletion that results in a frame shift) are generally obvious upon inspection.
In a significant minority of tests, however, sequence differences from wild type are difficult to interpret. These are ‘variants of unknown significance’ (VUS). Missense mutations that substitute one amino acid for another or changes near intron–exon boundaries can be particularly difficult to interpret. Myriad claims that the fraction of cases resulting in a VUS is 3% in its hands, and 20% for most European BRCA-testing services.1 This discrepancy is at least in part due to Myriad having sole possession of the information needed to interpret VUS results. Myriad has obtained this exclusivity by using its status as the sole BRCA1/2 test provider to develop, at its own cost, an extensive database that relates variants of uncertain significance to phenotype, details their frequency in various populations and includes genetic studies on patient families. Thus, Myriad’s proprietary database that contains information about variants, which is not found in public databases, is probably the major factor in explaining the company’s ability to interpret VUS results more successfully than others.
To its credit and the benefit of patients, Myriad has used its database to reduce the frequency with which it reports a VUS. When Myriad finds a new VUS – or one previously identified but whose clinical significance is not yet understood – it offers free testing to the patient’s family members (something that not all genetic testing laboratories do) in an effort to help determine the variant’s significance. Myriad encourages the person with the VUS to contact others in their family, providing a model letter that patients can send their relatives. Myriad collects data regarding the clinical outcome associated with that VUS, and a VUS may ultimately be reclassified as deleterious or neutral as more is learned; conversely, deleterious or neutral mutations are occasionally reclassified as VUSs.
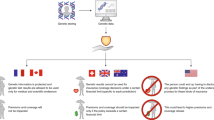
Myriad has access to public databases in interpreting mutations, but outsiders do not have access to Myriad’s database. This asymmetry has clinical impact: a woman might be able to receive BRCA testing from another laboratory in Malawi or Malta, where Myriad’s BRCA patent rights are not in force and testing is perfectly legal, but that laboratory will have no access to Myriad’s data and will thus be unable to interpret many VUS results. Geographic inequities are common in the market for medical products and services. But the fact that the inequity is based on the availability of basic scientific and medical information, rather than of a drug or product, changes the policy context, prompting a debate about keeping clinically relevant data proprietary when that secrecy makes independent verification of its medical significance impossible.
In an environment in which new technologies, including whole-genome and whole-exome sequencing, are already beginning to change clinical practices in genetic testing,9, 10, 11 a proprietary database gives Myriad indefinite exclusivity independent of patent protection. Even if Myriad’s patents are invalidated (for a summary of the ongoing court challenge, see Supplementary materials), or new alternative testing technologies do not infringe them, until the data and interpretive algorithms are re-created in publicly accessible form, competing services will be able to manage VUS results in only two ways: by having samples analyzed at Myriad, where it is interpreted in light of Myriad’s proprietary database, or by rendering inadequate interpretations based upon incomplete public data and algorithms. The former perpetuates Myriad’s exclusivity even after the expiration of its patent rights, while the latter is unacceptable from a clinical perspective. In either case, current practice permits the privatization of valuable clinical data obtained from patients.
Data-sharing practices
Myriad contributed data to public databases until late 2004, but since then its contributions have essentially stopped. Its last major deposit of data to the Breast Cancer Information Core (BIC, the largest database for BRCA mutations, maintained by the National Human Genome Research Institute) was in November 2004. Myriad officials explained to one of us (RC-D) that the decision not to share data was initially because of difficulties in matching data formats, but that after 2005, the company adopted a deliberate policy of retaining data as a trade secret.
Myriad has published some articles on VUS data since November 2004 when its public data-sharing stopped. Investigators with access to the Myriad database through 2006, did the most extensive analysis of VUS, reporting 18 deleterious and 100 neutral variants out of 1433 variants they studied.9 Those 118 sequence variants of known significance are now in the public literature. More than 1200 variants are mentioned in that publication, but the sequences are not listed, and the interpretive algorithms are not specified in detail or deposited where others can use them to interpret the data. Thus, Myriad’s general approach to ‘calling’ VUS results is described, but neither analytic algorithms nor underlying sequence data are available.9, 10, 11, 12, 13 This contrasts with recommendations of the National Academies in two reports that call for depositing data and methods sufficient for replication and interpretation.14, 15
Myriad’s exclusive US testing rights and its pursuit of cases of VUS have enabled it to accumulate data that confer a proprietary advantage in BRCA test interpretation worldwide. Some will surely point to this as a legitimate benefit bestowed by the patent system, part of Myriad’s just reward for innovating. Patents gave Myriad exclusive access to those seeking genetic testing, which enabled Myriad, in turn, to produce a valuable database. Others, however, are likely to consider the withholding of unpatented patient data to hinder rather than ‘to promote the progress of science and useful arts,’ the Constitutional mandate upon which the US patent system is founded. Arguments that focus on rewarding innovation, moreover, must also take into account that much of the work that led to the isolation of BRCA1/2 was done with public or nonprofit funding. The practical effect of retaining such data as a trade secret is to extend Myriad’s testing monopoly beyond the life of the patents on which it was founded.
Whole genome analysis stands poised to have a major impact on medical care if it can be harnessed appropriately. But the biggest challenge to its implementation is properly interpreting the variants found upon analyzing any individual’s genome. As whole-genome and whole-exome sequencing become commonplace, the rate of truly novel mutations will eventually decline. For the foreseeable future, however, each individual whose genome is sequenced will have vast numbers of variants of uncertain clinical significance.
Comprehensive databases such as The Human Gene Mutation Database in Cardiff, MutaDATABASE, the Human Variome Project database, the Leiden Open Variation Database, and other public databases will be essential resources for tracking and interpreting VUS data. Those databases depend, however, on sharing sufficient information to make genotype–phenotype correlations. Myriad, Prevention Genetics and Medical Neurogenetics are the only three laboratories not agreeing to contribute data on human genetic variants to MutaDATABASE, in contrast to over 100 services that have agreed to contribute mutation data (including GeneDx, Quest/Athena, LabCorp and other large commercial testing services).16 The Evidence-based Network for the Interpretation of Mutant Germline Alleles (ENIGMA) was funded as a US National Institutes of Health challenge grant in 2009 to focus on interpreting VUS results from BRCA genes in the public domain.17, 18, 19 It draws from databases and colleagues around the world to apply bioinformatic and laboratory biological methods to improve VUS interpretation. ENIGMA has access to data in Myriad’s database through 2006, but not from the past 5 years.
The objective of these databases and research consortia is to accumulate data and to refine interpretive methods to create a publicly available foundation for improving clinical interpretation of genetic testing. As these public resources accumulate data, the value of proprietary databases will eventually erode, but in the meantime clinicians will be ordering and health plans will be paying for many genetic results that cannot be accurately interpreted based on publicly available information.
Policy options
To deal with this conundrum, one set of policy options involves leveraging the influence of scientific journals and organizations by applying existing disclosure standards. Medical journals and scholars have legitimate claims on data and methods for clinical interpretation of mutations. The 2003 National Academies report on publication of genomic data recommended that ‘authors should include in their publications the data, algorithms or other information that is central or integral to the publication – that is, whatever is necessary to support the major claims of the paper and would enable one skilled in the art to verify or replicate the claims’.15 The Uniform Requirements for Submission of Manuscripts to Medical Journals mandates that authors ‘identify the methods… and procedures in sufficient detail to allow others to reproduce the results’.20 The importance of replicability and objective, independent access to data and algorithms was reiterated in the March 2012 report from the Institute of Medicine (IOM), which recommended that ‘data and metadata used for development of the candidate omics-based test should be made available in an independently managed database’.14 IOM also recommended that computer code and computational methods be fully shared, either through a public database, publication or in the process of regulatory review. These criteria imply a norm of access to data and analytical methods sufficient to make clinical inferences about VUS results.
Leveraging publication standards holds promise but also has limitations. Some journals already require public deposit of data sufficient to independently interpret reported mutations. But as noted above, Myriad’s publications gave the sequences of only 118 of more than 1400 mutations studied, and did not include the interpretive algorithms. Publication guidelines, moreover, apply only when the benefits of publication are sought; they obviously do not apply to unpublished VUS data.
As a related option, databases listing mutations or availability of genetic tests (e.g., the NIH’s nascent Genetic Testing Registry) could mandate test providers share sequence data and interpretive algorithms as a condition of listing their tests. In addition, physicians receiving results, or the organizations that collectively represent them, could also demand access to underlying data and algorithms. For example, standards for reporting such results could be established by the European Society for Human Genetics or international scientific and medical organizations.
Another set of options would rely on the power of payers and regulators. Payers currently reimburse bundled genetic tests and interpretive services. In cases when interpretation cannot be independently verified, payers would be on firm ground to request – or demand – the evidence underlying the clinical determinations.
National health systems, insurers and regulators have several policy tools at their disposal to ensure independent validation of clinical inferences about genomic variants. First, they could ask testing firms to voluntarily adopt policies to share mutation data publicly. Second, payers could refuse payment unless clinically relevant data are shared and subject to independent verification for both accuracy and validity of interpretation. This option further bifurcates into (1) disclosure only to payers or providers, or (2) full public disclosure. That is, if payers mandated data access, disclosure could be limited to regulatory authorities or to those making coverage and payment decisions. Alternatively, payers could require – as a condition of payment – deposit of data and interpretive algorithms into public databases to enable open and independent evaluation, building on the IOM recommendations. Similarly, national authorities that regulate genomic tests could mandate public disclosure as a condition of pre-market approval. Third, national and international institutions could fund research to re-create the data in proprietary databases by ensuring that results of genetic analysis get incorporated into large databases. Such an option, although redundant and thus expensive, might be accomplished through electronic health records that include genomic as well as clinical data, or by building out from consortia such as ENIGMA that have been established for just this purpose – to collect data and develop analytical methods as a public research and clinical resource. Fourth, national health systems could craft payment policies to create incentives for disclosure of data needed to interpret genetic tests – for example, establishing payment codes for public deposit and interpretation of genomic data, in addition to performing the test itself – thus rewarding firms that disclose valuable data.
Conclusion
Current practices of proprietary databases may hinder interpretation of genomic data and impede the advance of personalized medicine. Policies to reward or require data sharing can prevent some foreseeable problems caused by limited access to proprietary data about the clinical significance of genetic variations. Myriad Genetics, for example, has leveraged its BRCA patents to become the dominant BRCA testing service and, in turn, to create a valuable database. Myriad clearly sees its proprietary database as a source of competitive advantage, one that will persist after its underlying patents expire or are invalidated in court. Because of its public profile and explicit, data-based business plan, Myriad’s entry into Europe will force policy choices into stark relief, just as the reduced cost of full-genome analysis brings a worldwide deluge of genomic data. Payers in the United States did not foresee the problems of incomplete access to data, and did not put in place policies to ensure independent verification of clinical predictions. Hundreds of agreements have been signed between genetic testing firms and US payers that have apparently not required disclosure of the underlying data, which is ultimately derived from – and would benefit – patients seeking optimal treatment. Payers and regulators in Europe, South America, Asia and other markets need not be so passive. With the entry of Myriad into Europe in 2012, those making policy decisions about regulation, coverage and reimbursement of genetic tests in Europe can ensure that the data necessary to interpret the clinical significance of genetic variations are made public, where they can be subjected to scientific scrutiny and be available to benefit patients and health professionals around the world.
References
Myriad Genetics I. Written Comments on Genetic Diagnostic Testing Study (the Statement About One Million Tests is on Page 20; the 3 Percent VUS Rate for US Versus 20 Percent for Europe is Attributed to a Market Research Report Cited on Page 23). Salt Lake City, UT: Written statement supplementing oral testimony to the US Patent and Trademark Office pursuant to Section 27 of the America Invents Act of 2011, 2012; 27.
Davies K, White M : Breakthrough: The Race to Find the Breast Cancer Gene. New York: John Wiley, 1996.
Gold ER, Carbone J : Myriad genetics: in the eye of the policy storm. Genet Med 2010; 12 (4 Suppl): S39–S70.
Parthasarathy S : Building Genetic Medicine: Breast Cancer, Technology, and the Comparative Politics of Health Care. Cambridge, MA: MIT Press, 2007.
Cho MK, Illangasekare S, Weaver MA, Leonard DG, Merz JF : Effects of patents and licenses on the provision of clinical genetic testing services. J Mol Diagn 2003; 5: 3–8.
Myriad Genetics I: Form 10-Q: Quarterly Report Pursuant to Section 13 or 15(d) of the Securities Exchange Act of 1934 for the Quarterly Period Ended Septembera 30, 2011. Salt Lake City, Utah: Myriad Genetics, Inc., 2011.
Hopkins MM, Nightingale P : Strategic risk management using complementary assets: organizational capabilities and the commercialization of human genetic testing in the UK. Res Policy 2006; 35: 355–374.
UK Human Genetic Commission: Intellectual Property and DNA Diagnostics. London: Human Genetics Commission, 2010.
Easton DF, Deffenbaugh AM, Pruss D et al: A systematic genetic assessment of 1,433 sequence variants of unknown clinical significance in the BRCA1 and BRCA2 breast cancer-predisposition genes. Am J Hum Genet 2007; 81: 873–883.
Eisenbraun A, Wenstrup R, Hellerstedt B et al: Hereditary breast and ovarian cancer testing: integration and outcomes within community oncology practices. Commun Oncol 2010; 7: 75–81.
Hall MJ, Reid JE, Burbidge LA et al: BRCA1 and BRCA2 mutations in women of different ethnicities undergoing testing for hereditary breast-ovarian cancer. Cancer 2009; 115: 2222–2233.
Saam JBL, Bowles K, Roa B et al: Decline in rate of BRCA1/2 variants of uncertain significance: 2002-2008. 27th Annual Education Conference of the National Society of Genetic Counselors. Los Angeles, CA, 2008.
Wu K, Hinson SR, Ohashi A et al: Functional evaluation and cancer risk assessment of BRCA2 unclassified variants. Cancer Res 2005; 65: 417–426.
Micheel CM, Nass SJ, Omenn GS eds: Evolution of Translational Omics: Lessons Learned and the Path Forward. National Academies Press, 2012.
National Research Council: Sharing Publication-Related Data and Materials: Responsibilities of Authorship in the Life Sciences. Washington, DC: National Academies Press, 2003.
MutaDATABASE. ‘Participating Laboratories’ and ‘Non-Participating Laboratories’ lists at www.mutadatabase.org and personal communication, Patrick Willems, 18 April 2012; in Willems P ed. MutaDATABASE website Belgium, 2012.
Lindor NM, Guidugli L, Wang X et al: A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat 2012; 33: 8–21.
Radice P, De Summa S, Caleca L, Tommasi S : Unclassified variants in BRCA genes: guidelines for interpretation. Ann Oncol 2011; 22 (Suppl 1): i18–i23.
Tommasi S, De Summa S, Pilato B, Paradiso A : BRCA unclassified variants: how can they be classified? Curr Women Health Rev 2012; 8: 30–37.
International Committee of Medical Journal Editors: Uniform Requirements for Manuscripts Submitted to Biomedical Journals: Writing and Editing for Biomedical Publication 2010.
Acknowledgements
This work was supported in part by NIH grant P50 HG003391, the Ewing Marion Kauffman Foundation and Fondation Brocher. This manuscript benefited from the insightful comments of Richard Epstein, Michael Hopkins, Patrick Willems and anonymous reviewers.
Author contributions: All four authors were directly involved in all phases of writing and revising this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Copyright
This manuscript is based in part on US federally funded research, and thus subject to the PubMed Central deposition requirements. The authors will submit the authors’ version of the manuscript and pay for the ‘open publication’ option.
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Cook-Deegan, R., Conley, J., Evans, J. et al. The next controversy in genetic testing: clinical data as trade secrets?. Eur J Hum Genet 21, 585–588 (2013). https://doi.org/10.1038/ejhg.2012.217
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2012.217
This article is cited by
-
Patients v. Myriad or the GDPR Access Right v. the EU Database Right
European Journal of Human Genetics (2019)
-
Privacy in the age of medical big data
Nature Medicine (2019)
-
The challenges of the expanded availability of genomic information: an agenda-setting paper
Journal of Community Genetics (2018)
-
Outcomes of retesting BRCA negative patients using multigene panels
Familial Cancer (2017)
-
The changing life science patent landscape
Nature Biotechnology (2016)