Abstract
Loss-of-function mutations in the cystatin B (CSTB), a cysteine protease inhibitor, gene underlie progressive myoclonus epilepsy of Unverricht–Lundborg type (EPM1), characterized by myoclonic and tonic–clonic seizures, ataxia and a progressive course. A minisatellite repeat expansion in the promoter region of the CSTB gene is the most common mutation in EPM1 patients and leads to reduced mRNA levels. Seven other mutations altering the structure of CSTB, or predicting altered splicing, have been described. Using a novel monoclonal CSTB antibody and organelle-specific markers in human primary myoblasts, we show here that endogenous CSTB localizes not only to the nucleus and cytoplasm but also associates with lysosomes. Upon differentiation to myotubes, CSTB becomes excluded from the nucleus and lysosomes, suggesting that the subcellular distribution of CSTB is dependent on the differentiation status of the cell. Four patient mutations altering the CSTB polypeptide were transiently expressed in BHK-21 cells. The p.Lys73fsX2-truncated mutant protein shows diffuse cytoplasmic and nuclear distribution, whereas p.Arg68X is rapidly degraded. Two missense mutations, the previously described p.Gly4Arg affecting the highly conserved glycine, critical for cathepsin binding, and a novel mutation, p.Gln71Pro, fail to associate with lysosomes. These data imply an important lysosome-associated physiological function for CSTB and suggest that loss of this association contributes to the molecular pathogenesis of EPM1.
Similar content being viewed by others
Introduction
Progressive myoclonus epilepsy of Unverricht–Lundborg type (EPM1; OMIM 254800) is an autosomal recessive disorder characterized by the age of onset at 6–15 years, stimulus-sensitive myoclonus, tonic–clonic epileptic seizures and a progressive course.1,2 Mutations in the ubiquitously expressed gene encoding cystatin B (CSTB), a cysteine protease inhibitor, underlie EPM1.3 The majority of EPM1 patients are homozygous for a minisatellite repeat expansion mutation in the promoter region of the CSTB gene.4,5,6 This expanded dodecamer repeat downregulates CSTB gene expression in vitro and in vivo.3,5,6,7,8 Seven additional EPM1-associated mutations have been described.3,9,10,11 Five of them, occurring in compound heterozygous form with the minisatellite expansion, predict protein truncation either through nonsense (g.2388C>T, p.Arg68X), frameshift (g.2400_2402delTC, p.Lys73fsX2) or splicing alterations (g.1925G>C, g.2027G>A, g.2353A>G). The sixth mutation, g.426G>C, identified in homozygous form,9 results in the substitution of a conserved glycine at position four, critical for binding the proteases,12 to an arginine (p.Gly4Arg). A novel missense mutation, g.2398A>C, found in compound heterozygous form with the minisatellite expansion, results in the substitution of a glutamine at position 71 by a proline (p.Gln71Pro).11
Although CSTB has been shown to function in vitro as a reversible inhibitor of cathepsins B, H, L and S,13,14 its physiological function remains unknown. In EPM1 patient lymphoblastoid cells decreased CSTB inhibitory activity correlates with enhanced activity of cathepsins B, L and S, demonstrating that cathepsin activity is regulated by CSTB also in vivo.15 CSTB-deficient mice develop myoclonic seizures and ataxia, similar to human patients.16 The mice also show loss of cerebellar granule cells due to apoptotic cell death,16 neuronal atrophy and apoptosis outside the cerebellum and gliosis.17 In mice double-deficient for CSTB and cathepsin B, the amount of cerebellar apoptosis is significantly decreased, suggesting that cathepsin B contributes to the apoptotic phenotype in EPM1.18 Moreover, seizures induce upregulation of CSTB mRNA and protein in rats.19 Together, these findings strongly implicate a role for CSTB in neuroprotection.
Apart from the reduced CSTB mRNA expression due to the minisatellite expansion, consequences of the other EPM1 patient mutations at cellular level have not been investigated. We here first analysed the subcellular localization of CSTB in COS-1 cells, myoblasts and differentiated myotubes by using a novel, highly specific monoclonal CSTB antibody. We further focused to study four known mutations predicted to alter the CSTB polypeptide and demonstrate loss of lysosomal association of these EPM1 mutant CSTB proteins.
Materials and methods
Production of mouse monoclonal CSTB antibodies
CSTB was purified from human muscle and spleen as described previously,20 and antibodies were raised according to standard methods.21,22 The antigenic epitope was mapped with overlapping 13-mer peptides using the SPOT technique.23
Expression constructs
Mutations were introduced to the human CSTB cDNA in pcDNA3.1(+) vector (Invitrogen) using the QuikChange™ Site-Directed Mutagenesis Kit (Stratagene).
Cell culture and transfection
COS-1 and BHK-21 cells were cultured according to American Type Culture Collection's instructions. Epstein–Barr virus-transformed lymphoblastoid cells were cultured in RPMI 1640 with 10% FCS. Human pectoral myoblasts (kindly provided by Olli Carpén, University of Helsinki, Finland) were isolated as described before,24 cultured in DMEM with 15% FCS and 2% Ultroser G (Life Technologies) and differentiated into myotubes for 4 days in DMEM with 0.4% Ultroser G.
Cells were transfected using FuGENE 6 Transfection Reagent (Roche), incubated with 25 μ M lactacystin (Sigma) for 3 h and with 5 μ M LysoTracker® Green DND-26 (Molecular Probes) for 1 h.
In vitro translation, immunoprecipitation and Western blot
The cDNAs were translated using the TNT® Quick Coupled Transcription/Translation System and canine pancreatic microsomal membranes (Promega) and immunoprecipitated. For secretion analysis, CSTB was immunoprecipitated both from the total culture medium and cells lysed from a 25 cm2 flask. Proteins were separated on 14% SDS–PAGE gels under reducing conditions. Primary antibodies were detected using HRP-conjugated secondary antibodies (Dako) and enhanced chemiluminescence (Amersham Biosciences).
Immunofluorescence microscopy
The cells were fixed with 4% paraformaldehyde in PBS and permeabilized with either 0.1% saponin or 0.1% TX-100. Polyclonal antibodies against protein disulphide isomerase (PDI) were from StressGen Biotechnologies, TGN46 from Serotec and CSTB from Biogenesis; mSec13, mannose-6-phosphate receptor (MPR) and aspartylglucosaminidase (AGA) were kindly provided by Wanjin Hong (The National University of Singapore, Republic of Singapore), Kurt von Figura (University of Göttingen, Germany) and Anu Jalanko (National Public Health Institute, Finland), respectively. FITC-labelled Lentil lectin was from Sigma and Cy2- or Cy3-conjucated secondary antibodies were from Jackson Immunochemicals. The cells were viewed using Axioplan 2 microscope and photographed with AxioVision 3.1 (Zeiss).
Results
Production and specificity of mouse monoclonal CSTB antibodies
We raised monoclonal antibodies against CSTB purified from human muscle and spleen. In Western blots of COS-1 cell lysates and control lymphoblastoid cells, the CSTB antibody from hybridoma clone RJMW 2E7 (2E7) recognized a protein of approximately 12 kDa, the calculated molecular weight of CSTB, whereas in patient lymphoblastoid cells, no CSTB protein was detected (Figure 1). The 2E7 antibody did not crossreact with the homologous cystatin A (CSTA) in Western analyses (Figure 1) or in ELISA (data not shown). Epitope mapping (data not shown) revealed amino acids 31–39 (ENKKFPVFK) of the CSTB sequence as the epitope for the 2E7 antibody. With the exception of one amino acid, the sequence (TNETYGKLE) in the corresponding region of CSTA is different adding further proof to the specificity of 2E7.
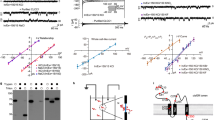
Specificity of the monoclonal CSTB antibody 2E7. 2E7 detected CSTB in COS-1 and control lymphoblastoid cells (lanes 1 and 2, respectively) and recognized purified CSTB (lane 5). There was no immunoreaction in lymphoblastoid cells of an EPM1 patient homozygous for the minisatellite repeat expansion (lane 3), of a patient compound heterozygous for the expansion and the g.2388C>T mutation predicting truncated p.Arg68X protein (lane 4) or against purified CSTA protein (lane 6). 15 μg (lanes 1–4) or 50 ng (lanes 5 and 6) of proteins were loaded per lane.
Subcellular localization of endogenous CSTB
Immunofluorescence analyses of endogenous CSTB in COS-1 cells and human primary myoblasts using the monoclonal 2E7 antibody showed that CSTB exists in several intracellular compartments (Figure 2). Detergents with different permeabilization properties were used to optimally visualize cellular pools of CSTB. We first used Triton X-100 to visualize both nuclear and cytoplasmic CSTB. Nuclear staining of CSTB was prominent in COS-1 cells (Figure 2a) as well as in proliferating myoblasts (Figure 2b). In addition, CSTB was localized diffusely and in punctate structures in the cytoplasm. In differentiated myotubes, CSTB was mostly excluded from the nucleus and mainly localized in the cytoplasm (Figure 2c). Numerous punctate cytoplasmic structures were observed in cells permeabilized with saponin that was used to optimally visualize the cytoplasmic pool (Figure 2d and e). The number of these structures was decreased upon differentiation to myotubes (Figure 2f).
Subcellular localization of endogenous CSTB protein. Fixed cells were permeabilized with either TX-100 (a–c) or saponin (does not permeabilize the nuclear envelope) (d–f) and stained with the 2E7 antibody. CSTB in COS-1 cells (a and d) and myoblasts (b and e) show a nuclear, diffuse cytoplasmic and punctate cytoplasmic staining pattern. In differentiated myotubes (shown by an arrow, c and f), CSTB is localized diffusely in the cytoplasm and excluded from the nucleus and punctate cytoplasmic structures.
A variety of organelle markers was used to identify the CSTB-positive punctate cytoplasmic structures in myoblasts. No overlap was observed with CSTB and a marker for the endoplasmic reticulum (ER), PDI (Figure 3a). CSTB was also excluded from the early secretory pathway as judged by complete separation of CSTB and an ER exit site marker mSec13 (Figure 3b) or a Golgi-marker Lentil lectin (Figure 3c). CSTB colocalized neither with a trans-Golgi network marker, TGN46 (Figure 3d), nor a late endosomal marker MPR (Figure 3e). However, CSTB displayed significant overlap (Figure 3f) with an antibody against a lysosomal enzyme AGA,25,26 as indicated by the yellow colour in merged images. Many of the punctate cytoplasmic structures positive for CSTB contained AGA, whereas not all AGA-positive structures stained with the CSTB antibody. This indicates that CSTB associates only with a subset of lysosomes. Colocalization of endogenous CSTB with AGA and lack of colocalization with other cellular markers was also observed in COS-1 cells (data not shown).
CSTB associates with lysosomes. Immunofluorescence analyses of CSTB (red) and organelle markers (green) in saponin-permeabilized myoblasts. CSTB does not colocalize with markers for ER (PDI) (a), ER exit site (mSec13) (b), Golgi (Lentil lectin) (c), trans-Golgi network (TGN46) (d) or late endosomes (MPR) (e), whereas an overlap with the lysosomal marker AGA (f) is observed.
CSTB is neither inserted into microsomal membranes in vitro nor secreted in cell cultures
In vitro translated CSTB was not enriched in microsomes as a result of cotranslational or post-translational addition of the membranes, but remained largely in the supernatants (Figure 4a), showing that CSTB is unable to translocate into microsomal membranes. On the contrary, a soluble lysosomal enzyme (AGA) and an integral membrane protein (p58/ERGIC-53) were translocated through or integrated in microsomal membranes, respectively (data not shown). In Western blot analyses of COS-1 cells and culture medium, CSTB was detected in the cell lysate but could not be immunoprecipitated from the medium (Figure 4b), indicating that endogenous CSTB is not secreted in significant amounts in these cells.
CSTB is neither inserted into membranes nor secreted. (a) CSTB was in vitro translated without microsomal membranes and immunoprecipitated with the 2E7 antibody (lane 1). When microsomes are added either cotranslationally (lanes 2 and 3) or post-translationally (lanes 4 and 5), CSTB enriches in the supernatant fractions (lanes 2 and 4), and is virtually absent from the microsomal pellets (lanes 3 and 5). The reason for CSTB protein migrating as doublets in SDS-PAGE is currently unknown. (b) In Western blot analyses of immunoprecipitated endogenous CSTB from COS-1 cells and culture medium, the CSTB protein is detected in the cell lysate but not in the culture medium, indicating that it is not secreted significantly in these cells.
Expression and intracellular targeting of mutant proteins representing EPM1 patient mutations
The subcellular localization of four CSTB polypeptides representing EPM1 mutations was investigated using transient expression in BHK cells: g.426G>C/p.Gly4Arg; g.2398A>C/p.Gln71Pro; g.2388C>T/p.Arg68X; and g.2400_2402delTC/p.Lys73fsX2. BHK cells were chosen for transfection experiments since no significant crossreaction with endogenous hamster protein was observed with the CSTB antibodies used. Since polyclonal antibodies recognizing a lysosomal protein in hamster cells were not available, we chose to visualize the lysosomes with LysoTracker Green. The overexpressed wild-type CSTB displayed a distribution similar to that of the endogenous protein, as it was detected in the nucleus, cytoplasm and lysosomes (Figure 5a). Both the p.Gly4Arg (Figure 5b) and p.Gln71Pro (Figure 5c) missense mutants were diffusely localized in the cytoplasm and nucleus, but no punctate structures overlapping with LysoTracker were observed (Figure 5b and c). The truncated mutant, p.Lys73fsX2 (Figure 5d), was seen diffusely distributed to the cytoplasm and nucleus with no colocalization with LysoTracker. The p.Arg68X mutant protein (Figure 5e) showed diffuse cellular staining but was detected only in the presence with a proteasomal inhibitor, lactacystin, indicating rapid degradation of the newly synthesized protein. In conclusion, the wild-type CSTB is localized in the cytoplasm, nucleus and lysosomes, while all mutant proteins studied failed to associate with lysosomes, but were distributed diffusely in the cytoplasm and nucleus.
Transient expression of CSTB polypeptides representing EPM1-associated mutations in BHK cells. (a) Transiently expressed wild-type CSTB (red) displays a distribution similar to that of the endogenous protein including localization to the nucleus, cytoplasm and lysosomes (visualized with LysoTracker Green). The p.Gly4Arg (b), p.Gln71Pro (c) and p.Lys73fsX2 (d) mutant proteins are detected in the nucleus and cytoplasm, but show no overlap with the LysoTracker and thus no association with lysosomes. The p.Arg68X (e) mutant protein is detected only in the presence of lactacystin, indicating rapid degradation by proteasomes. Immunofluorescence stainings were carried out with the monoclonal (2E7) (a–c) or polyclonal CSTB antibody (d and e). Even though the epitope recognized by 2E7 was preserved in both truncated mutant proteins, the antibody failed to detect them and a polyclonal CSTB antibody was used instead.
Discussion
CSTB, a soluble cytoplasmic cathepsin inhibitor, has been considered to counteract inappropriate proteolysis due to cathepsins that leak out of the lysosomes. Recent evidence shows that CSTB is also found in the nucleus and in cytoplasmic granular structures, representing either the Golgi complex or lysosomes, in a variety of cell lines.19,27,28,29 We show here, in line with previous studies, that CSTB resides in the nucleus and cytoplasm and associates with lysosomes in human primary myoblasts. In vitro translation experiments showed that CSTB was unable to translocate into microsomal membranes. In addition, we could not immunoprecipitate CSTB from cell culture medium or detect it in the cellular compartments of the early secretory pathway even when overexpressed. Although these results, together with the fact that CSTB lacks an ER signal sequence, suggest that CSTB is attached to the outer side of the lysosomal membrane, this would need confirmation by, for example, immunoelectronmicroscopy.
In differentiated myotubes, CSTB is excluded from the nucleus and lysosomes and localizes diffusely in the cytoplasm. This is in agreement with previous findings in which CSTB was found to reside in the nucleus in proliferating neuronal cells but distributed to the cytoplasm in differentiated human neuroblastoma and rat primary cerebellar granule cells.28 Since findings of the distribution of CSTB are contradictory in other neuronal cell types,29 further systematic evaluation is necessary to determine the role of CSTB during cell differentiation.
We investigated the consequences of EPM1 mutations that lead to altered CSTB protein sequence on intracellular targeting of CSTB. The prematurely truncated protein, p.Lys73fsX29 did not associate with lysosomes, but showed diffuse cytoplasmic and nuclear distribution. The p.Arg68X3 polypeptide was rapidly degraded and only detected in the presence of a proteasomal inhibitor. This finding is in line with the lack of CSTB immunoreactivity in patient cells harboring this mutation (Figure 1). Missense mutations p.Gly4Arg9 and p.Gln71Pro11 are predicted to affect functionally important amino acids. The evolutionary conserved glycine residue at position four has been shown to be important for interaction with the target proteases.12,30,31,32 The glutamine at position 71 does not interact directly with target proteases, but is located proximal to the second hairpin loop, which also contributes to protease binding.12,33 Mutation of glutamine 71 to a proline predicts shortening of the fourth beta strand by two residues, which would most likely result in changes in the second binding loop and altered binding affinities for cathepsins. Interestingly, both missense mutations also abolish concentration of CSTB at the lysosomal membrane, implying an important physiological function for this association. However, the possibility that the loss of lysosomal association is a consequence of altered cathepsin inhibitory function of CSTB remains speculative.
The lysosomal association of CSTB is compatible with the current view of CSTB having a protective role as a constitutive cytoplasmic inhibitor for cysteine proteases released from lysosomes. This view is strongly supported by the decreased cerebellar granule cell apoptosis detected in mice double-deficient for CSTB and cathepsin B.18 Yet, the severity of the ataxia and seizure phenotypes were not reduced,18 suggesting that CSTB might have other functions than cathepsin inhibition. The localization of CSTB both in the cytoplasm and nucleus allows one to speculate a role for CSTB in signalling events involved in apoptosis with downstream effects of transcriptional regulation. Data emerging from yeast two-hybrid studies (Di Giaimo et al34 and our unpublished data) suggests that CSTB may also be part of a complex molecular network within the cell and challenges further studies on CSTB function in normal physiological as well as in pathological conditions both in non-neuronal and neuronal cells.
References
Koskiniemi M, Donner M, Majuri H, Haltia M, Norio R : Progressive myoclonus epilepsy. A clinical and histopathological study. Acta Neurol Scand 1974; 50: 307–332.
Norio R, Koskiniemi M : Progressive myoclonus epilepsy: genetic and nosological aspects with special reference to 107 Finnish patients. Clin Genet 1979; 15: 382–398.
Pennacchio LA, Lehesjoki A-E, Stone NE et al: Mutations in the gene encoding cystatin B in progressive myoclonus epilepsy (EPM1). Science 1996; 271: 1731–1734.
Lafreniere RG, Rochefort DL, Chretien N et al: Unstable insertion in the 5′ flanking region of the cystatin B gene is the most common mutation in progressive myoclonus epilepsy type 1, EPM1. Nat Genet 1997; 15: 298–302.
Lalioti MD, Scott HS, Buresi C et al: Dodecamer repeat expansion in cystatin B gene in progressive myoclonus epilepsy. Nature 1997; 386: 847–851.
Virtaneva K, D'Amato E, Miao J et al: Unstable minisatellite expansion causing recessively inherited myoclonus epilepsy, EPM1. Nat Genet 1997; 15: 393–396.
Lalioti MD, Scott HS, Antonarakis SE : Altered spacing of promoter elements due to the dodecamer repeat expansion contributes to reduced expression of the cystatin B gene in EPM1. Hum Mol Genet 1999; 81: 791–798.
Alakurtti K, Virtaneva K, Joensuu T, Palvimo JJ, Lehesjoki A-E : Characterization of the cystatin B gene promoter harboring the dodecamer repeat expanded in progressive myoclonus epilepsy, EPM1. Gene 2000; 242: 65–73.
Lalioti MD, Mirotsou M, Buresi C et al: Identification of mutations in cystatin B, the gene responsible for the Unverricht–Lundborg type of progressive myoclonus epilepsy (EPM1). Am J Hum Genet 1997; 60: 342–351.
Kagitani-Shimono K, Imai K, Okamoto N, Ono NJ, Okada S : Unverricht–Lundborg disease with cystatin B gene abnormalities. Pediatr Neurol 2002; 26: 55–60.
de Haan GJ, Halley DJJ, Doelman CJ et al: Unverricht–Lundborg disease: Underdiagnosed in the Netherlands. Epilepsia 2004; 45: 1061–1063.
Stubbs MT, Laber B, Bode W et al: The refined 2.4 Å X-ray crystal structure of recombinant human stefin B in complex with the cystein protease papain: a novel type of proteinase inhibitor interaction. EMBO J 1990; 9: 1939–1947.
Green GD, Kembhavi AA, Davies ME, Barrett AJ : Cystatin-like cysteine proteinase inhibitors from human liver. Biochem J 1984; 218: 939–946.
Brömme D, Rinne R, Kirschke H : Tight-binding inhibition of cathepsin S by cystatins. Biomed Biochim Acta 1991; 50: 631–635.
Rinne R, Saukko P, Järvinen M, Lehesjoki A-E : Reduced cystatin B activity correlates with enhanced cathepsin activity in progressive myoclonus epilepsy. Ann Med 2002; 34: 380–385.
Pennacchio LA, Bouley DM, Higgins KM, Scott MP, Noebels JL, Myers RM : Progressive ataxia, myoclonic epilepsy and cerebellar apoptosis in cystatin B-deficient mice. Nat Genet 1998; 20: 251–258.
Shannon P, Pennacchio LA, Houseweart MK, Minassian BA, Myers RM : Neuropathological changes in a mouse model of progressive myoclonus epilepsy, cystatin B deficiency and Unverricht–Lundborg disease. J Neuropathol Exp Neurol 2002; 61: 1085–1091.
Houseweart MK, Pennacchio LA, Vilaythong A, Peters C, Noebels JL, Myers RM : Cathepsin B but not cathepsins L or S contributes to the pathogenesis of Unverricht–Lundborg progressive myoclonus epilepsy (EPM1). J Neurobiol 2003; 56: 315–327.
D'Amato E, Kokaia Z, Nanobashvili A et al: Seizures induce widespread upregulation of cystatin B, the gene mutated in progressive myoclonus epilepsy, in rat forebrain neurons. Eur J Neurosci 2000; 12: 1687–1695.
Järvinen M, Rinne A : Human spleen cysteine protease inhibitor. Purification, fractionation into isoelectric variants and some properties of the variants. Biochim Biophys Acta 1982; 708: 210–217.
Köhler G, Milstein C : Continuous culture of fused cells secreting antibodies of predefined specificity. Nature 1975; 56: 495–497.
Weber E, Bahn H, Günther D : Monoclonal antibodies against cathepsin L and procathepsin L of different species. Hybridoma 1997; 16: 159–166.
Frank R, Overwin H : Spot synthesis. Epitope analysis with arrays of synthetic peptides prepared on cellulose membranes. Methods Mol Biol 1996; 66: 149–169.
Yasin R, van Beers G, Nurse KCE, Al-Ani S, Landon DN, Thompson EJ : A quantitative technique for growing human adult skeletal muscle in culture starting from mononucleated cells. J Neurol Sci 1977; 32: 347–360.
Enomaa N, Heiskanen T, Halila R et al: Human aspartylglucosaminidase. A biochemical and immunocytochemical characterization of the enzyme in normal and aspartylglucosaminuria fibroblasts. Biochem J 1992; 286: 613–618.
Enomaa NE, Lukinmaa PL, Ikonen EM et al: Expression of aspartylglucosaminidase in human tissues from normal individuals and aspartylglucosaminuria patients. J Histochem Cytochem 1993; 41: 981–989.
Calkins CC, Sameni M, Koblinski J, Sloane BF, Moin K : Differential localization of cysteine protease inhibitors and a target cysteine protease, cathepsin B, by immuno-confocal microscopy. J Histochem Cytochem 1998; 46: 745–751.
Riccio M, Di Giaimo R, Pianetti S, Palmieri PP, Melli M, Santi S : Nuclear localization of cystatin B, the cathepsin inhibitor implicated in myoclonus epilepsy (EPM1). Exp Cell Res 2001; 262: 84–94.
Brännvall K, Hjelm H, Korhonen L, Lahtinen U, Lehesjoki A-E, Lindholm D : Cystatin-B is expressed by neural stem cells and by differentiated neurons and astrocytes. Biochem Biophys Res Commun 2003; 308: 369–374.
Estrada S, Nycander M, Hill NJ, Craven CJ, Waltho JP, Björk I : The role of Gly-4 of human cystatin A (stefin A) in the binding of target proteinases. Characterization by kinetic and equilibrium methods of the interactions of cystatin A Gly-4 mutants with papain, cathepsin B, and cathepsin L. Biochemistry 1998; 37: 7551–7560.
Pol E, Björk I : Role of the single cysteine residue, Cys 3, of human and bovine cystatin B (stefin B) in the inhibition of cysteine proteases. Protein Sci 2001; 10: 1729–1738.
Pol E, Björk I : Contributions of individual residues in the N-terminal region of cystatin B (stefin B) to inhibition of cysteine proteases. Biochim Biophys Acta 2003; 1645: 105–112.
Pol E, Björk I : Importance of the second binding loop and the C-terminal end of cystatin B (stefin B) for inhibition of cysteine proteinases. Biochemistry 1999; 38: 10519–10526.
Di Giaimo R, Riccio M, Santi S, Galeotti C, Ambrosetti DC, Melli M : New insights into the molecular basis of progressive myoclonus epilepsy: a multiprotein complex with cystatin B. Hum Mol Genet 2002; 11: 2941–2950.
Acknowledgements
We thank Paula Hakala, Teija-Tuulia Toivonen and Rita Medek for technical assistance, Jukka Kallijärvi, Nina Aula and Jorma Palvimo for advice, Dicky Halley for contribution in identifying the Gln71Pro mutation and Liisa Laakkonen for discussions. This study was supported by the Academy of Finland (Projects 43029 and 50011), the Sigrid Jusélius Foundation, the Folkhälsan Research Foundation, the Federal Ministry of Education and Research, Bonn, Germany (Project NBL 3/2-03), Dutch Epilepsy Fund (Grant 02-04) and the EC (Project QLG3-CT-2000-01405) and was partly carried out in the Center of Excellence in Disease Genetics of the Academy of Finland (Project 44870, Finnish Centre of Excellence Programme 2000-2005).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alakurtti, K., Weber, E., Rinne, R. et al. Loss of lysosomal association of cystatin B proteins representing progressive myoclonus epilepsy, EPM1, mutations. Eur J Hum Genet 13, 208–215 (2005). https://doi.org/10.1038/sj.ejhg.5201300
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201300
Keywords
This article is cited by
-
Cathepsin B abundance, activity and microglial localisation in Alzheimer’s disease-Down syndrome and early onset Alzheimer’s disease; the role of elevated cystatin B
Acta Neuropathologica Communications (2023)
-
Stefin B deficiency reduces tumor growth via sensitization of tumor cells to oxidative stress in a breast cancer model
Oncogene (2014)
-
New neuropathological findings in Unverricht–Lundborg disease: neuronal intranuclear and cytoplasmic inclusions
Acta Neuropathologica (2011)
-
Cystatin B: mutation detection, alternative splicing and expression in progressive myclonus epilepsy of Unverricht-Lundborg type (EPM1) patients
European Journal of Human Genetics (2007)