Abstract
Cells precisely control the formation of dynamic actin cytoskeleton networks to coordinate fundamental processes, including motility, division, endocytosis and polarization. To support these functions, actin filament networks must be assembled, maintained and disassembled at the correct time and place, and with proper filament organization and dynamics. Regulation of the extent of filament network assembly and of filament network organization has been largely attributed to the coordinated activation of actin assembly factors through signalling cascades. Here, we discuss an intriguing model in which actin monomer availability is limiting and competition between homeostatic actin cytoskeletal networks for actin monomers is an additional crucial regulatory mechanism that influences the density and size of different actin networks, thereby contributing to the organization of the cellular actin cytoskeleton.
Similar content being viewed by others
Main
Cells coordinate the formation of diverse actin filament (F-actin) networks from a general cytoplasmic pool of components, including monomeric actin (G-actin), at the correct time and place to facilitate crucial functions such as cytokinesis, motility and endocytosis (Fig. 1). F-actin adopts two main architectures, whose formation is mediated by distinct actin nucleators. The actin-related protein 2/3 (ARP2/3) complex facilitates the formation of dense, branched F-actin networks, which are necessary for generating pushing forces capable of deforming the cell membrane to promote processes like motility and endocytosis (Fig. 1Aa,Ba). Conversely, formins and Ena/VASP (Enabled/vasodilator-stimulated phosphoprotein)-homology proteins assemble linear actin filaments, which can be bundled to create networks such as the cytokinetic contractile ring, membrane-protruding filopodia and polarizing F-actin bundles (Fig. 1Ab–d,Bb,c). F-actin networks adopt specific architectures and dynamics dictated by the action of diverse sets of specific actin-binding proteins (ABPs) to perform their particular cellular tasks1,2 (Fig. 1).
A | Representative F-actin network organization in a motile mammalian cell. Aa | The actin-related protein 2/3 (ARP2/3) complex generates branched F-actin networks, such as lamellipodia and endocytic vesicles. Ab–e | Formins and Ena/VASP (Enabled/vasodilator-stimulated phosphoprotein)-homology proteins assemble F-actin that are found in linear bundles, such as filopodia (parts Ab–d) and focal adhesions (part Ae). Af | Myosin motors are associated with contractile antiparallel F-actin bundles, including stress fibres. B | Fission yeast cells contain three principal F-actin networks generated by different actin assembly factors. Ba | The Arp2/3 complex generates branched F-actin networks for endocytic actin patches. Bb | Formin 3 (For3) generates parallel F-actin bundles for polarizing actin cables. Bc | The formin Cdc12 generates antiparallel F-actin bundles for the cytokinetic contractile ring.
The size and density of F-actin networks seem to be critical for their proper function. For example, fission yeast contains three major F-actin networks, each composed of F-actin generated by a specific actin assembly factor (Fig. 1B). Endocytic actin patches require the Arp2/3 complex, whereas the formins Cdc12 and formin 3 (For3) assemble F-actin for the cytokinetic contractile ring and polarizing actin cables1. The amount of actin incorporated into endocytic patches and the contractile ring is remarkably consistent from network to network and from cell to cell, varying less than 50% between each structure3,4. Furthermore, experimental manipulation of the size of actin networks typically correlates with functional defects. For example, depletion of the formin mDIA2 (also known as DIAPH3) causes a 20% decrease of F-actin in the contractile ring of fibroblasts, which is coupled with cytokinesis defects5. Localized inhibition of Arp2/3 complex induces a change in direction of cell motility in fish keratocytes6, and general disruption of the ARP2/3 complex in fibroblasts coincides with the disappearance of lamellipodia7,8,9,10,11,12, a decrease in cell migration speed7,12 and reduced cell spreading efficiency10. Interestingly, this decrease in migration speed was not observed in ARPC3−/− fibroblasts (which are missing the ARP2/3 complex) but instead the motility directionality was severely decreased, indicating that various cells can also respond differently to the perturbation of their actin networks11. As another example, expression of constitutively active formin constructs in budding yeast increases actin cable number and causes defects in polarized cell growth13,14.
How does the cell generate diverse F-actin networks of proper functional sizes within a common cytoplasm? It has been largely accepted that the amount of actin assembly is regulated primarily by the activation of actin assembly factors via signalling cascades, such as those involving RHO GTPases (Fig. 2). Here, we offer an updated model, in which the free cytoplasmic G-actin content is limited and therefore becomes depleted as cytoskeletal networks grow, thereby limiting the size and density of other (competing) F-actin networks. The release of G-actin through network disassembly is thus critical for promoting the subsequent growth of other coexisting networks and for the assembly of new networks. We propose that F-actin networks exist in collective homeostasis, whereby their size is controlled, at least in part, through this competition. Traditionally, actin homeostasis has referred to the general ratio of G-actin to F-actin2, without taking into account that F-actin is distributed between distinct networks with different functions. The regulation of diverse actin networks through competition for G-actin has received little attention, but a substantial amount of recent experimental data indicates that competition is in fact an important additional regulator of actin cytoskeletal network size, thereby supporting our model.
Signalling cues bind membrane receptors, activating RHO GTPase signalling pathways. RHO GTPases activate actin assembly factors directly, with RHOA·GTP directly relieving autoinhibition of formins, or indirectly, through nucleation-promoting factors like Wiskott–Aldrich syndrome protein (WASP), which activates the actin-related protein 2/3 (ARP2/3) complex following its own activation, which is induced by CDC42·GTP binding. Actin assembly factors then facilitate the nucleation and/or elongation of G-actin into branched (ARP2/3 complex) or linear (formins) actin filaments. A reserve of unassembled actin is maintained by proteins that prevent nucleation (profilin) or nucleation and elongation (thymosin β4). F-actin networks 'age' through ATP hydrolysis to ADP on actin monomers incorporated into the filament, which leads to filament disassembly. Disassembly of filaments is also facilitated by a suite of severing and depolymerizing factors, and this depolymerization collectively replenishes the pool of G-actin. ADF, actin-depolymerizing family; AIP1, actin-interacting protein 1; CA, acidic domain; CAP, adenylyl cyclase-associated protein (also known as SRV2); DAD, Diaphanous autoregulatory domain; DID, DAD-interacting domain; FH, formin homology; GBD, GTPase-binding domain; PP, polyproline domain; V, WH2 domain.
Classic view of actin turnover
Here, we define F-actin network size as the number of actin subunits a network contains. A network's size is determined by the cumulative difference between the amount of incorporation and dissociation of G-actin, which in turn is controlled by the regulation of actin assembly factors through signalling pathways (Fig. 2). F-actin polymerization occurs principally by the addition of ATP-bound G-actin to the dynamic barbed end rather than to the slow-growing pointed end15. Upon barbed-end subunit addition, ATP is hydrolysed rapidly (half-life: ∼2 seconds) to become ADP and inorganic phosphate (Pi)16. A slower release of Pi follows (half-life: ∼6 minutes), resulting in ADP- F-actin15. F-actin elongates, and the network expands through the addition of more ATP-bound G-actin onto the free barbed ends (Fig. 2); 'older' ADP-bound F-actin networks are disassembled by severing and depolymerization (Fig. 2). These aspects are described further in Box 1.
The initiation of F-actin network assembly is primarily controlled by signalling pathways, and in the classical view it is believed that signalling principally regulates actin cytoskeleton organization by promoting polymerization within defined networks. Signals initiate from external stimuli, such as cell–cell junctions and chemical cues (Fig. 2), or internal stimuli, such as the activation of formin-mediated F-actin assembly by mictrotubules17,18,19,20. Because the main actin assembly factors — the ARP2/3 complex and the formins — are normally inhibited and therefore intrinsically inactive, unwanted F-actin assembly is prevented (Fig. 2). Only on localized activation mediated by small RHO GTPases do these proteins engage in actin polymerization, and this controls spatial and temporal F-actin network assembly21,22,23,24 (Fig. 2). Three well-studied RHO GTPases (RAC1, CDC42 and RHOA) are known to be involved in actin nucleator activation. For example, RAC1 and CDC42 release the inhibition of the ARP2/3 complex activators WAVE (also known as WASF) and Wiskott–Aldrich syndrome protein (WASP)21,25,26,27 (Fig. 2). Conversely, RHOA releases autoinhibition of some formins, such as mDIA1 (also known as DIAPH1)24,28,29 (Fig. 2).
It must be noted that the role of signalling pathways in actin assembly regulation is much more complicated than described here — it can be highly context-dependent, and a considerable amount of crosstalk among different pathways can occur. For example, CDC42 can activate both the ARP2/3 complex and formin-like 1 (FMNL1) in macrophages, and RAC1 may activate both WAVE and Dictyostelium discoideum formin dDia2 (also known as ForH)30,31.
Competition for the G-actin pool
In addition to signalling, other factors and mechanisms can potentially contribute to the regulation of actin polymerization within networks and can affect the distribution of actin between them. As the total amount of actin synthesized in a given cell is believed to remain constant32, it is possible that coexisting F-actin networks are engaged in a competitive crosstalk whereby they compete for a limiting pool of G-actin. Although some data suggest that G-actin is not limiting (discussed below), there is a growing body of evidence that internetwork competition for G-actin is important. We therefore further propose a new model of the regulation of cellular F-actin networks, in which — in addition to signalling-based actin regulators — competition between homeostatic F-actin networks for a limited amount of G-actin is critical for regulating network size and density33.
Basic concept of internetwork competition. The concept that a limiting number of building blocks contributes to the regulation of the size and density of organelles, such as centrosomes, flagella and mitotic and meiotic spindles is well documented and has been the subject of several recent reviews34,35. In these examples, the limited availability of components typically restricts the size or density of a specific organelle or structure (such as a mitotic spindle or the nucleus). A limited actin pool represents a more sophisticated mechanism of size regulation, as functionally diverse networks with different architectures compete for the same resource.
In addition to the classic signalling-based hypothesis of actin turnover (Fig. 2), we propose a modified model including competition for G-actin that posits that generation of new F-actin networks also depends on the disassembly of older networks to regenerate the G-actin pool. For example, the size of the G-actin pool has been shown to be important for multiple actin-based processes, such as the migration of PtK2 cells and the in vitro reconstituted motility of the bacterium Listeria monocytogenes36,37. The concentration of G-actin is crucial for F-actin network assembly, as the nucleation rate of assembly factors, such as the ARP2/3 complex and the formins, is enhanced with an increase in the concentration of free G-actin29,38. Furthermore, an experimental increase of G-actin concentration induces ARP2/3 complex- and formin-mediated actin assembly in fission yeast and fibroblasts, resulting in a twofold increase of ARP2/3 complex-mediated actin patches and a fivefold increase in processive formin–mDIA1 molecules, respectively33,39,40,41.
If homeostatic F-actin networks compete for G-actin, a simple prediction is that the disruption of one network will increase the amount of G-actin consumed by the remaining networks (Fig. 3a,b). A systematic test of competition among F-actin networks has recently been performed in fission yeast33. However, there are numerous other examples published over the past 20 years that indicate that modification of the amount of F-actin in one type of network has an effect on the size and density of other networks, suggesting competitive crosstalk between the networks. These examples are briefly described below.
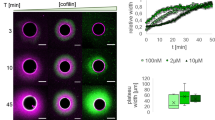
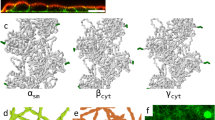
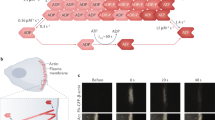
a,b | Representative graphs illustrating our model of F-actin network homeostasis. Network sizes are simulated deterministically using coupled rate equations, and each network is distinguished by a set of parameter values. The starting values of fraction of total actin are based on estimates from fission yeast33. As these graphs are illustrative, time is presented in arbitrary units (a.u.). a | Model of actin reorganization on branched network disassembly. The disassembly of an actin-related protein 2/3 (ARP2/3) complex-mediated network (of branched F-actin) via genetic or small molecule inhibitor perturbation releases a pool of G-actin that is incorporated into remaining formin- and or Ena/VASP (Enabled/vasodilator-stimulated phosphoprotein)-homology protein-mediated linear F-actin networks. Through ARP2/3 complex-mediated branched F-actin network disassembly, G-actin concentration initially increases and then is followed by a reduction owing to its incorporation in remaining F-actin networks. b | Model of actin homeostasis during cytokinetic ring assembly and disassembly. Assembly of the formin-mediated contractile ring requires a reduction in ARP2/3 complex-mediated F-actin networks (such as endocytic actin patches in fission yeast). The amount of F-actin incorporated into the ring will equal its decrease in actin patches. After ring disassembly, the level of actin consumed by actin patches is restored. In this example, the assembly of a formin-mediated contractile ring during mitosis coincides with a partial disassembly of ARP2/3 complex-mediated endocytic actin patches, necessary for the incorporation of G-actin in the formin-mediated contractile ring. c–e | Examples of F-actin network homeostasis in cells and in vivo evidence for internetwork competition. c | Fission yeast expressing the general F-actin marker Lifeact–GFP. Wild-type cells (left panel) contain three principal F-actin networks: endocytic actin patches (mediated by the Arp2/3 complex), polarizing actin cables and contractile rings (mediated by formins). Inhibition of the Arp2/3 complex using the small molecule inhibitor CK-666 induces the disappearance of patches, coupled with an accumulation of actin in formin-associated networks (middle panel). Genetic inhibition of formin-mediated F-actin networks induces an increase in the number of Arp2/3 complex-mediated endocytic patches (right panel). d | A fibroblast expressing the general F-actin marker Lifeact–EGFP (left panel). Disappearance of lamellipodia by ARP2/3 complex sequestration through injection of the acidic domain (WCA) of WAVE1 in the presence of constitutively active RAC1 (WCA/RAC) induces an increase in formin- and/or Ena/VASP-generated filopodia (right panel). e | A Drosophila melanogaster S2 culture cell expressing actin–GFP (left panel). Inhibition of the Arp2/3 complex by the addition of Arp2 double-stranded RNA (dsRNA) induces a disappearance of Arp2/3 complex-mediated lamellipodia and an increase in Ena/VASP- and/or formin-mediated filopodia formation (right panel). Part c is adapted with permission from Ref. 33, Elsevier. Part d is republished with permission of American Society of Cell Biology, from Ref. 47; permission conveyed through Copyright Clearance Center, Inc. Part e is modified from ©2013 Ingerman et al. Journal of Cell Biology. 206:763–777. doi:10.1083/jcb.201211069.
Experimental evidence for competition. Modification of actin incorporation into dendritic networks generated by the ARP2/3 complex directly influences actin incorporation into linear networks generated by formins and Ena/VASP. As shown in yeast, pharmacological depletion of Arp2/3 complex-mediated endocytic actin patches causes a 20-fold increase in actin assembled into formin-associated actin cables and contractile rings33,42,43 (Fig. 3c). Conversely, increasing endocytic actin patch number by overexpressing the Arp2/3 complex activator Las17 decreases formin-mediated actin cables by 40%13. Similarly, modification of formin-mediated actin assembly affects Arp2/3 complex-mediated networks. The inhibition of formins and the disappearance of their associated networks (contractile ring and polarizing actin cables) increases the consumption of actin by Arp2/3 complex-mediated actin patches by 50%33,44,45 (Fig. 3c). Overexpressing constitutively active formin stimulates the excessive formation of formin-mediated cables, coupled with a 35% decrease in Arp2/3 complex-mediated endocytic patches13. Notably, the aforementioned studies have been performed with a large range of fluorescent F-actin labels (including Lifeact, phalloidin, calponin homology domain, antibodies and actin fused with fluorescent proteins), excluding the possibility that the reported phenotypes are simply unintended artefacts caused by F-actin biomakers46.
F-actin network homeostasis is also important in multicellular systems, and the inhibition of ARP2/3 complex-mediated networks often leads to an increase in the density of formin- and/or Ena/VASP-mediated structures, such as filopodia (Fig. 3d,e). For example, depletion or inactivation of the ARP2/3 complex in fibroblasts results in the disappearance of lamellipodia and an increase in filopodia formation, probably by incorporation of the released pool of G-actin by formins or Ena/VASP7,12,47,48 (Fig. 3d). A similar ARP2/3 complex-inactivation phenotype has been reported in other cell types, such as Drosophila melanogaster S2, human osteosarcoma U2OS, rat adenocarcinoma MTLn3, coelomocytes and Aplysia californica neuronal growth cones49,50,51,52,53,54 (Fig. 3e). Interestingly, in A. californica neuronal growth cones, ARP2/3 complex inhibition does not increase the length of filopodia but instead increases the rate of both filopodia elongation and retrograde flow54. An increase in G-actin concentration after ARP2/3 complex inhibition could account for this phenotype. This example suggests that competition between F-actin networks for G-actin is relevant even without changes in filament length and corresponding network size, and therefore competition for G-actin may be critical for most F-actin networks. For example, microvilli, which depend on F-actin assembled by the WH2 domain family actin assembly factor Cordon-bleu, are longer when the ARP2/3 complex is inhibited55.
In addition to the interference with distinct F-actin networks, disruption of F-actin disassembly mediated by proteins of the actin-depolymerizing factor (ADF)/cofilin family affects the reincorporation of G-actin into different networks, most likely through interference with the proper regeneration of the G-actin pool (alternatively, ADF/cofilin is also thought to have a direct role in F-actin assembly by generating new barbed ends through severing56,57). For example, F-actin severing by ADF/cofilin has been shown to promote the formation of migratory cell lamellipodia in mammalian cells58,59. Conversely, depletion of ADF/cofilin in fission yeast prevented endocytic actin patch disassembly and contractile ring assembly33,60.
In these examples from yeast and animal cells, actin assembly factor activity was artificially modulated (Fig. 3c–e). An important question is whether cells use competition to intrinsically regulate F-actin-dependent processes. It seems likely that this is the case, as the assembly of formin-mediated contractile rings during division in fission yeast correlates with a decrease in actin incorporated into ARP2/3 complex-mediated endocytic actin patches61. Nevertheless, further studies are needed to determine how cells use homeostasis-driven internetwork competition to regulate F-actin assembly. Additional studies are also required to determine whether other actin regulators, such as the ABP profilin12,39, may also be limiting and contribute to the size regulation of competing networks.
Mediators of competition
If F-actin networks are in competition for a limited pool of G-actin, how do cells actively tune this competition to spatially and temporally favour particular networks? Activation of actin nucleators at particular times and places via distinct signalling pathways (Fig. 2) presumably plays a major part in how cells regulate competition between F-actin networks. For example, specific formin isoforms are activated during mitosis to generate the cytokinetic contractile ring1,62, and these must compete with other F-actin networks for G-actin to ensure robust assembly of the ring. A major question then is how G-actin is properly distributed between F-actin networks competing for the pool of monomers. Potentially, any ABP that differentially affects competitive F-actin networks could have a role in this process and mediate the competition and its outcome. We focus below on profilin, capping protein and myosin II because of recent evidence of their participation in F-actin network competition, but we suspect that other actin regulators are also important.
Role of profilin. Recent articles highlight the major contribution of the small ABP profilin to the proper segregation of G-actin between competing networks. Profilin has long been considered to be a general actin cytoskeleton 'housekeeping' protein that helps maintain a pool of unpolymerized actin that is equally usable by diverse F-actin networks2. However, recent work revealed that, through opposing effects on competing actin assembly factors, profilin potentially constitutes a molecular switch that mediates network competition. Profilin enhances formin-mediated actin elongation up to 15-fold63,64,65 (Fig. 4) and may slightly increase Ena/VASP-mediated filament elongation66,67,68. Conversely, profilin inhibits ARP2/3 complex-mediated nucleation (branch formation)12,39,69,70,71 (Fig. 4). Therefore, the presence of high levels of profilin favours G-actin incorporation into F-actin networks consisting of linear F-actin, such as formin-generated filaments in the contractile ring and polarizing actin cables in fission yeast or filopodial protrusions in fibroblasts made by Ena/VASP proteins12,39 (Fig. 4).
Extracellular stimuli activate RHO GTPase signalling pathways that activate formin-, actin-related protein 2/3 (ARP2/3) complex- and/or Ena/VASP (Enabled/vasodilator-stimulated phosphoprotein)-homology protein-mediated F-actin networks, promoting actin assembly within these networks. These networks are in competition for G-actin from a common limited pool. Actin associated with sequestering proteins or incorporated into other networks (such as stress fibres stabilized by myosin motors) cannot participate in competition, and this has a large effect on the internetwork competition by further restricting the amount of actin available for polymerization. Profilin favours incorporation of G-actin into formin- and Ena/VASP-dependent F-actin networks by enhancing filament elongation (but at the same time inhibits formin-mediated nucleation of new filaments), while inhibiting ARP2/3 complex activation by competing with the Wiskott–Aldrich syndrome protein (WASP) VCA domain (WASP-VCA) for G-actin binding, and tips the balance towards the formation of linear actin networks. Capping protein competes with formins and Ena/VASP for barbed ends, whereas capping protein rapidly binds free barbed ends generated by the ARP2/3 complex. Disassembly of F-actin networks replenishes the G-actin pool, allowing more-robust polymerization and growth of networks.
Interestingly, the overexpression of profilin favours the formation of formin-mediated contractile rings, whereas the overexpression of actin favours the formation of ARP2/3 complex-mediated patches33,39. These results indicate that the ratio of profilin to G-actin may govern the competition between ARP2/3 complex- and formin-mediated F-actin networks. Two non-mutually exclusive mechanisms have been proposed to contribute to the inhibition of ARP2/3 complex-mediated branch formation by profilin. First, profilin may directly compete with the ARP2/3 complex activator WASP for binding to G-actin39,71,72, although this mechanism may not explain the full extent of ARP2/3 complex inhibition70. Second, profilin association with F-actin barbed ends may inhibit ARP2/3 complex binding70 and thereby ARP2/3 activity and branching at barbed ends; however, the generally accepted view is that branches are primarily formed from the side of actin filaments rather than from the barbed end15. In addition, phosphorylation of mouse profilin 1 at Tyr129 increases its affinity to G-actin and its ability to compete with actin-sequestering thymosin β4 for G-actin73, a mechanism that is expected to further contribute to the favouring of formin- over ARP2/3 complex-mediated actin assembly by profilin. At extremely high concentrations, free profilin may also compete with barbed end-binding proteins, such as capping protein, formins and Ena/VASP70, thereby allowing profilin to similarly inhibit competing actin assembly factors. However, the concentrations of profilin and unassembled G-actin in most cell types are similar15,39, which is why profilin probably favours formins and Ena/VASP over the ARP2/3 complex in most cells12,39.
We propose that regulation of profilin activity and/or localization probably represents an important mechanism by which cells control the balance between ARP2/3 complex- and formin (or Ena/VASP)- mediated F-actin assembly. For example, fission yeast profilin seems to be locally concentrated at the tips of interphase cells, thereby potentially favouring For3- over Arp2/3 complex-mediated actin assembly at these sites61. Similarly, profilin may be more concentrated at the division site of dividing fission yeast cells, facilitating Cdc12-mediated actin assembly61. In fibroblasts, profilin may be enriched at the ARP2/3 complex-mediated lamellipodia74, where it could limit ARP2/3 complex-mediated nucleation, thereby regulating the extension of these protrusions. Most multicellular eukaryotes have multiple profilin isoforms that are expressed at different times, at varied cellular locations and in specific tissues. At the same time, these different isoforms have been differentially tailored to facilitate actin assembly for distinct F-actin networks75,76. Further experiments are necessary to determine whether and how cells actively modulate profilin activity to govern F-actin network competition.
Linear versus dendritic array competition involves barbed-end capping. Barbed-end cappers, such as capping protein, are important regulators of F-actin network size (Fig. 4). Capping protein has a high affinity (∼0.1 nM) for free F-actin barbed ends77, competing with formins and Ena/VASP for F-actin barbed ends66,78,79,80,81 and promoting the formation of short-branched filaments in ARP2/3 complex-mediated F-actin networks15. Interestingly, deletion of capping protein in budding and fission yeast increases actin incorporation into Arp2/3 complex-mediated patches by at least 35%82,83,84,85. Capping protein localizes to ARP2/3 complex-mediated actin patches, where it presumably associates with F-actin barbed ends and keeps filaments short by terminating their elongation (Fig. 4). In the absence of barbed-end capping, the uninterrupted growth of F-actin is expected to rapidly deplete the monomer pool, presumably decreasing the availability of G-actin for formin-dependent processes. In agreement, the number of formin-mediated actin cables decreases dramatically in the absence of capping protein in fission yeast and budding yeast82,85,86. However, the function of capping protein may be more complicated and might be context dependent, as its depletion in melanoma cells and neurons decreases F-actin in ARP2/3 complex-mediated lamellipodia and increases formin- and Ena/VASP-mediated filopodia number fivefold87,88, contrary to what has been observed in yeast. In line with this, capping protein has been proposed to enhance ARP2/3 complex-mediated branching by steering monomers to new branch nuclei rather than to existing barbed ends89.
It is possible that the observed differences in the role of capping protein in regulating the size of competing networks depends on whether the competing ARP2/3 complex- and formin- or Ena/VASP-mediated networks are physically linked. An example of such physically linked networks is found in the migratory protrusions of mammalian cells, in which filopodia are thought to elongate from the lamellipodia90. In this case, inhibition of capping protein may facilitate recruitment of Ena/VASP and formins to branched networks, which would subsequently drive actin assembly into linear networks and cause filopodia extension, thereby decreasing the pool of G-actin available for the ARP2/3 complex. Regulation of capping protein activity might therefore also be critical for balancing competitive F-actin networks. Notably, cells harbour various regulators of capping protein, including steric inhibitors, such as the protein myotrophin (also known as V1) or the phospholipid phosphatidylinositol-4,5-bisphosphate (PtdIns(4,5)P2), which bind directly to capping protein and inhibit its capping activity91. Furthermore, localization and activity of capping protein can be influenced by association with CARMIL (also known as LRRC16A), which contains a capping protein-interaction motif92.
Actomyosin networks contribute to internetwork competition. The molecular motor myosin II, which associates with F-actin to form contractile actomyosin bundles such as cell stress fibres, may also be an important regulator of F-actin network homeostasis and thus may influence internetwork competition. A recent study uncovered a competitive relationship between ARP2/3 complex-mediated networks and actomyosin networks in epithelial cells that may help regulate the transition of epithelial cells from static cells to migratory polarized cells93. Inhibition of non-muscle myosin II by the small molecule inhibitor blebbistatin increases the concentration of G-actin by approximately 50% within 15 minutes; this G-actin is subsequently consumed by ARP2/3 complex-mediated networks to trigger cell polarization and motility (Fig. 4). These observations may be relevant, as non-muscle myosin II can be activated or inhibited by phosphorylation of specific residues in both its regulatory light chain and its heavy chain94. In addition to its important role in force generation, myosin II may therefore be a molecular switch that controls the assembly of actin into actomyosin bundles over ARP2/3 complex-mediated lamellipodia. In addition to myosin II, another component of actomyosin bundles, tropomyosin, contributes to the competitive crosstalk between actin networks. For example, in general, tropomyosins seem to be important regulators of actin filament identity and dynamics, and they have been implicated in the inhibition of cofilin-mediating binding and severing95. In addition, tropomyosins have been suggested to bind preferentially to formin- over ARP2/3 complex-mediated F-actin networks96,97, and microinjection of skeletal tropomyosin into PtK1 epithelial cells has been shown to inhibit ARP2/3 complex-mediated lamellipodia while promoting filopodia formation98.
Homeostatic control of actin
Traditionally, actin homeostasis has referred to the maintenance of a stable ratio of total F-actin to G-actin in cells2. The principal driving force of F-actin network formation is the signalling pathway-mediated spatial and temporal activation of actin assembly factors. However, how the signalling activates particular networks leading to their expansion while at the same time allowing the maintenance of actin homeostasis has remained elusive.
A model of homeostatic internetwork competition for G-actin. Based on recent observations from primary data discussed above, we propose here an updated view of actin homeostasis, whereby competition between coexisting actin networks for a limited G-actin pool is an additional important factor regulating the distribution of actin among these networks and thus also controlling their relative growth and size (Figs 3,4). We propose that, owing to the limited G-actin availability, to support growth of defined networks, activated assembly factors require constant replenishment of G-actin, which occurs though G-actin release by the steady disassembly of other existing networks15. In our model, this internetwork competition for limited G-actin is critical for regulating F-actin network size and density and can be influenced by specific regulatory ABPs. Because profilin inhibits ARP2/3 complex-mediated branching while enhancing formin-mediated elongation, and formins and Ena/VASP antagonize capping protein, profilin and capping protein prevent G-actin incorporation into ARP2/3 complex-mediated F-actin networks more than formin- and Ena/VASP-mediated F-actin networks12,39,80,81,83,84,85. Myosin II can stabilize and sequester actin subunits in F-actin network reserves, which upon signalling can quickly release G-actin that subsequently incorporates into ARP2/3 complex-mediated lamellipodia and trigger polarized epithelial cell motility93,94. We therefore hypothesize that modulating the activity of these different ABPs by signalling pathways could serve to direct the assembly of actin into specific F-actin networks.
Alternative possibilities and validity of the proposed model. The model we propose here provides an attractive explanation of homeostatic control of F-actin networks in vivo. However, future studies are needed to better understand the principles governing the regulated distribution of actin between coexisting networks.
Traditionally, it was thought that the number of free F-actin barbed ends, not the G-actin pool, is the limiting factor for the regulation of F-actin network size and density. For example, Acanthamoeba castellanii and neutrophil extracts contain filaments that do not elongate because their barbed ends are strongly capped, and the addition of activated RHO GTPase CDC42 to these extracts triggers ARP2/3 complex-mediated nucleation of new branched filaments that rapidly assemble99,100. This indicates that the limitation of barbed-end availability might indeed play an important part in the regulation of network size. However, it is noteworthy that this regulatory mechanism is not in direct contradiction with our model and that both mechanisms might be at play in vivo.
Our model of F-actin networks competing for a limited pool of G-actin postulates that the amount of actin incorporated into one network will indirectly and inversely influence the amount of actin incorporated into other networks, thereby affecting their size. However, this relationship between F-actin networks was not observed in several studies. For example, in melanoma B16-F1 cells, the induced disappearance of ARP2/3 complex-mediated lamellipodia by RNAi knockdown of WAVE or the ARP2/3 complex does not seem to modify the number of Ena/VASP- and/or formin-generated filopodia9. In Jurkat T cells, the inhibition of the ARP2/3 complex via the small molecule inhibitor CK-666 induces a significant decrease in formin-mediated filopodia formation101. In D. melanogaster BG2-c2 cells, knockdown of the ARP2/3 complex activator Scar leads to disappearance of the lamellipodia coupled with an eightfold decrease in the total F-actin concentration, suggesting that G-actin released by disassembled lamellipodia is not fully reincorporated into the remaining F-actin networks, such as filopodia102. Finally, in D. discoideum, depletion of the formin ForA, which generates F-actin networks at the cell cortex, is coupled with an increase in cell motility rate103. The authors concluded that the motility rate increase was dependent on myosin II activity, rather than on a rise of actin incorporation into ARP2/3 complex-mediated networks. It is thus possible that the increase of G-actin concentration is not sufficient to incorporate into existing F-actin networks.
So, how can these observations be reconciled in light of the model we propose? It might be that internetwork competition for G-actin may not contribute to actin network regulation in all systems and/or cellular contexts, perhaps because actin is not limiting in all systems or because additional regulatory mechanisms are at play that dilute the importance of G-actin limitation. For example, in the lamellipodia of differentiated neuroblastoma cells, G-actin bound to thymosin β4 targets formin-mediated polymerization instead of the ARP2/3 complex, indicating that the ratio of G-actin to G-actin-binding proteins, including profilin and thymosin β4, is a critical buffer of changes in unpolymerized actin104. It is also likely that the nature (and the associated timescale in particular) of the methods used to inhibit particular actin network could have an impact on their effects on F-actin network competition and therefore the final phenotype observed. Long-term perturbations, such as genetic mutants, could allow adaptations (such as changes in actin expression) over numerous generations that mask homeostatic competition. Conversely, acute perturbations, such as the use of small molecule inhibitors, will probably elicit only short-lasting homeostatic responses, which can be overlooked or misinterpreted. As an example of different phenotypes resulting from distinct perturbations, rapid small molecule inhibition of the Arp2/3 complex in fission yeast has a much stronger and evident effect on the formation of additional formin-mediated networks in comparison with Arp2/3 complex deletion in mutant cells33. It is also possible that some perturbations to the actin cytoskeleton are efficiently buffered, counter-balanced by additional mechanisms or cannot be easily detected owing to the nature and/or complexity of the particular redistribution event. For example, it is possible that, in some conditions, the released G-actin from inhibited ARP2/3 complex-mediated networks is redistributed to multiple formin- and Ena/VASP-mediated F-actin networks, such as focal adhesion (Fig. 1Ae) and stress fibres (Fig. 1Af), that are not as easily quantifiable as filopodia (Fig. 1Ab–d). Additionally, it has been reported that ARP2/3 complex-mediated F-actin network formation may be necessary to generate the barbed ends used to form filopodia90 and can account for an observed decrease in filopodia formation on ARP2/3 complex inhibition.
Conclusions and perspectives
We have outlined here the novel idea that G-actin is a limiting resource in F-actin network assembly. We propose that, in addition to signalling pathways, internetwork competition for this limited G-actin pool regulates network size and density. We further suggest that specific ABPs, by modulating filament assembly and disassembly rates, influence the efficiency of actin incorporation into competing networks, thereby serving as mediators (or effectors) of internetwork competition. This new competitive F-actin network homeostasis model generates several challenging questions that remain to be addressed. For example, the concept of homeostasis-driven internetwork competition for G-actin has emerged based on experiments involving artificial manipulations of F-actin networks through the use of inhibitory drugs and genetic approaches (Fig. 3c–e). Therefore, an important question is whether competition for G-actin to favour the assembly of particular networks underlies inherent, physiological processes, such as actin cytoskeleton remodelling on the initiation or cessation of cell migration. A good model to start addressing the importance of internetwork competition in the physiological context is cytokinetic contractile ring formation during mitosis in fission yeast, in which approximately 20% of the F-actin content has to be redistributed from Arp2/3 complex-mediated endocytic patches to the formin-generated contractile ring33,61. During this process, is the activation of formins by signalling sufficient to allow effective competition for G-actin with pre-existing Arp2/3 complex-mediated networks, or is a reduced amount of Arp2/3 complex activation also necessary? It will also be important to quantitatively measure how redistribution of actin to competing networks affects the loss or incorporation of ABPs into these competing networks. Do F-actin networks that expand by inhibition of competing networks incorporate more ABPs? Conversely, what happens to ABPs from inhibited networks? Finally, what are the regulatory mechanisms governing segregation of ABPs to diverse F-actin networks in cells?
Bearing in mind the important role of ABPs and their regulation for the competitive crosstalk between actin networks, it is also possible that ABPs function context-specifically and that different ABPs are required for distinct responses. In vivo and in vitro experiments have revealed that profilin favours the assembly of formin- and Ena/VASP- over ARP2/3 complex-mediated networks12,39 (Fig. 4), and cells could potentially temporally and spatially activate profilin via signalling pathways, phosphorylation and sequestration to elicit the desired changes in actin cytoskeleton organization. Furthermore, different isoforms of profilin, formin or ARP2/3 complex subunits are present in cells and may be more or less tailored for competition in different contexts. Nevertheless, it is currently not known how the cell regulates the activities of these proteins to control the size of specific networks at any given time. The next challenge will be to determine whether such regulatory mechanisms exist and how they are used to facilitate growth of particular F-actin networks.
We also predict that the contribution of ABPs to internetwork competition may be species-dependent. For example, in worms, profilin is able to stimulate F-actin elongation rates mediated by the cytokinesis-specific formin CYK-1 up to six times more than in the case of equivalent formin Cdc12 from fission yeast105. Hence, it is possible that the impact of profilin on the competition between formins and the ARP2/3 complex in these two species differs. In addition, the particular profilin-to-actin ratio or profilin-binding affinity to actin may alter profilin's contribution to competition. It will be thus interesting to investigate how the expression levels of various ABPs contribute to actin network homeostasis.
Homeostasis-driven internetwork competition for G-actin also suggests that the expression level of actin may be critical. For example, inducing fivefold changes in actin expression dramatically tips the balance of network formation in fission yeast cells33,39. Overexpressing actin favours the Arp2/3 complex, whereas lowering actin expression favours the formins, a result that seems to be largely due to altering the ratio of actin to profilin39. Surprisingly, these drastic variations of actin expression levels affect only the number of Arp2/3 complex endocytic patches, not their size33. It would be interesting to investigate how cells maintain the constant size of these structures in such differing conditions. Perhaps actin expression is carefully monitored and tightly regulated in cells so that competition for G-actin can be used to facilitate the assembly of particular F-actin networks as cells respond to changing needs. It will be important to study in more detail how cells respond to changes in actin expression, how robustly they can maintain homeostasis on changes in actin levels and whether they can actively control actin expression to regulate network competition.
Our competition model also predicts that F-actin network assembly in a cell may be perturbed on infection with bacterial pathogens, including L. monocytogenes, Shigella flexneri and Rickettsia spp., which rely on actin-mediated motility and hijack host actin to generate F-actin networks driving their own propulsion106,107,108. It will be interesting to determine whether and how these bacteria efficiently compete with host-cell F-actin networks for a limited pool of G-actin. It has been reported that L. monocytogenes cells move slower in host-cell compartments containing dense F-actin networks109, indicating that, indeed, in this context, competition for G-actin might be at play and that incorporation of actin into host networks locally depletes the G-actin pool, thereby restricting bacterial propulsion.
An extremely interesting question is whether competition between F-actin networks provides any additional benefit for the cell compared to regulation of network density and size by signalling alone. One possibility is that competition allows for consistent F-actin elongation rates and allows cells to maintain a constant G-actin concentration without the need for unnecessary and slow changes in actin expression levels. A limited pool of G-actin also avoids having excessive F-actin assembly without requiring concomitant inhibition through signalling pathways, as assembly rates into networks will naturally decrease as the cytoplasmic G-actin concentration decreases. In addition, competition mediated by ABPs, such as profilin and thymosin β4, may be necessary to ensure that formin- and Ena/VASP-mediated F-actin networks can still be generated in the presence of an excess of the ARP2/3 complex12,39,104 — a scenario probably observed in most cells. Furthermore, the rapid generation of G-actin on acute disassembly of F-actin networks, such as through the inhibition of myosins or activation of ADF/cofilin, may be necessary for rapid transitions between static and motile cells93 (Fig. 4).
Finally, the homeostatic F-actin network competition model has significant repercussions for interpreting experiments in which actin assembly factors and/or their associated F-actin networks are disrupted using small molecule inhibitors or genetic approaches. For example, disruption of ARP2/3 complex-mediated lamellipodia leads to an increase of filopodia at the leading edge of migrating cells7,12,48. One interpretation may be that ARP2/3 complex-generated branched actin filaments are not critical for filopodia assembly9. Alternatively, it is possible that the release of excess G-actin through ARP2/3 complex inhibition artificially leads to ectopic formin- and/or Ena/VASP-mediated filopodia by alternative mechanisms. Care must therefore be taken when interpreting experiments that interfere with the physiological homeostatic balance of actin, and we think that considering the possibility of the existence of the competitive crosstalk between networks can bring novel understanding of experimental data.
References
Kovar, D. R., Sirotkin, V. & Lord, M. Three's company: the fission yeast actin cytoskeleton. Trends Cell Biol. 21, 177–187 (2011).
Blanchoin, L., Boujemaa-Paterski, R., Sykes, C. & Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 94, 235–263 (2014).
Sirotkin, V., Berro, J., Macmillan, K., Zhao, L. & Pollard, T. D. Quantitative analysis of the mechanism of endocytic actin patch assembly and disassembly in fission yeast. Mol. Biol. Cell 21, 2894–2904 (2010).
Wu, J.-Q. & Pollard, T. D. Counting cytokinesis proteins globally and locally in fission yeast. Science 310, 310–314 (2005).
Watanabe, S. et al. mDia2 induces the actin scaffold for the contractile ring and stabilizes its position during cytokinesis in NIH 3T3 cells. Mol. Biol. Cell 19, 2328–2338 (2008).
Dang, I. et al. Inhibitory signalling to the Arp2/3 complex steers cell migration. Nature 503, 281–284 (2013).
Wu, C. et al. Arp2/3 is critical for lamellipodia and response to extracellular matrix cues but is dispensable for chemotaxis. Cell 148, 973–987 (2012).
Machesky, L. M. & Insall, R. H. Scar1 and the related Wiskott–Aldrich syndrome protein, WASP, regulate the actin cytoskeleton through the Arp2/3 complex. Curr. Biol. 8, 1347–1356 (1998).
Steffen, A. et al. Filopodia formation in the absence of functional WAVE- and Arp2/3-complexes. Mol. Biol. Cell 17, 2581–2591 (2006).
Nicholson-Dykstra, S. M. & Higgs, H. N. Arp2 depletion inhibits sheet-like protrusions but not linear protrusions of fibroblasts and lymphocytes. Cell Motil. Cytoskeleton 65, 904–922 (2008).
Suraneni, P. et al. The Arp2/3 complex is required for lamellipodia extension and directional fibroblast cell migration. J. Cell Biol. 197, 239–251 (2012).
Rotty, J. D. et al. Profilin-1 serves as a gatekeeper for actin assembly by Arp2/3-dependent and -independent pathways. Dev. Cell 32, 54–67 (2015).
Gao, L. & Bretscher, A. Analysis of unregulated formin activity reveals how yeast can balance F-actin assembly between different microfilament-based organizations. Mol. Biol. Cell 19, 1474–1484 (2008).
Sagot, I., Klee, S. K. & Pellman, D. Yeast formins regulate cell polarity by controlling the assembly of actin cables. Nat. Cell Biol. 4, 42–50 (2002).
Pollard, T. D., Blanchoin, L. & Mullins, R. D. Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu. Rev. Biophys. Biomol. Struct. 29, 545–576 (2000).
Blanchoin, L. & Pollard, T. D. Hydrolysis of ATP by polymerized actin depends on the bound divalent cation but not profilin. Biochemistry 41, 597–602 (2002).
Martin, S. G., Rincón, S. A., Basu, R., Pérez, P. & Chang, F. Regulation of the formin For3p by Cdc42p and Bud6p. Mol. Biol. Cell 18, 4155–4167 (2007).
Martin, S. G., McDonald, W. H., Yates, J. R. 3rd & Chang, F. Tea4p links microtubule plus ends with the formin For3p in the establishment of cell polarity. Dev. Cell 8, 479–491 (2005).
Piekny, A., Werner, M. & Glotzer, M. Cytokinesis: Welcome to the Rho zone. Trends Cell Biol. 15, 651–658 (2005).
Rincón, S. A. et al. Pob1 participates in the Cdc42 regulation of fission yeast actin cytoskeleton. Mol. Biol. Cell 20, 4390–4399 (2009).
Eden, S., Rohatgi, R., Podtelejnikov, A. V., Mann, M. & Kirschner, M. W. Mechanism of regulation of WAVE1-induced actin nucleation by Rac1 and Nck. Nature 418, 790–793 (2002).
Leung, D. W. & Rosen, M. K. The nucleotide switch in Cdc42 modulates coupling between the GTPase-binding and allosteric equilibria of Wiskott–Aldrich syndrome protein. Proc. Natl Acad. Sci. USA 102, 5685–5690 (2005).
Otomo, T., Otomo, C., Tomchick, D. R., Machius, M. & Rosen, M. K. Structural basis of rho GTPase-mediated activation of the formin mDia1. Mol. Cell 18, 273–281 (2005).
Watanabe, N. et al. p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein for Rho small GTPase and is a ligand for profilin. EMBO J. 16, 3044–3056 (1997).
Higgs, H. N. & Pollard, T. D. Activation by Cdc42 and Pip2 of Wiskott–Aldrich syndrome protein (Wasp) stimulates actin nucleation by Arp2/3 complex. J. Cell Biol. 150, 1311–1320 (2000).
Rohatgi, R. et al. The interaction between N-WASP and the Arp2/3 complex links Cdc42-dependent signals to actin assembly. Cell 97, 221–231 (1999).
Wu, Y. I. et al. A genetically encoded photoactivatable Rac controls the motility of living cells. Nature 461, 104–108 (2009).
Alberts, A. S. Identification of a carboxyl-terminal diaphanous-related formin homology protein autoregulatory domain. J. Biol. Chem. 276, 2824–2830 (2001).
Li, F. & Higgs, H. N. The mouse formin mDia1 is a potent actin nucleation factor regulated by autoinhibition. Curr. Biol. 13, 1335–1340 (2003).
Schirenbeck, A., Bretschneider, T., Arasada, R., Schleicher, M. & Faix, J. The Diaphanous-related formin dDia2 is required for the formation and maintenance of filopodia. Nat. Cell Biol. 7, 619–625 (2005).
Seth, A., Otomo, C. & Rosen, M. K. Autoinhibition regulates cellular localization and actin assembly activity of the diaphanous-related formins FRLα and mDia1. J. Cell Biol. 174, 701–713 (2006).
Leavitt, J. et al. Expression of transfected mutant beta-actin genes: alterations of cell morphology and evidence for autoregulation in actin pools. Mol. Cell. Biol. 7, 2457–2466 (1987).
Burke, T. A. et al. Homeostatic actin cytoskeleton networks are regulated by assembly factor competition for monomers. Curr. Biol. 24, 579–585 (2014).
Chan, Y.-H. M. & Marshall, W. F. How cells know the size of their organelles. Science 337, 1186–1189 (2012).
Goehring, N. W. & Hyman, A. A. Organelle growth control through limiting pools of cytoplasmic components. Curr. Biol. 22, R330–R339 (2012).
Cramer, L. P. Role of actin-filament disassembly in lamellipodium protrusion in motile cells revealed using the drug jasplakinolide. Curr. Biol. 9, 1095–1105 (1999).
Loisel, T. P., Boujemaa, R., Pantaloni, D. & Carlier, M.-F. Reconstitution of actin-based motility of Listeria and Shigella using pure proteins. Nature 401, 613–616 (1999).
Higgs, H. N., Blanchoin, L. & Pollard, T. D. Influence of the C terminus of Wiskott–Aldrich syndrome protein (WASp) and the Arp2/3 complex on actin polymerization. Biochemistry 38, 15212–15222 (1999).
Suarez, C. et al. Profilin regulates F-actin network homeostasis by favoring formin over Arp2/3 complex. Dev. Cell 32, 43–53 (2015).
Higashida, C. et al. G-actin regulates rapid induction of actin nucleation by mDia1 to restore cellular actin polymers. J. Cell Sci. 121, 3403–3412 (2008).
Higashida, C. et al. F- and G-actin homeostasis regulates mechanosensitive actin nucleation by formins. Nat. Cell Biol. 15, 395–405 (2013).
Basu, R. & Chang, F. Characterization of Dip1p reveals a switch in Arp2/3-dependent actin assembly for fission yeast endocytosis. Curr. Biol. 21, 905–916 (2011).
Nakano, K., Kuwayama, H., Kawasaki, M., Numata, O. & Takaine, M. GMF is an evolutionarily developed Adf/cofilin-super family protein involved in the Arp2/3 complex-mediated organization of the actin cytoskeleton. Cytoskeleton 67, 373–382 (2010).
Jose, M. et al. A quantitative imaging-based screen reveals the exocyst as a network hub connecting endo- and exocytosis. Mol. Biol. Cell 26, 2519–2534 (2015).
Willet, A. H. et al. The F-BAR Cdc15 promotes contractile ring formation through the direct recruitment of the formin Cdc12. J. Cell Biol. 208, 391–399 (2015).
Courtemanche, N., Pollard, T. D. & Chen, Q. Avoiding artefacts when counting polymerized actin in live cells with LifeAct fused to fluorescent proteins. Nat. Cell Biol. 18, 676–683 (2016).
Koestler, S. A. et al. Arp2/3 complex is essential for actin network treadmilling as well as for targeting of capping protein and cofilin. Mol. Biol. Cell 24, 2861–2875 (2013).
Suraneni, P. et al. A mechanism of leading edge protrusion in the absence of Arp2/3 complex. Mol. Biol. Cell 26, 901–912 (2015).
Henson, J. H. et al. Arp2/3 complex inhibition radically alters lamellipodial actin architecture, suspended cell shape, and the cell spreading process. Mol. Biol. Cell 26, 887–900 (2015).
Hotulainen, P. & Lappalainen, P. Stress fibers are generated by two distinct actin assembly mechanisms in motile cells. J. Cell Biol. 173, 383–394 (2006).
Ingerman, E., Hsiao, J. Y. & Mullins, R. D. Arp2/3 complex ATP hydrolysis promotes lamellipodial actin network disassembly but is dispensable for assembly. J. Cell Biol. 200, 619–633 (2013).
Rogers, S. L., Wiedemann, U., Stuurman, N. & Vale, R. D. Molecular requirements for actin-based lamella formation in Drosophila S2 cells. J. Cell Biol. 162, 1079–1088 (2003).
Sarmiento, C. et al. WASP family members and formin proteins coordinate regulation of cell protrusions in carcinoma cells. J. Cell Biol. 180, 1245–1260 (2008).
Yang, Q., Zhang, X.-F., Pollard, T. D. & Forscher, P. Arp2/3 complex-dependent actin networks constrain myosin II function in driving retrograde actin flow. J. Cell Biol. 197, 939–956 (2012).
Grega-Larson, N. E., Crawley, S. W., Erwin, A. L. & Tyska, M. J. Cordon bleu promotes the assembly of brush border microvilli. Mol. Biol. Cell 26, 3803–3815 (2015).
Chen, Q. & Pollard, T. D. Actin filament severing by cofilin is more important for assembly than constriction of the cytokinetic contractile ring. J. Cell Biol. 195, 485–498 (2011).
Ghosh, M. et al. Cofilin promotes actin polymerization and defines the direction of cell motility. Science 304, 743–746 (2004).
Kiuchi, T., Ohashi, K., Kurita, S. & Mizuno, K. Cofilin promotes stimulus-induced lamellipodium formation by generating an abundant supply of actin monomers. J. Cell Biol. 177, 465–476 (2007).
Kiuchi, T., Nagai, T., Ohashi, K. & Mizuno, K. Measurements of spatiotemporal changes in G-actin concentration reveal its effect on stimulus-induced actin assembly and lamellipodium extension. J. Cell Biol. 193, 365–380 (2011).
Nakano, K. & Mabuchi, I. Actin-depolymerizing protein Adf1 is required for formation and maintenance of the contractile ring during cytokinesis in fission yeast. Mol. Biol. Cell 17, 1933–1945 (2006).
Balasubramanian, M. K., Hirani, B. R., Burke, J. D. & Gould, K. L. The Schizosaccharomyces pombe cdc3+ gene encodes a profilin essential for cytokinesis. J. Cell Biol. 125, 1289–1301 (1994).
Balasubramanian, M. K., Bi, E. & Glotzer, M. Comparative analysis of cytokinesis in budding yeast, fission yeast and animal cells. Curr. Biol. 14, R806–R818 (2004).
Kovar, D. R., Harris, E. S., Mahaffy, R., Higgs, H. N. & Pollard, T. D. Control of the assembly of ATP- and ADP-actin by formins and profilin. Cell 124, 423–435 (2006).
Romero, S. et al. Formin is a processive motor that requires profilin to accelerate actin assembly and associated ATP hydrolysis. Cell 119, 419–429 (2004).
Higashida, C. et al. Actin polymerization-driven molecular movement of mDia1 in living cells. Science 303, 2007–2010 (2004).
Barzik, M. et al. Ena/VASP proteins enhance actin polymerization in the presence of barbed end capping proteins. J. Biol. Chem. 280, 28653–28662 (2005).
Pasic, L., Kotova, T. & Schafer, D. A. Ena/VASP proteins capture actin filament barbed ends. J. Biol. Chem. 283, 9814–9819 (2008).
Hansen, S. D. & Mullins, R. D. VASP is a processive actin polymerase that requires monomeric actin for barbed end association. J. Cell Biol. 191, 571–584 (2010).
Machesky, L. M. et al. Scar, a WASp-related protein, activates nucleation of actin filaments by the Arp2/3 complex. Proc. Natl Acad. Sci. USA 96, 3739–3744 (1999).
Pernier, J., Shekhar, S., Jegou, A., Guichard, B. & Carlier, M.-F. Profilin interaction with actin filament barbed end controls dynamic instability, capping, branching, and motility. Dev. Cell 36, 201–214 (2016).
Rodal, A. A., Manning, A. L., Goode, B. L. & Drubin, D. G. Negative regulation of yeast WASp by two SH3 domain-containing proteins. Curr. Biol. 13, 1000–1008 (2003).
Marchand, J.-B., Kaiser, D. A., Pollard, T. D. & Higgs, H. N. Interaction of WASP/Scar proteins with actin and vertebrate Arp2/3 complex. Nat. Cell Biol. 3, 76–82 (2001).
Fan, Y. et al. Stimulus-dependent phosphorylation of profilin-1 in angiogenesis. Nat. Cell Biol. 14, 1046–1056 (2012).
Buss, F., Temm-Grove, C., Henning, S. & Jockusch, B. M. Distribution of profilin in fibroblasts correlates with the presence of highly dynamic actin filaments. Cell Motil. Cytoskeleton 22, 51–61 (1992).
Mouneimne, G. et al. Differential remodeling of actin cytoskeleton architecture by profilin isoforms leads to distinct effects on cell migration and invasion. Cancer Cell 22, 615–630 (2012).
Neidt, E. M., Scott, B. J. & Kovar, D. R. Formin differentially utilizes profilin isoforms to rapidly assemble actin filaments. J. Biol. Chem. 284, 673–684 (2009).
Schafer, D. A., Jennings, P. B. & Cooper, J. A. Dynamics of capping protein and actin assembly in vitro: uncapping barbed ends by polyphosphoinositides. J. Cell Biol. 135, 169–179 (1996).
Winkelman, J. D., Bilancia, C. G., Peifer, M. & Kovar, D. R. Ena/VASP Enabled is a highly processive actin polymerase tailored to self-assemble parallel-bundled F-actin networks with Fascin. Proc. Natl Acad. Sci. USA 111, 4121–4126 (2014).
Bear, J. E. et al. Antagonism between ena/VASP proteins and actin filament capping regulates fibroblast motility. Cell 109, 509–521 (2002).
Shekhar, S. et al. Formin and capping protein together embrace the actin filament in a ménage à trois. Nat. Commun. 6, 8730 (2015).
Bombardier, J. P. et al. Single-molecule visualization of a formin-capping protein 'decision complex' at the actin filament barbed end. Nat. Commun. 6, 8707 (2015).
Amatruda, J. F., Cannon, J. F., Tatchell, K., Hug, C. & Cooper, J. A. Disruption of the actin cytoskeleton in yeast capping protein mutants. Nature 344, 352–354 (1990).
Berro, J. & Pollard, T. D. Synergies between Aip1p and capping protein subunits (Acp1p and Acp2p) in clathrin-mediated endocytosis and cell polarization in fission yeast. Mol. Biol. Cell 25, 3515–3527 (2014).
Kim, K., Yamashita, A., Wear, M. A., Maéda, Y. & Cooper, J. A. Capping protein binding to actin in yeast biochemical mechanism and physiological relevance. J. Cell Biol. 164, 567–580 (2004).
Kovar, D. R., Wu, J.-Q. & Pollard, T. D. Profilin-mediated competition between capping protein and formin Cdc12p during cytokinesis in fission yeast. Mol. Biol. Cell 16, 2313–2324 (2005).
Sizonenko, G. I., Karpova, T. S., Gattermeir, D. J. & Cooper, J. A. Mutational analysis of capping protein function in Saccharomyces cerevisiae. Mol. Biol. Cell 7, 1–15 (1996).
Mejillano, M. R. et al. Lamellipodial versus filopodial mode of the actin nanomachinery: pivotal role of the filament barbed end. Cell 118, 363–373 (2004).
Sinnar, S. A., Antoku, S., Saffin, J.-M., Cooper, J. A. & Halpain, S. Capping protein is essential for cell migration in vivo and for filopodial morphology and dynamics. Mol. Biol. Cell 25, 2152–2160 (2014).
Akin, O. & Mullins, R. D. Capping protein increases the rate of actin-based motility by promoting filament nucleation by the Arp2/3 complex. Cell 133, 841–851 (2008).
Svitkina, T. M. et al. Mechanism of filopodia initiation by reorganization of a dendritic network. J. Cell Biol. 160, 409–421 (2003).
Edwards, M. et al. Capping protein regulators fine-tune actin assembly dynamics. Nat. Rev. Mol. Cell Biol. 15, 677–689 (2014).
Edwards, M., McConnell, P., Schafer, D. A. & Cooper, J. A. CPI motif interaction is necessary for capping protein function in cells. Nat. Commun. 6, 8415 (2015).
Lomakin, A. J. et al. Competition for actin between two distinct F-actin networks defines a bistable switch for cell polarization. Nat. Cell Biol. 17, 1435–1445 (2015).
Vicente-Manzanares, M., Ma, X., Adelstein, R. S. & Horwitz, A. R. Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol. 10, 778–790 (2009).
Gunning, P. W., Hardeman, E. C., Lappalainen, P. & Mulvihill, D. P. Tropomyosin — master regulator of actin filament function in the cytoskeleton. J. Cell Sci. 128, 2965–2974 (2015).
Johnson, M., East, D. A. & Mulvihill, D. P. Formins determine the functional properties of actin filaments in yeast. Curr. Biol. 24, 1525–1530 (2014).
Hsiao, J. Y., Goins, L. M., Petek, N. A. & Mullins, R. D. Arp2/3 complex and cofilin modulate binding of tropomyosin to branched actin networks. Curr. Biol. 25, 1573–1582 (2015).
Gupton, S. L. et al. Cell migration without a lamellipodium. J. Cell Biol. 168, 619–631 (2005).
Mullins, R. D. & Pollard, T. D. Rho-family GTPases require the Arp2/3 complex to stimulate actin polymerizationin Acanthamoeba extracts. Curr. Biol. 9, 405–415 (1999).
Zigmond, S. H. et al. Mechanism of Cdc42-induced actin polymerization in neutrophil extracts. J. Cell Biol. 142, 1001–1012 (1998).
Young, L. E., Heimsath, E. G. & Higgs, H. N. Cell type-dependent mechanisms for formin-mediated assembly of filopodia. Mol. Biol. Cell 26, 4646–4659 (2015).
Biyasheva, A., Svitkina, T., Kunda, P., Baum, B. & Borisy, G. Cascade pathway of filopodia formation downstream of SCAR. J. Cell Sci. 117, 837–848 (2004).
Ramalingam, N. et al. A resilient formin-derived cortical actin meshwork in the rear drives actomyosin-based motility in 2D confinement. Nat. Commun. 6, 8496 (2015).
Vitriol, E. A. et al. Two functionally distinct sources of actin monomers supply the leading edge of lamellipodia. Cell Rep. 11, 433–445 (2015).
Neidt, E. M., Skau, C. T. & Kovar, D. R. The cytokinesis formins from the nematode worm and fission yeast differentially mediate actin filament assembly. J. Biol. Chem. 283, 23872–23883 (2008).
Gouin, E., Welch, M. D. & Cossart, P. Actin-based motility of intracellular pathogens. Curr. Opin. Microbiol. 8, 35–45 (2005).
Haglund, C. M., Choe, J. E., Skau, C. T., Kovar, D. R. & Welch, M. D. Rickettsia Sca2 is a bacterial formin-like mediator of actin-based motility. Nat. Cell Biol. 12, 1057–1063 (2010).
Madasu, Y., Suarez, C., Kast, D. J., Kovar, D. R. & Dominguez, R. Rickettsia Sca2 has evolved formin-like activity through a different molecular mechanism. Proc. Natl Acad. Sci. USA 110, E2677–E2686 (2013).
Lacayo, C. I. & Theriot, J. A. Listeria monocytogenes actin-based motility varies depending on subcellular location: a kinematic probe for cytoarchitecture. Mol. Biol. Cell 15, 2164–2175 (2004).
Moseley, J. B. & Goode, B. L. The yeast actin cytoskeleton: from cellular function to biochemical mechanism. Microbiol. Mol. Biol. Rev. 70, 605–645 (2006).
Hannapel, E. & van Kampen, M. Determination of thymosin β4 in human blood cells and serum. J. Chromatogr. A 397, 279–285 (1987).
Safer, D., Elzinga, M. & Nachmias, V. T. Thymosin β4 and Fx, an actin-sequestering peptide, are indistinguishable. J. Biol. Chem. 266, 4029–4032 (1991).
Carlier, M. F., Jean, C., Rieger, K. J., Lenfant, M. & Pantaloni, D. Modulation of the interaction between G-actin and thymosin β4 by the ATP/ADP ratio: possible implication in the regulation of actin dynamics. Proc. Natl Acad. Sci. USA 90, 5034–5038 (1993).
Goldschmidt-Clermont, P. J., Machesky, L. M., Doberstein, S. K. & Pollard, T. D. Mechanism of the interaction of human platelet profilin with actin. J. Cell Biol. 113, 1081–1089 (1991).
Goldschmidt-Clermont, P. J. et al. The control of actin nucleotide exchange by thymosin β4 and profilin. A potential regulatory mechanism for actin polymerization in cells. Mol. Biol. Cell 3, 1015–1024 (1992).
Kang, F., Purich, D. L. & Southwick, F. S. Profilin promotes barbed-end actin filament assembly without lowering the critical concentration. J. Biol. Chem. 274, 36963–36972 (1999).
Kaiser, D. A., Vinson, V. K., Murphy, D. B. & Pollard, T. D. Profilin is predominantly associated with monomeric actin in Acanthamoeba. J. Cell Sci. 112, 3779–3790 (1999).
Pollard, T. D. & Cooper, J. A. Quantitative analysis of the effect of Acanthamoeba profilin on actin filament nucleation and elongation. Biochemistry 23, 6631–6641 (1984).
Pring, M., Weber, A. & Bubb, M. R. Profilin–actin complexes directly elongate actin filaments at the barbed end. Biochemistry 31, 1827–1836 (1992).
Harris, E. S., Li, F. & Higgs, H. N. The mouse formin, FRLα, slows actin filament barbed end elongation, competes with capping protein, accelerates polymerization from monomers, and severs filaments. J. Biol. Chem. 279, 20076–20087 (2004).
Moseley, J. B. et al. A conserved mechanism for Bni1- and mDia1-induced actin assembly and dual regulation of Bni1 by Bud6 and profilin. Mol. Biol. Cell 15, 896–907 (2004).
Zigmond, S. H. et al. Formin leaky cap allows elongation in the presence of tight capping proteins. Curr. Biol. 13, 1820–1823 (2003).
Campellone, K. G. & Welch, M. D. A nucleator arms race: Cellular control of actin assembly. Nat. Rev. Microbiol. 11, 237–251 (2010).
Carlier, M.-F., Husson, C., Renault, L. & Didry, D. in International Review of Cell and Molecular Biology (ed. Jeon, K. W.) 290, 55–85 (Elsevier, 2011).
Goley, E. D. & Welch, M. D. The ARP2/3 complex: an actin nucleator comes of age. Nat. Rev. Mol. Cell Biol. 7, 713–726 (2006).
Goode, B. L. & Eck, M. J. Mechanism and function of formins in the control of actin assembly. Annu. Rev. Biochem. 76, 593–627 (2007).
Bernstein, B. W. & Bamburg, J. R. ADF/cofilin: a functional node in cell biology. Trends Cell Biol. 20, 187–195 (2010).
Blanchoin, L. & Pollard, T. D. Mechanism of interaction of Acanthamoeba actophorin (ADF/cofilin) with actin filaments. J. Biol. Chem. 274, 15538–15546 (1999).
McCullough, B. R., Blanchoin, L., Martiel, J.-L. & De La Cruz, E. M. Cofilin increases the bending flexibility of actin filaments: implications for severing and cell mechanics. J. Mol. Biol. 381, 550–558 (2008).
Fujiwara, I., Remmert, K. & Hammer, J. A. Direct observation of the uncapping of capping protein-capped actin filaments by CARMIL homology domain 3. J. Biol. Chem. 285, 2707–2720 (2010).
Miyoshi, T. et al. Actin turnover-dependent fast dissociation of capping protein in the dendritic nucleation actin network: evidence of frequent filament severing. J. Cell Biol. 175, 947–955 (2006).
Carlier, M.-F. et al. Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility. J. Cell Biol. 136, 1307–1322 (1997).
Hotulainen, P., Paunola, E., Vartiainen, M. K. & Lappalainen, P. Actin-depolymerizing factor and cofilin-1 play overlapping roles in promoting rapid F-actin depolymerization in mammalian nonmuscle cells. Mol. Biol. Cell 16, 649–664 (2005).
Jansen, S. et al. Single-molecule imaging of a three-component ordered actin disassembly mechanism. Nat. Commun. 6, 7202 (2015).
Mikati, M. A., Breitsprecher, D., Jansen, S., Reisler, E. & Goode, B. L. Coronin enhances actin filament severing by recruiting cofilin to filament sides and altering F-actin conformation. J. Mol. Biol. 427, 3137–3147 (2015).
Mohri, K., Vorobiev, S., Fedorov, A. A., Almo, S. C. & Ono, S. Identification of functional residues on Caenorhabditis elegans actin-interacting protein 1 (UNC-78) for disassembly of actin depolymerizing factor/cofilin-bound actin filaments. J. Biol. Chem. 279, 31697–31707 (2004).
Ono, S., Mohri, K. & Ono, K. Microscopic evidence that actin-interacting protein 1 actively disassembles actin-depolymerizing factor/cofilin-bound actin filaments. J. Biol. Chem. 279, 14207–14212 (2004).
Chaudhry, F. et al. Srv2/cyclase-associated protein forms hexameric shurikens that directly catalyze actin filament severing by cofilin. Mol. Biol. Cell 24, 31–41 (2013).
Johnston, A. B., Collins, A. & Goode, B. L. High-speed depolymerization at actin filament ends jointly catalysed by Twinfilin and Srv2/CAP. Nat. Cell Biol. 17, 1504–1511 (2015).
Yamashiro, S. et al. New single-molecule speckle microscopy reveals modification of the retrograde actin flow by focal adhesions at nanometer scales. Mol. Biol. Cell 25, 1010–1024 (2014).
Acknowledgements
The authors thank P. McCall (University of Chicago, Illinois, USA) and members of the Kovar laboratory, including J. Winkelman, J. Christensen and T. Burke for discussions and comments. Work on actin cytoskeleton regulation in the Kovar laboratory is supported by US National Institutes of Health grant R01 GM079265. C.S. has also been supported in part by the US Department of Defence and the US Army Research Laboratory's Army Research Office through a multidisciplinary university research initiative (MURI) grant, number W911NF1410403.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Suarez, C., Kovar, D. Internetwork competition for monomers governs actin cytoskeleton organization. Nat Rev Mol Cell Biol 17, 799–810 (2016). https://doi.org/10.1038/nrm.2016.106
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm.2016.106
This article is cited by
-
Glia maturation factor beta deficiency protects against diabetic osteoporosis by suppressing osteoclast hyperactivity
Experimental & Molecular Medicine (2023)
-
Mitochondrial fission links ECM mechanotransduction to metabolic redox homeostasis and metastatic chemotherapy resistance
Nature Cell Biology (2022)
-
Fascin1 empowers YAP mechanotransduction and promotes cholangiocarcinoma development
Communications Biology (2021)
-
Two macrophages, osteoclasts and microglia: from development to pleiotropy
Bone Research (2021)
-
The role of mode switching in a population of actin polymers with constraints
Journal of Mathematical Biology (2021)