Key Points
-
Toxicology is the science of poisons. The global biological responses that result from toxicant-induced molecular perturbations are kinetic and dynamic phenomena that depend on the dose of the toxicant and the period of exposure to it.
-
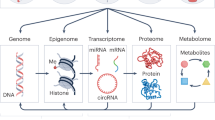
By contrast, toxicogenomics combines genetics, the analysis of genome-scale mRNA expression (transcriptomics), cell and tissue-wide protein expression (proteomics), metabolite profiling (metabolomics and metabonomics), and bioinformatics with conventional toxicology to understand the role of gene–environment interactions in disease.
-
Gene, protein and metabolite-expression profiles can be thought of as 'snapshots' of a currently poorly mapped molecular landscape. The ultimate aim of toxicogenomics is to fully map this landscape and to realize a systems toxicology.
-
Systems toxicology is the description of all the toxicological interactions within a living system. Like systems biology, systems toxicology attempts to define the behaviour and relationships of all of the components of a biological system on the premise that global-molecular data can be integrated and modelled computationally.
-
The science of systems toxicology must capture data from experiments in molecular expression and toxicology and convert them into knowledge about the toxicological responses of cells and organisms under stress. This will be accomplished through the development of knowledgebases that support the integration of data from multiple domains, as well as through computational modelling.
-
Toxicogenomics experiments have succeeded in defining several molecular signatures of exposure to drugs and chemicals and expression patterns corresponding to various histopathologies and diseases; however, much more remains to be done to delineate the toxicity response of many target organs and to predict disease outcomes following exposures to toxicants.
Abstract
Toxicogenomics combines transcript, protein and metabolite profiling with conventional toxicology to investigate the interaction between genes and environmental stress in disease causation. The patterns of altered molecular expression that are caused by specific exposures or disease outcomes have revealed how several toxicants act and cause disease. Despite these success stories, the field faces noteworthy challenges in discriminating the molecular basis of toxicity. We argue that toxicology is gradually evolving into a systems toxicology that will eventually allow us to describe all the toxicological interactions that occur within a living system under stress and use our knowledge of toxicogenomic responses in one species to predict the modes-of-action of similar agents in other species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mattes, W. B., Pettit, S. D., Sansone, S. A., Bushel, P. R. & Waters, M. D. Database development in toxicogenomics: issues and efforts. Environ. Health Perspect. 112, 495–505 (2004). This paper describes the current status of development of databases related to toxicogenomics, with an emphasis on those that are public.
Aardema, M. J. & MacGregor, J. T. Toxicology and genetic toxicology in the new era of 'toxicogenomics': impact of '-omics' technologies. Mutat. Res. 499, 13–25 (2002).
Afshari, C. A. Perspective: microarray technology, seeing more than spots. Endocrinology 143, 1983–1989 (2002).
Ulrich, R. & Friend, S. H. Toxicogenomics and drug discovery: will new technologies help us produce better drugs? Nature Rev. Drug Discov. 1, 84–88 (2002).
Fielden, M. R. & Zacharewski, T. R. Challenges and limitations of gene expression profiling in mechanistic and predictive toxicology. Toxicol. Sci. 60, 6–10 (2001).
Hamadeh, H. K., Amin, R. P., Paules, R. S. & Afshari, C. A. An overview of toxicogenomics. Curr. Issues Mol. Biol. 4, 45–56 (2002).
Tennant, R. W. The National Center for Toxicogenomics: using new technologies to inform mechanistic toxicology. Environ. Health Perspect. 110, A8–A10 (2002). The author describes the field of toxicogenomics and the goals of the NCT, as well as important concepts such as phenotypic anchoring.
Thomas, R. S. et al. Identification of toxicologically predictive gene sets using cDNA microarrays. Mol. Pharmacol. 60, 1189–1194 (2001).
Olden, K. & Guthrie, J. Genomics: implications for toxicology. Mutat. Res. 473, 3–10 (2001).
Waters, M. D., Olden, K. & Tennant, R. W. Toxicogenomic approach for assessing toxicant-related disease. Mutat. Res. 544, 415–424 (2003).
Waters, M. D. et al. Systems toxicology and the chemical effects in biological systems knowledge base. Environ. Health Perspect. 111, 811–824 (2003). This paper proposes the concept and strategy for building a true toxicogenomics knowledgebase, integrating -omics and conventional data domains, and anchoring data in sequence, phenotype and chemical structure.
Lobenhofer, E. K., Bushel, P. R., Afshari, C. A. & Hamadeh, H. K. Progress in the application of DNA microarrays. Environ. Health Perspect. 109, 881–891 (2001).
Burchiel, S. W. et al. Analysis of genetic and epigenetic mechanisms of toxicity: potential roles of toxicogenomics and proteomics in toxicology. Toxicol. Sci. 59, 193–195 (2001).
Merrick, B. A. & Tomer, K. B. Toxicoproteomics: a parallel approach to identifying biomarkers. Environ. Health Perspect. 111, A578–A579 (2003).
Petricoin, E. F. et al. Toxicoproteomics: serum proteomic pattern diagnostics for early detection of drug induced cardiac toxicities and cardioprotection. Toxicol. Pathol. 32 (Suppl. 1), 122–130 (2004).
Wilkins, M. R. et al. From proteins to proteomes: large scale protein identification by two-dimensional electrophoresis and amino acid analysis. Biotechnology (NY) 14, 61–65 (1996).
Ideker, T., Galitski, T. & Hood, L. A new approach to decoding life: systems biology. Annu. Rev. Genomics Hum. Genet. 2, 343–372 (2001).
Nurse, P. Understanding cells. Nature Biotechnol. 424, 883 (2003).
Hamadeh, H. K. et al. Methapyrilene toxicity: anchorage of pathologic observations to gene expression alterations. Toxicol. Pathol. 30, 470–482 (2002). The authors describe the methods that have been developed to anchor expression changes to pathology phenotypes, to identify expression changes seen before the detection of overt histopathology and to explore the mechanism of toxicity of a reference hepatotoxicant.
Zweiger, G. Knowledge discovery in gene-expression-microarray data: mining the information output of the genome. Trends Biotechnol. 17, 429–436 (1999).
Waring, J. F. et al. Clustering of hepatotoxins based on mechanism of toxicity using gene expression profiles. Toxicol. Appl. Pharmacol. 175, 28–42 (2001).
Mortuza, G. B., Neville, W. A., Delaney, J., Waterfield, C. J. & Camilleri, P. Characterisation of a potential biomarker of phospholipidosis from amiodarone-treated rats. Biochim. Biophys. Acta 1631, 136–146 (2003).
Kramer, J. A. et al. Acute molecular markers of rodent hepatic carcinogenesis identified by transcription profiling. Chem. Res. Toxicol. 17, 463–470 (2004). This paper identifies molecular markers seen after acute exposure to chemical carcinogens from various classes, making possible a short-term test for potential rodent carcinogens.
Heinloth, A. N. et al. Gene expression profiling of rat livers reveals indicators of potential adverse effects. Toxicol. Sci. 80, 193–202 (2004).
Hamadeh, H. K. et al. Prediction of compound signature using high density gene expression profiling. Toxicol. Sci. 67, 232–240 (2002).
Bulera, S. J. et al. RNA expression in the early characterization of hepatotoxicants in Wistar rats by high-density DNA microarrays. Hepatology 33, 1239–1258 (2001).
Bartosiewicz, M. J., Jenkins, D., Penn, S., Emery, J. & Buckpitt, A. Unique gene expression patterns in liver and kidney associated with exposure to chemical toxicants. J. Pharmacol. Exp. Ther. 297, 895–905 (2001).
Cutler, P. et al. An integrated proteomic approach to studying glomerular nephrotoxicity. Electrophoresis 20, 3647–3658 (1999).
Coen, M. et al. Integrated application of transcriptomics and metabonomics yields new insight into the toxicity due to paracetamol in the mouse. J. Pharm. Biomed. Anal. 35, 93–105 (2004).
Donald, S. et al. Hepatobiliary damage and changes in hepatic gene expression caused by the antitumor drug ecteinascidin-743 (ET-743) in the female rat. Cancer Res. 62, 4256–4562 (2002).
Fertuck, K. C., Eckel, J. E., Gennings, C. & Zacharewski, T. R. Identification of temporal patterns of gene expression in the uteri of immature, ovariectomized mice following exposure to ethynylestradiol. Physiol. Genomics 15, 127–141 (2003). This article describes the united expression changes and temporal patterns of physiological responses to an oestrogenic agent, with the aim of producing a mechanism-based understanding of effects on cell cycle, proliferation, tissue remodelling and related processes.
Fountoulakis, M. et al. Modulation of gene and protein expression by carbon tetrachloride in the rat liver. Toxicol. Appl. Pharmacol. 183, 71–80 (2002).
Hamadeh, H. K. et al. Gene expression analysis reveals chemical-specific profiles. Toxicol. Sci. 67, 219–231 (2002).
Huang, Q. et al. Assessment of cisplatin-induced nephrotoxicity by microarray technology. Toxicol. Sci. 63, 196–207 (2001).
Kramer, J. A. et al. Transcription profiling distinguishes dose-dependent effects in the livers of rats treated with clofibrate. Toxicol. Pathol. 31, 417–431 (2003).
Peterson, R. L. et al. Mechanistic toxicogenomic analysis of WAY-144122 administration in Sprague-Dawley rats. Toxicol. Appl. Pharmacol. 196, 80–94 (2004).
Ruepp, S. U., Tonge, R. P., Shaw, J., Wallis, N. & Pognan, F. Genomics and proteomics analysis of acetaminophen toxicity in mouse liver. Toxicol. Sci. 65, 135–150 (2002).
Waring, J. F. et al. Identifying toxic mechanisms using DNA microarrays: evidence that an experimental inhibitor of cell adhesion molecule expression signals through the aryl hydrocarbon nuclear receptor. Toxicology 181–182, 537–550 (2002). This paper reports for the first time the methodology of toxicant profiling; a panel of responses to 15 reference hepatotoxicants was used to identify the probable mechanism of toxicity of a novel agent. The prediction was confirmed through other laboratory tests.
Wagenaar, G. T. M. et al. Gene expression profile and histopathology of experimental bronchopulmonary dysplasia induced by prolonged oxidative stress. Free Radic. Biol. Med. 36, 782–801 (2004).
Lu, T. et al. Application of cDNA microarray to the study of arsenic-induced liver diseases in the population of Guizhou, China. Toxicol. Sci. 59, 185–192 (2001).
Hamadeh, H. K. et al. Integration of clinical and gene expression endpoints to explore furan-mediated hepatotoxicity. Mutat. Res. 549, 169–183 (2004).
Iida, M. et al. Changes in global gene and protein expression during early mouse liver carcinogenesis induced by non-genotoxic model carcinogens oxazepam and Wyeth-14,643. Carcinogenesis 24, 757–770 (2003).
Ellinger-Ziegelbauer, H., Stuart, B., Wahle, B., Bomann, W. & Ahr, H. J. Characteristic expression profiles induced by genotoxic carcinogens in rat liver. Toxicol. Sci. 77, 19–34 (2004).
Twigger, S. et al. Rat Genome Database (RGD): mapping disease onto the genome. Nucleic Acids Res. 30, 125–128 (2002).
Higgins, M. A. et al. Gene expression analysis of the acute phase response using a canine microarray. Toxicol. Sci. 74, 470–484 (2003). The authors describe the development of the first canine array. It was used to characterize the acute phase response to inflammation that is seen in parallel with other phenotypic markers in the beagle, an important model organism in late stage, pharmaceutical toxicity testing.
Ulrich, R. G., Rockett, J. C., Gibson, G. G. & Pettit, S. D. Overview of an interlaboratory collaboration on evaluating the effects of model hepatotoxicants on hepatic gene expression. Environ. Health Perspect. 112, 423–427 (2004).
Pennie, W., Pettit, S. D. & Lord, P. G. Toxicogenomics in risk assessment: an overview of an HESI collaborative research program. Environ. Health Perspect. 112, 417–419 (2004). This provides an overview of the toxicogenomics consortium supported by the Health and Environmental Sciences Institute: the ILSI Committee on the Application of Toxicogenomics to Risk Assessment.
Lindon, J. C. et al. Contemporary issues in toxicology the role of metabonomics in toxicology and its evaluation by the COMET project. Toxicol. Appl. Pharmacol. 187, 137–146 (2003). This paper led COMET to apply metabonomics studies to toxicogenomics
Nuwaysir, E. F., Bittner, M., Trent, J., Barrett, J. C. & Afshari, C. A. Microarrays and toxicology: the advent of toxicogenomics. Mol. Carcinog. 24, 153–159 (1999).
Heller, R. A. et al. Discovery and analysis of inflammatory disease-related genes using cDNA microarrays. Proc. Natl Acad. Sci. USA 94, 2150–2155 (1997).
Loeb, W. F. & Quimby, F. W. (eds) The Clinical Chemistry of Laboratory Animals (Taylor and Francis, 1999).
Travlos, G. S. et al. Frequency and relationships of clinical chemistry and liver and kidney histopathology findings in 13-week toxicity studies in rats. Toxicology 107, 17–29 (1996).
Paules, R. Phenotypic anchoring: linking cause and effect. Environ. Health Perspect. 111, A338–A339 (2003). The author describes the use of the term 'phenotypic anchoring', first mentioned by Tennant in reference 7 to describe the process of using a pathology finding or toxicological endpoint to provide context for expression changes.
Searfoss, G. H. et al. Adipsin, a biomarker of gastrointestinal toxicity mediated by a functional γ-secretase inhibitor. J. Biol. Chem. 278, 46107–46116 (2003). For the first time, the authors apply the process of identifying a molecular marker of toxicity to support pre-clinical studies, and define the mechanism of toxicity associated with a novel class of toxicant.
Newton, R. K., Aardema, M. & Aubrecht, J. The utility of DNA microarrays for characterizing genotoxicity. Environ. Health Perspect. 112, 420–422 (2004).
Petricoin, E. F. 3rd et al. Serum proteomic patterns for detection of prostate cancer. J. Natl Cancer Inst. 94, 1576–1578 (2002).
Thompson, K. L. et al. Identification of platform-independent gene expression markers of cisplatin nephrotoxicity. Environ. Health Perspect. 112, 488–494 (2004).
Kramer, J. A. et al. Overview on the application of transcription profiling using selected nephrotoxicants for toxicology assessment. Environ. Health Perspect. 112, 460–464 (2004).
Baker, V. A. et al. Clofibrate-induced gene expression changes in rat liver: a cross-laboratory analysis using membrane cDNA arrays. Environ. Health Perspect. 112, 428–438 (2004).
Chu, T. M., Deng, S., Wolfinger, R., Paules, R. S. & Hamadeh, H. K. Cross-site comparison of gene expression data reveals high similarity. Environ. Health Perspect. 112, 449–455 (2004).
Waring, J. F. et al. Interlaboratory evaluation of rat hepatic gene expression changes induced by methapyrilene. Environ. Health Perspect. 112, 439–448 (2004).
Smith, L. L. Key challenges for toxicologists in the 21st century. Trends Pharmacol. Sci. 22, 281–285 (2001).
Hogstrand, C., Balesaria, S. & Glover, C. N. Application of genomics and proteomics for study of the integrated response to zinc exposure in a non-model fish species, the rainbow trout. Comp. Biochem. Physiol. B 133, 523–535 (2002).
Juan, H. -F. et al. Biomic study of human meyloid leukemia cells differentiation to macrophages using DNA array, proteomic, and bioinformatic analytical methods. Electrophoresis 23, 2490–2504 (2002).
Amin, R. P. et al. Genomic interrogation of mechanism(s) underlying cellular responses to toxicants. Toxicology 181–182, 555–563 (2002).
Kita, Y. et al. Implications of circadian gene expression in kidney, liver and the effects of fasting on pharmacogenomic studies. Pharmacogenetics 12, 55–65 (2002).
Kaput, J. & Rodriguez, R. L. Nutritional genomics: the next frontier in the postgenomic era. Physiol. Genomics 16, 166–177 (2004).
Kaput, J. Diet–disease gene interactions. Nutrition 20, 26–31 (2004).
Klein, T. E. and Altman, R. B. PharmGKB: the pharmacogenetics nad pharmacogenomics knowledge base. Pharmacogenomics J. 4, 1 (2004).
Wittliff, J. L. & Erlander, M. G. Laser capture microdissection and its applications in genomics and proteomics. Methods Enzymol. 356, 12–25 (2002).
Jain, K. K. Application of laser capture microdissection to proteomics. Methods Enzymol. 356, 157–167 (2002).
Emmert-Buck, M. R. et al. Laser capture microdissection. Science 274, 998–1001 (1996).
Bonner, R. F. et al. Laser capture microdissection: molecular analysis of tissue. Science 278, 1481–1483 (1997).
Karsten, S. L., Van Deerlin, V. M., Sabatti, C., Gill, L. H. & Geschwind, D. H. An evaluation of tyramide signal amplification and archived fixed and frozen tissue in microarray gene expression analysis. Nucleic Acids Res. 30, E4 (2002).
Hamadeh, H. K. et al. Detection of diluted gene expression alterations using cDNA microarrays. Biotechniques 32, 322, 324, 326–329 (2002).
Huang, R. P. Detection of multiple proteins in an antibody-based protein microarray system. J. Immunol. Methods 255, 1–13 (2001).
Merchant, M. & Weinberger, S. R. Recent advancements in surface-enhanced laser desorption/ionization-time of flight-mass spectrometry. Electrophoresis 21, 1164–1177 (2000).
Liotta, L. & Petricoin, E. Molecular profiling of human cancer. Nature Rev. Genet. 1, 48–56 (2000).
Quackenbush, J. Computational analysis of microarray data. Nature Rev. Genet. 2, 418–427 (2001).
Farland, W. H. The U. S. Environmental Protection Agency's Risk Assessment Guidelines: current status and future directions. Toxicol. Ind. Health 8, 205–212 (1992).
Farland, W. H. Cancer risk assessment: evolution of the process. Prev. Med. 25, 24–25 (1996).
Larsen, J. C., Farland, W. & Winters, D. Current risk assessment approaches in different countries. Food Addit. Contam. 17, 359–369 (2000).
Begley, T. J., Rosenbach, A. S., Ideker, T. & Samson, L. D. Damage recovery pathways in Saccharomyces cerevisiae revealed by genomic phenotyping and interactome mapping. Mol. Cancer Res. 1, 103–112 (2002).
Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Lewis, F., Maughan, N. J., Smith, V., Hillan, K. & Quirke, P. Unlocking the archive — gene expression in paraffin-embedded tissue. J. Pathol. 195, 66–71 (2001).
Wetmore, B. A. & Merrick, B. A. Toxicoproteomics: proteomics applied to toxicology and pathology. Toxicol. Pathol. 32, 619–642 (2004).
Stuart, G. W. & Berry, M. W. A comprehensive whole genome bacterial phylogeny using correlated peptide motifs defined in a high dimensional vector space. J. Bioinform. Comput. Biol. 1, 475–493 (2003).
Chaussabel, D. & Sher, A. Mining microarray expression data by literature profiling. Genome Biol. 3, research0055.1–research0055.16 (2002).
Sluka, J. P. in Methods of Microarray Data Analysis II (eds Lin, S. M. & Johnson, K. F.) (Kluwer Academic Publishers, Boston, 2002).
Patterson, S. D. & Aebersold, R. H. Proteomics: the first decade and beyond. Nature Genet. 33, S311–S323 (2003).
Nicholson, J. K., Connelly, J., Lindon, J. C. & Holmes, E. Metabonomics: a platform for studying drug toxicity and gene function. Nature Rev. Drug Discov. 1, 153–161 (2002).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Mattingly, C. J., Colby, G. T., Rosenstein, M. C., Forrest, J. N. Jr & Boyer, J. L. Promoting comparative molecular studies in environmental health research: an overview of the comparative toxicogenomics database (CTD). Pharmacogenomics J. 4, 5–8 (2004). The authors proposed and developed a public database to house gene sequences and toxicology information in support of comparative toxicogenomics across species, and to inform human risk assessment.
Brazma, A. et al. ArrayExpress — a public repository for microarray gene expression data at the EBI. Nucleic Acids Res. 31, 68–71 (2003).
Ball, C. A. et al. Standards for microarray data. Science 298, 539 (2002).
Brazma, A. et al. Minimum information about a microarray experiment (MIAME) — toward standards for microarray data. Nature Genet. 29, 365–371 (2001).
Ikeo, K., Ishi-i, J., Tamura, T., Gojobori, T. & Tateno, Y. CIBEX: center for information biology gene expression database. C. R. Biol. 326, 1079–1082 (2003).
Orchard, S. et al. Common interchange standards for proteomics data: public availability of tools and schema. Proteomics 4, 490–491 (2004).
Bartosiewicz, M., Trounstine, M., Barker, D., Johnston, R. & Buckpitt, A. Development of a toxicological gene array and quantitative assessment of this technology. Arch. Biochem. Biophys. 376, 66–73 (2000).
Jiang, X. S. et al. A high-throughput approach for subcellular proteome: identification of rat liver proteins using subcellular fractionation coupled with two-dimensional liquid chromatography tandem mass spectrometry and bioinformatic analysis. Mol. Cell. Proteomics 3, 441–455 (2004).
Vitorino, R. et al. Identification of human whole saliva protein components using proteomics. Proteomics 4, 1109–1115 (2004).
Talamo, F. et al. Proteins from bovine tissues and biological fluids: defining a reference electrophoresis map for liver, kidney, muscle, plasma and red blood cells. Proteomics 3, 440–460 (2003).
Reddy, P. H. et al. Gene expression profiles of transcripts in amyloid precursor protein transgenic mice: up-regulation of mitochondrial metabolism and apoptotic genes is an early cellular change in Alzheimer's disease. Hum. Mol. Genet. (2004).
Heijne, W. H., Stierum, R. H., Slijper, M., van Bladeren, P. J. & van Ommen, B. Toxicogenomics of bromobenzene hepatotoxicity: a combined transcriptomics and proteomics approach. Biochem. Pharmacol. 65, 857–875 (2003).
Weiss, A., Delproposto, J. & Giroux, C. N. High-throughput phenotypic profiling of gene–environment interactions by quantitative growth curve analysis in Saccharomyces cerevisiae. Anal. Biochem. 327, 23–34 (2004).
Fountoulakis, M. et al. Two-dimensional database of mouse liver proteins: changes in hepatic protein levels following treatment with acetaminophen or its nontoxic regioisomer 3-acetamidophenol. Electrophoresis 21, 2148–2161 (2000).
Lee, W. M. Drug-induced hepatotoxicity. N. Engl. J. Med. 349, 474–485 (2003).
Xie, T. et al. Changes in gene expression linked to methamphetamine-induced dopaminergic neurotoxicity. J. Neurosci. 22, 274–283 (2002).
Dam, K., Seidler, F. J. & Slotkin, T. A. Transcriptional biomarkers distinguish between vulnerable periods for developmental neurotoxicity of chlorpyrifos: implications for toxicogenomics. Brain Res. Bull. 59, 261–265 (2003).
Hamadeh, H. K., Trouba, K. J., Amin, R. P., Afshari, C. A. & Germolec, D. Coordination of altered DNA repair and damage pathways in arsenite-exposed keratinocytes. Toxicol. Sci. 69, 306–316 (2002).
Hu, D., Cao, K., Peterson-Wakeman, R. & Wang, R. Altered profile of gene expression in rat hearts induced by chronic nicotine consumption. Biochem. Biophys. Res. Commun. 297, 729–736 (2002).
Waring, J. F., Ciurlionis, R., Jolly, R. A., Heindel, M. & Ulrich, R. G. Microarray analysis of hepatotoxins in vitro reveals a correlation between gene expression profiles and mechanisms of toxicity. Toxicol. Lett. 120, 359–368 (2001).
Hamadeh, H. K., Bushel, P., Paules, R. & Afshari, C. A. Discovery in toxicology: mediation by gene expression array technology. J. Biochem. Mol. Toxicol. 15, 231–242 (2001).
Witzmann, F. A. et al. Analysis of rat testicular protein expression following 91-day exposure to JP-8 jet fuel vapor. Proteomics 3, 1016–1027 (2003).
Huang, Q. et al. Gene expression profiling reveals multiple toxicity endpoints induced by hepatotoxicants. Mutat. Res. 549, 147–167 (2004).
Adachi, T. et al. Toxicogenomic difference between diethylstilbestrol and 17β-estradiol in mouse testicular gene expression by neonatal exposure. Mol. Reprod. Dev. 67, 19–25 (2004).
Nadadur, S. S., Schladweiler, M. C. & Kodavanti, U. P. A pulmonary rat gene array for screening altered expression profiles in air pollutant-induced lung injury. Inhal. Toxicol. 12, 1239–1254 (2000).
Akhtar, R. A. et al. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr. Biol. 12, 540–550 (2002).
Heijne, W. H. et al. Bromobenzene–induced hepatotoxicity at the transcriptome level. Toxicol. Sci. 79, 411–422 (2004).
Bessems, J. G. & Vermeulen, N. P. Paracetamol (acetaminophen)-induced toxicity: molecular and biochemical mechanisms, analogues and protective approaches. Crit. Rev. Toxicol. 31, 55–138 (2001).
Reilly, T. P. et al. Expression profiling of acetaminophen liver toxicity in mice using microarray technology. Biochem. Biophys. Res. Commun. 282, 321–328 (2001).
de Longueville, F. et al. Use of a low-density microarray for studying gene expression patterns induced by hepatotoxicants on primary cultures of rat hepatocytes. Toxicol. Sci. 75, 378–392 (2003).
Xirasagar, S. et al. CEBS object model for systems biology data, SysBio–OM. Bioinformatics 20, 2004–2015 (2004).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Petricoin, E. F. 3rd et al. Medical applications of microarray technologies: a regulatory science perspective. Nature Genet, 32, S474–S479 (2002).
Hermjakob, H. et al. The HUPO PSI's molecular interaction format — a community standard for the representation of protein interaction data. Nature Biotechnol. 22, 177–183 (2004).
Jones, A., Hunt, E., Wastling, J., Pizarro, A. & Stoeckert, C. J. Jr. An object model and database for functional genomics. Bioinformatics (2004).
Acknowledgements
We are indebted to B. Alex Merrick, Richard S. Paules and Raymond W. Tennant for their consultation and assistance with this manuscript, and to Kenneth Olden, Samuel Wilson, Lutz Birnbaumer and the staff of the National Center for Toxicogenomics for their continuing support and involvement with this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
OMIM
FURTHER INFORMATION
Chemical Effects in Biological Systems
Clinical Data Interchange Standards Consortium
EMBL-European Bioinformatics Institute
'From OMICS to systems biology' poster
ILSI Health and Environmental Services Institute
MGED Reporting Structure for Biological Investigations (RSBI)
Glossary
- NECROSIS
-
The localized death of living cells.
- MODE-OF-ACTION
-
The sequence of events from the absorption of a compound into an organism to a toxic outcome or death.
- PROTEIN CHIP
-
A genomic set of proteins that are arrayed on a solid surface without denaturation.
- NUCLEAR MAGNETIC RESONANCE
-
An analytical chemistry technique that is used to study molecular structure and dynamics; it explores spectral differences that are caused by the differential alignment of atomic spins in the presence of a strong magnetic field.
- TRANSCRIPTOMICS
-
Techniques that measure the full complement of activated genes, mRNAs or transcripts in a particular tissue at a particular time, typically through the use of cDNA or oligonucleotide microarrays.
- PROTEOMICS
-
A collection of techniques used to measure the structural and functional properties of proteins through the use of 2-dimensional gel electrophoresis or liquid chromatography; typically followed by protein identification using some form of mass spectrometry.
- METABONOMICS
-
Techniques that detect changes in the concentration of low-molecular-weight metabolites present in a cell or organism at a given time (the metabonome) by using nuclear magnetic resonance or mass spectrometry coupled to gas or liquid chromatography.
- KNOWLEDGEBASE
-
An archival and computational system that uses data, information and knowledge captured from experts to carry out tasks that create new information and new understanding.
- SYSTEMS TOXICOLOGY
-
The study of the perturbation of biological systems by chemicals and stressors, monitoring changes in molecular expression and conventional toxicological parameters, and iteratively integrating response data to describe the functioning organism.
- SYSTEMS BIOLOGY
-
The integrated study of biological systems (cells, tissues, organs or entire organisms) at the molecular level. It involves perturbing systems, monitoring molecular expression, integrating response data and modelling the molecular structure and network function of the system.
- BIOMARKER
-
A pharmacological or physiological measurement that is used to predict a toxic event in an animal.
- TOXICOINFORMATICS
-
The description of a toxicological stress and the annotation of the dose-dependent molecular responses that are elicited over time.
- INFORMATION SCIENCE
-
The systematic study and analysis of the sources, development, collection, organization, dissemination, evaluation, use and management of information in all its forms, including the media (formal and informal) and technology used in its communication.
- METABOLOMICS
-
The directed use of quantitative analytical methods for analysing the entire metabolic content of a cell or organism at a given time (the metabolome).
- FUNCTIONAL GENOMICS
-
The development and application of global (genome-wide or system-wide) experimental approaches to assess gene function by making use of information and reagents provided by physical mapping and sequencing of genomes.
- TANDEM MASS SPECTROMETRY
-
The use of two mass spectrometers in series to detect and identify substances on the basis of mass and charge.
- SYNOVIOCYTES
-
Cells believed to be responsible for the production of synovial-fluid components in joints, for absorption from the joint cavity, and for blood/synovial fluid exchanges.
- CHONDROCYTES
-
Cartilage cells that produce the structural components of cartilage.
- LEAD COMPOUNDS
-
Chemicals or drugs that show promise for commercialization.
- LONGITUDINAL DATA MINING
-
The process of locating previously unknown patterns and relationships within data that result from multiple observations of a population of genes, animals or patients.
- PRINCIPAL-COMPONENT ANALYSIS
-
A statistical method that seeks to reduce the dimensionality of a data set by projecting the data onto new axes that align with the variability in the data.
- NUTRIGENOMICS
-
The study of the nutritional environment and related cellular or genetic processes at the level of the genome.
- PHYSIOLOGICALLY-BASED PHARMACOKINETIC MODELLING
-
Involves deriving a set of mathematical (differential) equations that are structured to provide a time course for a chemical's mass–balance disposition (wherein all inputs, outputs and changes in total mass of the chemical are accounted for) in preselected anatomical compartments.
- PHARMACODYNAMIC MODELLING
-
Involves the development of a mathematical description of a toxicological or disease outcome after therapy.
- TARGET TISSUE
-
The tissue or tissues that are damaged as a result of exposure to a toxicant or stressor.
- REAL-TIME PCR
-
A process that allows the amount of PCR product to be quantified during each cycle of a PCR reaction. The product concentration, as a function of cycle number, provides a good estimation of the relative quantity of the mRNA being tested.
- PARSING
-
The process of determining the syntactic structure of a sentence or string of symbols in a language.
- RNA INTERFERENCE
-
An ancient natural antiviral mechanism that directs silencing of gene expression in a sequence-specific manner and can be exploited artificially to inhibit the expression of any gene of interest.
- BIOLOGICALLY-BASED DOSE-RESPONSE MODELLING
-
The science of establishing dose-response models based on underlying biological processes.
Rights and permissions
About this article
Cite this article
Waters, M., Fostel, J. Toxicogenomics and systems toxicology: aims and prospects. Nat Rev Genet 5, 936–948 (2004). https://doi.org/10.1038/nrg1493
Issue Date:
DOI: https://doi.org/10.1038/nrg1493
This article is cited by
-
Astrocyte-Like Cells Transcriptome Changes After Exposure to a Low and Non-cytotoxic MeHg Concentration
Biological Trace Element Research (2023)
-
GWAS and Beyond: Using Omics Approaches to Interpret SNP Associations
Current Genetic Medicine Reports (2019)
-
JAK/STAT and TGF-ß activation as potential adverse outcome pathway of TiO2NPs phototoxicity in Caenorhabditis elegans
Scientific Reports (2017)
-
Insights into the molecular mechanisms of Polygonum multiflorum Thunb-induced liver injury: a computational systems toxicology approach
Acta Pharmacologica Sinica (2017)
-
Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions
Nature Communications (2017)