Abstract
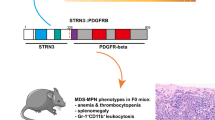
Various histological subtypes of leukaemia and lymphoma are associated with diagnostic chromosome translations1–4, and substantial strides have been made in determining the specific oncogenes targetted by those translocations. We report the cloning of a novel fusion oncogene associated with a unique leukaemia/lymphoma syndrome. Patients afflicted with this syndrome present with lymphoblastic lymphoma and a myelopro I iterative disorder, often accompanied by pronounced peripheral eosinophilia and/or prominent eosinophilic infiltrates in the affected bone marrow, which generally progress to full-blown acute myelogenous leukaemia within a year of diagnosis5–9. A specific chromosome translocation, t(8;13)(p11;q11–12), is found in both lymphoma and myeloid leukaemia cells from these patients, supporting bi-lineage differentiation from a transformed stem cell6–8. We find that the 8p11 translocation breakpoints, in each of four patients, interrupt intron 8 of the fibroblast growth factor receptor 1 gene (FGFR1). These translocations are associated with aberrant transcripts in which four predicted zinc-finger domains, contributed by a novel and widely expressed chromosome-13 gene (ZNF198), are fused to the FGFR1 tyrosine-kinase domain. Transient expression studies show that the ZNF198–FGFR1 fusion transcript directs the synthesis of an approximately 87-kD polypeptide, localizing predominantly to the cytoplasm. Our studies demonstrate an FGFR1 oncogenic role and suggest a tumorigenic mechanism in which ZNF198–FGFR1 activation results from ZNF198 zinc-finger-mediated homodimerization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Rowley, J.D. Molecular cytogenetics: Rosetta stone for understanding cancer: twenty-ninth G.H.A. Clowes memorial award lecture. Cancer Res. 50, 3816–3825 (1990).
Rabbitts, T.H. Chromosomal translocations in human cancer. Nature 372, 143–149 (1994).
Offit, K. & Chaganti, R.S. Chromosomal aberrations in non-Hodgkin′s lymphoma: biologic and clinical correlations. Hematol. Oncol. Clin. North Am. 5, 853–869 (1991).
Croce, CM. Molecular biology of lymphomas. Semin. Oncol. 20, 31–46 (1993).
Abruzzo, L.V. et al. T-cell lymphoblastic lymphoma with eosinophilia associated with subsequent myeloid malignancy. Am. J. Surg. Pathol. 16, 236–245 (1992).
Inhorn, R.C. et al. A syndrome of lymphoblastic lymphoma, eosinophilia, and myeloid hyperplasia/malignancy associated with t(8;13)(p11;q11): description of a distinctive clinicopathologic entity. Blood 85, 1881–1887 (1995).
Pagan, K., Hyde, S. & Harrison, P. Translocation (8; 13) and T-cell lymphoma: a case report. Cancer Genet. Cytogenet. 65, 71–73 (1993).
Naeem, R., Singer, S. & Fletcher, J.A. Translocation t(8;13)(p11;q11-12) in stem cell leukemia/lymphoma of T-cell and myeloid lineages. Genes Chromosom. Cancer 12, 148–151 (1995).
Macdonald, D., Aguiar, R.C, Mason, P.J., Goldman, J.M. & Cross, N.C. A new myeloproliferative disorder associated with chromosomal translocations involving 8p11: a review. Leukemia 9, 1628–1630 (1995).
Dib, A. et al. Characterization of the region of the short arm of chromosome 8 amplified in breast carcinoma. Oncogene 10, 995–1001 (1995).
Itoh, N. Terachi, T., Ohta, M. & Seo, M.K. The complete amino acid sequence of the shorter form of human basic fibroblast growth factor receptor deduced from itscDNA. Biochem. Biophys, Res. Commun. 169, 680–685 (1990).
van der Maarel, S.M. et al. Cloning and characterization of DXS6673E, a candidate gene for X-linked mental retardation in Xq 13.1. Hum. Mol. Genet. 5, 887–897 (1996).
Verdier, A.S. et al. Chromosomal mapping of two novel human FGF genes, FGF11 and FGF12. Genomics 40, 151–154 (1996).
Burgess, W.H. & Maciag, T. The heparin-binding (fibroblast) growth factor family of proteins. Annu. Rev. Biochem. 58, 575–606 (1989).
Gabbianelli, M. et al. “Pure” human hematopoietic progenitors: permissive action of basic fibroblast growth factor. Science 249, 1561–1564 (1990).
Gabrilove, J.L., White, K., Rahman, Z. & Wilson, E.L. Stem cell factor and basic fibroblast growth factor are synergistic in augmenting committed myeloid progenitor cell growth. Blood 83, 907–910 (1994).
Wilson, E.L., Rifkin, D.B., Kelly, F., Hannocks, MJ. & Gabrilove, J.L. Basic fibroblast growth factor stimulates myelopoiesis in long-term human bone marrow cultures. Blood 77, 954–960 (1991).
Avraham, H., Banu, N., Scadden, D.T., Abraham, J. & Groopman, J.E. Modulation of megakaryocytopoiesis by human basic fibroblast growth factor. Blood 83, 2126–2132 (1994
Kouhara, H. et al. A lipid-anchored Grb2-binding protein that links FGF-receptor activation to the Ras/MAPK signaling pathway. Cell 89, 693–702 (1997
Carroll, M., Tomasson, M.H., Barker, G.F., Golub, T.R. & Gilliland, D.G. The TEL/platelet-derived growth factor b receptor (PDGFbR) fusion in chronic myelomonocytic leukemia is a transforming protein that self-associates and activates PDGF(3R kinase-dependent signaling pathways. Proc. Natl. Acad. Sci. USA 93, 14845–14850 (1996).
Rodrigues, G.A. & Park, M. Dimerization mediated through a leucine zipper activates the oncogenic potential of the met receptor tyrosine kinase. Mol. Cell Biol. 13, 6711–6722 (1993).
Santoro, M. et al. Activation of RET as a dominant transforming gene by germline mutations of MEN2A and MEN2B. Science 267, 381–383 (1995).
Tong, Q., Xing, S. & Jhiang, S.M. Leucine zipper-mediated dimerization is essential for the PTC1 oncogenic activity. J. Biol. Chem. 272, 9043–9047 (1997).
Luisi, B.F . et al. Crystallographic analysis of the interaction of the glucocorticoid receptor with DNA. Nature 352, 497–505 (1991).
Naski, M.C., Wang, Q., Xu, J. & Ornitz, D.M. Graded activation of fibroblast growth factor receptor 3 by mutations causing achondroplasia and thanatophoric dysplasia. Nature Genet. 13, 233–237 (1996).
Fletcher, J.A. et al. Diagnostic relevance of clonal cytogenetic aberrations in malignant soft-tissue tumors. N. Engl. J. Med. 324, 436–442 (1991.).
Xiao, S., Renshaw, A.A., Cibas, E.S.,Hudson, T.J. & Fletcher, J.A. Novel fluorescence in situ hybridization approaches in solid tumors: characterization of frozen specimens, touch preparations, and cytologic preparations. Am. J. Pathol. 147, 896–904 (1995).
Parimoo, S., Kolluri, R. & Weissman, S.M. cDNA selection from total yeast DNA containing YACs. Nucleic Acids Res. 21, 4422–4423 (1993
Pear, W.S., Nolan, G.P., Scott, M.L & Baltimore, D. Production of high-titer helper-free retroviruses by transient transfection. Proc. Natl. Acad. Sci. USA 90, 8392–8396 (1993).
Aster, J.C. et al. Oncogenic forms of NOTCH1 lacking either the primary binding site for RBP-J or nuclear localization sequences retain the ability to associate with RBP-J and activate transcription. J. Biol. Chem. 272, 11336–11343 (1997).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Xiao, S., Nalabolu, S., Aster, J. et al. FGFR1 is fused with a novel zinc-finger gene, ZNF198, in the t(8;13) leukaemia/lymphoma syndrome. Nat Genet 18, 84–87 (1998). https://doi.org/10.1038/ng0198-84
Issue Date:
DOI: https://doi.org/10.1038/ng0198-84
This article is cited by
-
Identification of a novel HOOK3-FGFR1 fusion gene involved in activation of the NF-kappaB pathway
Cancer Cell International (2022)
-
A proximity biotinylation-based approach to identify protein-E3 ligase interactions induced by PROTACs and molecular glues
Nature Communications (2022)
-
Fibroblast growth factor receptor fusions in cancer: opportunities and challenges
Journal of Experimental & Clinical Cancer Research (2021)
-
Serum fibroblast growth factor 19 serves as a potential novel biomarker for hepatocellular carcinoma
BMC Cancer (2019)
-
Detection of novel fusion-transcripts by RNA-Seq in T-cell lymphoblastic lymphoma
Scientific Reports (2019)