Abstract
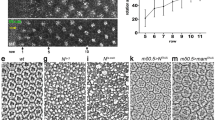
Polycomb Group (PcG) proteins silence critical developmental genes and modulate cell proliferation. Using the Drosophila melanogaster eye as a model system, we show that cells with mutations in the gene locus (ph) that encodes the PcG protein Polyhomeotic (PH) overproliferate and lose both the ability to differentiate and their normal polarity. They invade the neighboring tissues and, when combined with an activated form of the Ras proto-oncogene, they trigger the formation of metastases. PcG proteins bind to multiple genes in the Notch pathway and control their transcription as well as Notch signaling. The massive cell-autonomous overproliferation of ph mutant cell clones can be rescued by ectopic expression of a dominant negative form of Notch or by RNA interference (RNAi)-mediated repression of Notch. Conversely, overexpression of ph induces a small-eye phenotype that is rescued by activation of Notch signaling. These data show that ph is a tumor suppressor locus that controls cellular proliferation by silencing multiple Notch signaling components.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. & Cavalli, G. Genome regulation by polycomb and trithorax proteins. Cell 128, 735–745 (2007).
Sparmann, A. & van Lohuizen, M. Polycomb silencers control cell fate, development and cancer. Nat. Rev. Cancer 6, 846–856 (2006).
Bracken, A.P. et al. The Polycomb group proteins bind throughout the INK4A-ARF locus and are disassociated in senescent cells. Genes Dev. 21, 525–530 (2007).
Jacobs, J.J., Kieboom, K., Marino, S., DePinho, R.A. & van Lohuizen, M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 397, 164–168 (1999).
Bracken, A.P., Dietrich, N., Pasini, D., Hansen, K.H. & Helin, K. Genome-wide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 20, 1123–1136 (2006).
Ohm, J.E. et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat. Genet. 39, 237–242 (2007).
Widschwendter, M. et al. Epigenetic stem cell signature in cancer. Nat. Genet. 39, 157–158 (2007).
Bruggeman, S.W. et al. Bmi1 controls tumor development in an ink4a/arf-independent manner in a mouse model for glioma. Cancer Cell 12, 328–341 (2007).
Beuchle, D., Struhl, G. & Muller, J. Polycomb group proteins and heritable silencing of Drosophila Hox genes. Development 128, 993–1004 (2001).
Martinez, A.M., Colomb, S., Dejardin, J., Bantignies, F. & Cavalli, G. Polycomb group-dependent Cyclin A repression in Drosophila. Genes Dev. 20, 501–513 (2006).
Oktaba, K. et al. Dynamic regulation by polycomb group protein complexes controls pattern formation and the cell cycle in Drosophila. Dev. Cell 15, 877–889 (2008).
Brumby, A.M. & Richardson, H.E. scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila. EMBO J. 22, 5769–5779 (2003).
Pagliarini, R.A. & Xu, T. A genetic screen in Drosophila for metastatic behavior. Science 302, 1227–1231 (2003).
Shao, Z. et al. Stabilization of chromatin structure by PRC1, a Polycomb complex. Cell 98, 37–46 (1999).
Vaccari, T. & Bilder, D. The Drosophila tumor suppressor vps25 prevents nonautonomous overproliferation by regulating notch trafficking. Dev. Cell 9, 687–698 (2005).
Bryant, P.J. & Levinson, P. Intrinsic growth control in the imaginal primordia of Drosophila, and the autonomous action of a lethal mutation causing overgrowth. Dev. Biol. 107, 355–363 (1985).
Klebes, A. & Knust, E. A conserved motif in Crumbs is required for E-cadherin localisation and zonula adherens formation in Drosophila. Curr. Biol. 10, 76–85 (2000).
Pagliarini, R.A., Quinones, A.T. & Xu, T. Analyzing the function of tumor suppressor genes using a Drosophila model. Methods Mol. Biol. 223, 349–382 (2003).
Gateff, E. Malignant neoplasms of genetic origin in Drosophila melanogaster. Science 200, 1448–1459 (1978).
Gonzalez, C. Spindle orientation, asymmetric division and tumour suppression in Drosophila stem cells. Nat. Rev. Genet. 8, 462–472 (2007).
Nègre, N. et al. Chromosomal distribution of PcG proteins during Drosophila development. PLoS Biol. 4, e170 (2006).
Schuettengruber, B. et al. Functional anatomy of polycomb and trithorax chromatin landscapes in Drosophila embryos. PLoS Biol. 7, e13 (2009).
Schwartz, Y.B. et al. Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat. Genet. 38, 700–705 (2006).
Tolhuis, B. et al. Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster. Nat. Genet. 38, 694–699 (2006).
Randsholt, N.B., Maschat, F. & Santamaria, P. polyhomeotic controls engrailed expression and the hedgehog signaling pathway in imaginal discs. Mech. Dev. 95, 89–99 (2000).
Domínguez, M. & de Celis, J.F. A dorsal/ventral boundary established by Notch controls growth and polarity in the Drosophila eye. Nature 396, 276–278 (1998).
Furriols, M. & Bray, S. A model Notch response element detects Suppressor of Hairless-dependent molecular switch. Curr. Biol. 11, 60–64 (2001).
Hariharan, I.K. & Bilder, D. Regulation of imaginal disc growth by tumor-suppressor genes in Drosophila. Annu. Rev. Genet. 40, 335–361 (2006).
Moberg, K.H., Bell, D.W., Wahrer, D.C., Haber, D.A. & Hariharan, I.K. Archipelago regulates Cyclin E levels in Drosophila and is mutated in human cancer cell lines. Nature 413, 311–316 (2001).
Dow, L.E. et al. The tumour-suppressor Scribble dictates cell polarity during directed epithelial migration: regulation of Rho GTPase recruitment to the leading edge. Oncogene 26, 2272–2282 (2007).
Wodarz, A. & Nathke, I. Cell polarity in development and cancer. Nat. Cell Biol. 9, 1016–1024 (2007).
Reynolds-Kenneally, J. & Mlodzik, M. Notch signaling controls proliferation through cell-autonomous and non-autonomous mechanisms in the Drosophila eye. Dev. Biol. 285, 38–48 (2005).
Dominguez, M., Ferres-Marco, D., Gutierrez-Avino, F.J., Speicher, S.A. & Beneyto, M. Growth and specification of the eye are controlled independently by Eyegone and Eyeless in Drosophila melanogaster. Nat. Genet. 36, 31–39 (2004).
Martinez, A.M. & Cavalli, G. The role of polycomb group proteins in cell cycle regulation during development. Cell Cycle 5, 1189–1197 (2006).
Ferres-Marco, D. et al. Epigenetic silencers and Notch collaborate to promote malignant tumours by Rb silencing. Nature 439, 430–436 (2006).
Hurlbut, G.D., Kankel, M.W., Lake, R.J. & Artavanis-Tsakonas, S. Crossing paths with Notch in the hyper-network. Curr. Opin. Cell Biol. 19, 166–175 (2007).
Bolós, V., Grego-Bessa, J. & de la Pompa, J.L. Notch signaling in development and cancer. Endocr. Rev. 28, 339–363 (2007).
Lee, N., Maurange, C., Ringrose, L. & Paro, R. Suppression of Polycomb group proteins by JNK signalling induces transdetermination in Drosophila imaginal discs. Nature 438, 234–237 (2005).
Wang, J., Lee, C.H., Lin, S. & Lee, T. Steroid hormone-dependent transformation of polyhomeotic mutant neurons in the Drosophila brain. Development 133, 1231–1240 (2006).
Bello, B., Holbro, N. & Reichert, H. Polycomb group genes are required for neural stem cell survival in postembryonic neurogenesis of Drosophila. Development 134, 1091–1099 (2007).
Lee, T.I. et al. Control of developmental regulators by polycomb in human embryonic stem cells. Cell 125, 301–313 (2006).
Deshpande, A.M. et al. PHC3, a component of the hPRC-H complex, associates with E2F6 during G0 and is lost in osteosarcoma tumors. Oncogene 26, 1714–1722 (2007).
Tokimasa, S. et al. Lack of the Polycomb-group gene rae28 causes maturation arrest at the early B-cell developmental stage. Exp. Hematol. 29, 93–103 (2001).
Dura, J.M. et al. A complex genetic locus, polyhomeotic, is required for segmental specification and epidermal development in D. melanogaster. Cell 51, 829–839 (1987).
Rebay, I., Fehon, R.G. & Artavanis-Tsakonas, S. Specific truncations of Drosophila Notch define dominant activated and dominant negative forms of the receptor. Cell 74, 319–329 (1993).
Haerry, T.E., Khalsa, O., O'Connor, M.B. & Wharton, K.A. Synergistic signaling by two BMP ligands through the SAX and TKV receptors controls wing growth and patterning in Drosophila. Development 125, 3977–3987 (1998).
Lee, J.D. et al. An acylatable residue of Hedgehog is differentially required in Drosophila and mouse limb development. Dev. Biol. 233, 122–136 (2001).
Heitzler, P. & Simpson, P. The choice of cell fate in the epidermis of Drosophila. Cell 64, 1083–1092 (1991).
Hadorn, E. in The Genetics and Biology of Drosophila Vol. 2c (eds. Ashburner, M. & Wright, T.R.F.) 557–558 (Academic Press, New York, 1978).
Caussinus, E. & Gonzalez, C. Induction of tumor growth by altered stem-cell asymmetric division in Drosophila melanogaster. Nat. Genet. 37, 1125–1129 (2005).
Johnston, L.A. & Schubiger, G. Ectopic expression of wingless in imaginal discs interferes with decapentaplegic expression and alters cell determination. Development 122, 3519–3529 (1996).
Nègre, N., Lavrov, S., Hennetin, J., Bellis, M. & Cavalli, G. Mapping the distribution of chromatin proteins by ChIP on Chip. Methods Enzymol. 410, 316–341 (2006).
Comet, I. et al. PRE-mediated bypass of two Su(Hw) insulators targets PcG proteins to a downstream promoter. Dev. Cell 11, 117–124 (2006).
Neufeld, T.P., de la Cruz, A.F., Johnston, L.A. & Edgar, B.A. Coordination of growth and cell division in the Drosophila wing. Cell 93, 1183–1193 (1998).
Acknowledgements
We are grateful to N. Azpiazu, M. Dominguez, F. Maschat, J. Treisman, L. Jan, S. Artavanis-Tsakonas, J.-M. Dura, Y.N. Jan, J. Müller, M. O'Connor, F. Schweisguth, T. Xu, the Bloomington Stock Center (Indiana University) and the Developmental Studies Hybridoma Bank (University of Iowa) for fly stocks and antibodies (see Supplementary Acknowledgments for details). We thank C. Cazevieille, N. Lautredou and C. Duperray for technical assistance. We also thank F. Bantignies for his contribution on PC foci analysis, T. Sexton for critical reading of the manuscript and members of the Cavalli lab for discussion. G.C.'s research was supported by grants from the CNRS, the Human Frontier Science Program Organization, the European Union FP6 (Network of Excellence the Epigenome and STREP 3D Genome), the Agence Nationale de la Recherche, the Association pour la Recherche sur le Cancer, the Fondation pour la Recherche Médicale and the Ministère de l'Enseignement Supérieur.
Author information
Authors and Affiliations
Contributions
A.-M.M. and G.C. designed experiments and wrote the manuscript. A.-M.M. and S.S. did clonal analysis and immunostaining experiments. B.S. performed RT-PCR and ChIP experiments. A.J. and C.G. performed transplantation experiments. All authors discussed the text of the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Acknowledgments, Supplementary Figures 1–15 and Supplementary Tables 1 and 2 (PDF 4454 kb)
Rights and permissions
About this article
Cite this article
Martinez, AM., Schuettengruber, B., Sakr, S. et al. Polyhomeotic has a tumor suppressor activity mediated by repression of Notch signaling. Nat Genet 41, 1076–1082 (2009). https://doi.org/10.1038/ng.414
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.414