Abstract
To cope with proteotoxic stress, cells attenuate protein synthesis. However, the precise mechanisms underlying this fundamental adaptation remain poorly defined. Here we report that mTORC1 acts as an immediate cellular sensor of proteotoxic stress. Surprisingly, the multifaceted stress-responsive kinase JNK constitutively associates with mTORC1 under normal growth conditions. On activation by proteotoxic stress, JNK phosphorylates both RAPTOR at S863 and mTOR at S567, causing partial disintegration of mTORC1 and subsequent translation inhibition. Importantly, HSF1, the central player in the proteotoxic stress response (PSR), preserves mTORC1 integrity and function by inactivating JNK, independently of its canonical transcriptional action. Thereby, HSF1 translationally augments the PSR. Beyond promoting stress resistance, this intricate HSF1–JNK–mTORC1 interplay, strikingly, regulates cell, organ and body sizes. Thus, these results illuminate a unifying mechanism that controls stress adaptation and growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Balch, W. E., Morimoto, R. I., Dillin, A. & Kelly, J. W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Lindquist, S. The heat-shock response. Annu. Rev. Biochem. 55, 1151–1191 (1986).
Morimoto, R. I. The heat shock response: systems biology of proteotoxic stress in aging and disease. Cold Spring. Harb. Symp. Quant. Biol. 76, 91–99 (2011).
Morimoto, R. I. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes Dev. 22, 1427–1438 (2008).
Dai, C., Dai, S. & Cao, J. Proteotoxic stress of cancer: implication of the heat-shock response in oncogenesis. J. Cell. Physiol. 227, 2982–2987 (2012).
Min, J. N., Huang, L., Zimonjic, D. B., Moskophidis, D. & Mivechi, N. F. Selective suppression of lymphomas by functional loss of Hsf1 in a p53-deficient mouse model for spontaneous tumors. Oncogene 26, 5086–5097 (2007).
Dai, C., Whitesell, L., Rogers, A. B. & Lindquist, S. Heat shock factor 1 is a powerful multifaceted modifier of carcinogenesis. Cell 130, 1005–1018 (2007).
Meng, L., Gabai, V. L. & Sherman, M. Y. Heat-shock transcription factor HSF1 has a critical role in human epidermal growth factor receptor-2-induced cellular transformation and tumorigenesis. Oncogene 29, 5204–5213 (2010).
Jin, X., Moskophidis, D. & Mivechi, N. F. Heat shock transcription factor 1 is a key determinant of HCC development by regulating hepatic steatosis and metabolic syndrome. Cell Metab. 14, 91–103 (2011).
Dai, C. et al. Loss of tumor suppressor NF1 activates HSF1 to promote carcinogenesis. J. Clin. Invest. 122, 3742–3754 (2012).
Dai, S. et al. Suppression of the HSF1-mediated proteotoxic stress response by the metabolic stress sensor AMPK. EMBO J. 34, 275–293 (2015).
Tang, Z. et al. MEK guards proteome stability and inhibits tumor-suppressive amyloidogenesis via HSF1. Cell 160, 729–744 (2015).
Lindquist, S. Regulation of protein synthesis during heat shock. Nature 293, 311–314 (1981).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Inoki, K., Zhu, T. & Guan, K. L. TSC2 mediates cellular energy response to control cell growth and survival. Cell 115, 577–590 (2003).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Kim, E., Goraksha-Hicks, P., Li, L., Neufeld, T. P. & Guan, K. L. Regulation of TORC1 by Rag GTPases in nutrient response. Nat. Cell Biol. 10, 935–945 (2008).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008).
Derijard, B. et al. JNK1: a protein kinase stimulated by UV light and Ha-Ras that binds and phosphorylates the c-Jun activation domain. Cell 76, 1025–1037 (1994).
Kawaguchi, Y. et al. The deacetylase HDAC6 regulates aggresome formation and cell viability in response to misfolded protein stress. Cell 115, 727–738 (2003).
Leu, J. I., Pimkina, J., Frank, A., Murphy, M. E. & George, D. L. A small molecule inhibitor of inducible heat shock protein 70. Mol. Cell 36, 15–27 (2009).
Massey, A. J. et al. A novel, small molecule inhibitor of Hsc70/Hsp70 potentiates Hsp90 inhibitor induced apoptosis in HCT116 colon carcinoma cells. Cancer Chemother. Pharmacol. 66, 535–545 (2010).
Pickart, C. M. & Eddins, M. J. Ubiquitin: structures, functions, mechanisms. Biochim. Biophys. Acta 1695, 55–72 (2004).
Zhang, T. et al. Discovery of potent and selective covalent inhibitors of JNK. Chem. Biol. 19, 140–154 (2012).
Manning, A. M. & Davis, R. J. Targeting JNK for therapeutic benefit: from junk to gold? Nat. Rev. Drug Discov. 2, 554–565 (2003).
Kyriakis, J. M. & Avruch, J. Mammalian MAPK signal transduction pathways activated by stress and inflammation: a 10-year update. Phys. Rev. 92, 689–737 (2012).
Geiger, P. C., Wright, D. C., Han, D. H. & Holloszy, J. O. Activation of p38 MAP kinase enhances sensitivity of muscle glucose transport to insulin. Am. J. Physiol. Endocrinol. Metab. 288, E782–E788 (2005).
Kim, D. H. et al. mTOR interacts with raptor to form a nutrient-sensitive complex that signals to the cell growth machinery. Cell 110, 163–175 (2002).
Hara, K. et al. Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110, 177–189 (2002).
Soliman, G. A. et al. mTOR Ser-2481 autophosphorylation monitors mTORC-specific catalytic activity and clarifies rapamycin mechanism of action. J. Biol. Chem. 285, 7866–7879 (2010).
Gupta, S. et al. Selective interaction of JNK protein kinase isoforms with transcription factors. EMBO J. 15, 2760–2770 (1996).
Lei, K. et al. The Bax subfamily of Bcl2-related proteins is essential for apoptotic signal transduction by c-Jun NH(2)-terminal kinase. Mol. Cell. Biol. 22, 4929–4942 (2002).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl Acad. Sci. USA 109, 413–418 (2012).
Fingar, D. C., Salama, S., Tsou, C., Harlow, E. & Blenis, J. Mammalian cell size is controlled by mTOR and its downstream targets S6K1 and 4EBP1/eIF4E. Genes Dev. 16, 1472–1487 (2002).
Um, S. H., D’Alessio, D. & Thomas, G. Nutrient overload, insulin resistance, and ribosomal protein S6 kinase 1, S6K1. Cell Metab. 3, 393–402 (2006).
Hirosumi, J. et al. A central role for JNK in obesity and insulin resistance. Nature 420, 333–336 (2002).
Clausson, C. M. et al. Increasing the dynamic range of in situ PLA. Nat. Methods 8, 892–893 (2011).
Rosner, M. & Hengstschlager, M. Cytoplasmic and nuclear distribution of the protein complexes mTORC1 and mTORC2: rapamycin triggers dephosphorylation and delocalization of the mTORC2 components rictor and sin1. Hum. Mol. Genet. 17, 2934–2948 (2008).
Bogoyevitch, M. A. & Kobe, B. Uses for JNK: the many and varied substrates of the c-Jun N-terminal kinases. Microbiol. Mol. Biol. Rev. 70, 1061–1095 (2006).
Qian, S. B. et al. mTORC1 links protein quality and quantity control by sensing chaperone availability. J. Biol. Chem. 285, 27385–27395 (2010).
Jez, J. M., Chen, J. C., Rastelli, G., Stroud, R. M. & Santi, D. V. Crystal structure and molecular modeling of 17-DMAG in complex with human Hsp90. Chem. Biol. 10, 361–368 (2003).
Jewell, J. L., Russell, R. C. & Guan, K. L. Amino acid signalling upstream of mTOR. Nat. Rev. Mol. Cell. Biol. 14, 133–139 (2013).
Xiao, X. et al. HSF1 is required for extra-embryonic development, postnatal growth and protection during inflammatory responses in mice. EMBO J. 18, 5943–5952 (1999).
Jorgensen, P. & Tyers, M. How cells coordinate growth and division. Curr. Biol. 14, R1014–R1027 (2004).
Eilers, M. & Eisenman, R. N. Myc’s broad reach. Genes Dev. 22, 2755–2766 (2008).
Freeman, A. et al. Improved detection of hepatocyte proliferation using antibody to the pre-replication complex: an association with hepatic fibrosis and viral replication in chronic hepatitis C virus infection. J. Viral Hepat. 10, 345–350 (2003).
Goyal, L., Muzumdar, M. D. & Zhu, A. X. Targeting the HGF/c-MET pathway in hepatocellular carcinoma. Clin. Cancer Res. 19, 2310–2318 (2013).
Sancak, Y. et al. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Betz, C. & Hall, M. N. Where is mTOR and what is it doing there? J. Cell. Biol. 203, 563–574 (2013).
Cui, H. et al. Enhanced expression of asparagine synthetase under glucose-deprived conditions protects pancreatic cancer cells from apoptosis induced by glucose deprivation and cisplatin. Cancer Res. 67, 3345–3355 (2007).
Chaveroux, C. et al. Identification of a novel amino acid response pathway triggering ATF2 phosphorylation in mammals. Mol. Cell. Biol. 29, 6515–6526 (2009).
Taipale, M., Jarosz, D. F. & Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat. Rev. Mol. Cell. Biol. 11, 515–528 (2010).
Trepel, J., Mollapour, M., Giaccone, G. & Neckers, L. Targeting the dynamic HSP90 complex in cancer. Nat. Rev. Cancer 10, 537–549 (2010).
Zhang, H. et al. Hsp90/p50cdc37 is required for mixed-lineage kinase (MLK) 3 signaling. J. Biol. Chem. 279, 19457–19463 (2004).
Chen, J., Miller, E. M. & Gallo, K. A. MLK3 is critical for breast cancer cell migration and promotes a malignant phenotype in mammary epithelial cells. Oncogene 29, 4399–4411 (2010).
Dong, C. et al. Defective T cell differentiation in the absence of Jnk1. Science 282, 2092–2095 (1998).
Yang, D. D. et al. Differentiation of CD4 + T cells to Th1 cells requires MAP kinase JNK2. Immunity 9, 575–585 (1998).
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007).
Postic, C. et al. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic beta cell-specific gene knock-outs using Cre recombinase. J. Biol. Chem. 274, 305–315 (1999).
Kwiatkowski, D. J. et al. A mouse model of TSC1 reveals sex-dependent lethality from liver hemangiomas, and up-regulation of p70S6 kinase activity in Tsc1 null cells. Hum. Mol. Genet. 11, 525–534 (2002).
Acknowledgements
We would like to thank members of the Dai laboratory for discussions and technical assistance. This work was supported by a grant from the NIH (1DP1OD006438) to I.J.B.; and The Jackson Laboratory Cancer Center Support Grant (3P30CA034196), and grants from the NIH (1DP2OD007070) and the Ellison Medical Foundation (AS-NS-0599-09) to C.D.
Author information
Authors and Affiliations
Contributions
K.-H.S., J.C., Z.T. and S.D. designed and performed the experiments. Y.H. performed statistical analyses and generated graphs. I.J.B. provided mice and cell lines and actively engaged in discussions. C.D. conceived and oversaw this study. S.B.S. and C.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Perturbation of proteostasis by proteotoxic stressors, related to Fig. 1.
(a) Proteotoxic stressors perturb proteostasis via diverse mechanisms of action. Ub: ubiquitin. (b) HEK293T cells were treated with diverse proteotoxic stressors at the concentrations described in Fig. 1B and 1C for 4 and 8 h. Cell viability was measured by a Guava EasyCyte cytometer using ViaCount reagents (mean of 4 wells of cells per group from one experiment). (c) HEK293T cells were treated with 500 nM MG132 for indicated time. JNK and p38 MAPK phosphorylation were detected by immunoblotting. (d) Following pre-incubation with 10 μM SB202190 for 30 min, HEK293T cells were treated with 500 nM MG132 for indicated time. Phosphorylation of p38 MAPK and mTORC1 signalling were detected by immunoblotting. Uncropped images of blots are shown in Supplementary Fig. 8. Source data for Supplementary Fig. 1b can be found in Supplementary Table 1.
Supplementary Figure 2 JNK is a negative regulator of mTORC1 signalling, translation, and cell size, related to Fig. 2.
(a) HEK293T cells stably transduced with lentiviral scramble or JNK1/2-targeting shRNAs were treated with and without 500 nM MG132 for 4 h. Phosphorylated and total JNK1/2 levels were detected by immunoblotting. (b) Following pre-incubation with 3 μM JNK-IN-8 for 60 min, HEK293T cells were treated with and without 500 nM MG132 for 4 h. Endogenous mTOR-RAPTOR associations and mTORC1 kinase activity were measured by coIP. (c) and (d) Primary MEFs were treated with 200 nM MG132 for 6 h, and cell viability was measured using ViaCount reagents (mean of6 wells of cells per group from one experiment). (e) Following transfection with 10 nM three independent MKK7-targetting siRNAs for 4 days, MKK7 and phosphorylated S6K proteins were detected by immunoblotting in HEK293T cells. (f) Following pre-incubation with 3 μM JNK-IN-8 for 60 min, phosphorylated S6K proteins were detected by immunoblotting in HEK293T cells transfected with LacZ, JNK1CA, or JNK2A2 plasmid. (g) Endogenous mTOR-RAPTOR associations were detected by coIP in HEK293T cells transfected with LacZ, JNK1CA, or JNK2A2 plasmid. (h) Following 1 μM AZD8055 treatment overnight, HEK293T cells were labelled with 6-FAM-dc-puromycin for 30 min and analysed by flow cytometry. Unlabelled cells served as negative controls. (i) Following transfection with JNK1/2-targeting siRNAs (A: siJNK1_2 and siJNK2_2; B: siJNK1_3 and siJNK2_3), total JNK1/2 levels were detected by immunoblotting. (j) Following transfection with indicated plasmids, HeLa cells were treated with DMSO or 1 μM AZD8055 for 2 days and analysed for cell size using a Multisizer 3 Coulter Counter. Changes in cell size distribution, compared to the LacZ control, are statistically significant (Kolmogorov-Smirnov test, P < 0.01). (k) and (l) JNK and mTORC1 signalling were detected by immunoblotting in mouse skin and kidneys. (m)–(o) Whole-body weight and composition were measured in 6-week-old male mice (mean ± s.d., n = 5, 5, or 6 mice per genotype, One-way ANOVA). Statistical significance: ∗P < 0.05; ∗∗P < 0.01; ∗∗∗P < 0.001; n.s.: not significant. Uncropped images of blots are shown in Supplementary Fig. 8. Source data for Supplementary Fig. 2c, d can be found in Supplementary Table 1.
Supplementary Figure 3 JNK physically interacts with RAPTOR and mTOR, related to Fig. 3.
(a)–(c) Validations of mouse monoclonal anti-JNK1 (JM2671), rabbit monoclonal anti-mTOR (7C10), and mouse monoclonal anti-RAPTOR (1H6.2) antibodies by immunostaining in HeLa cells stably expressing shRNAs. Scale bars: 50 μm. Images are representative of 2 independent experiments. (d) Endogenous mTORC1 was precipitated using anti-RAPTOR antibodies from HEK293T cells. Precipitates were resolved on SDS-PAGE under non-reducing conditions. Precipitated mTORC1 components were visualized by silver staining (Silver Stain Kit, Thermo Fisher Scientific). To avoid obscuring of mTOR by the staining of abundant whole IgG, the portion containing IgG was separated from the rest of the gel and stained for less time. The rest of the gel was silver-stained for the same period of time to reveal both mTOR and JNK proteins. The p46 isoforms of JNK proteins could not be clearly identified due to the obscuration by IgG heavy chains (HC). MK: molecular weight marker. The staining intensities of mTOR and JNK proteins were quantitated by ImageJ software, and normalized against their total numbers of amino acids. (e) The consensus JNK phosphorylation sequence is derived from curated JNK substrates (www.phosphosite.org) and plotted using the online WebLoGo software (www.weblogo.berkeley.edu). Predicted JNK phosphorylation sites, RAPTOR Ser863 and mTOR Ser567, are highlighted in red. (f) Endogenous mTORC1 complexes were immunoprecipitated from HEK293T cells. Immediately following addition of 100 ng recombinant GST or GST-JNK1 proteins to equal amounts of precipitates, indicated concentrations of JNK-IN-8 (JIN8) or AZD8055 (AZD) were added to the kinase mixtures and incubated at 30 °C for 30 min for in vitro JNK kinase assays. (g) and (h) Following treatment with 500 nM MG132 for 4 h, endogenous RAPTOR and mTOR proteins were precipitated from HEK293T cells stably expressing scramble or JNK1/2-targeting shRNAs. RAPTOR Ser863 and mTOR Ser567 phosphorylation was detected by immunoblotting. (i) and (j) HEK293T cells depleted of endogenous RAPTOR or mTOR proteins due to stably shRNA expression were transfected with indicated plasmids and treated with DMSO or 500 nM MG132 for 4 h. JNK and mTORC1 signalling were detected by immunoblotting.
Supplementary Figure 4 HSF1 suppresses JNK activation and sustains mTORC1 signalling, related to Figs 4 and 5.
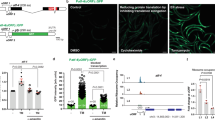
(a) Schematic depiction of the generation of conditional Hsf1fl/fl alleles on the C57BL/6J genetic background. Expression of Cre recombinase results in deletion of exon2-9 (Δ2-9) of the Hsf1 gene. (b) Cells were treated as described in Fig. 4A. Cell viability was measured by a Guava EasyCyte cytometer using ViaCount reagents (mean of4 or 5 wells of cells per group from one experiment). (c) Validations of rabbit polyclonal anti-JNK1/3 (C-17) and mouse monoclonal anti-HSF1 (E-4) antibodies by immunostaining in primary Jnk1+/+ and Jnk1−/− MEFs (left panel), and in immortalized Rosa26-CreERT2; Hsf1fl/fl MEFs treated with 1 μM 4-OHT or EtOH for 7 days (right panel). Scale bars: 50 μm. (d) Endogenous HSF1-JNK1 interactions were detected by PLA in immortalized Rosa26-CreERT2; Hsf1fl/fl MEFs treated with and without 1 μM 4-OHT for 7 days, using mouse anti-HSF1 (E-4) and rabbit anti-JNK1/3 (C-17) antibodies. Scale bars: 10 μm. (e) HEK293T cells were transfected with LacZ or FLAG-tagged HSF1 plasmids. Following IP of endogenous JNK1 proteins, co-precipitated FLAG-tagged HSF1 and endogenous RAPTOR proteins were immunoblotted. (f) Schematic depiction of the proposed sequestration of JNK apart from mTORC1 by HSF1. Uncropped images of blots are shown in Supplementary Fig. 8. Source data for Supplementary Fig. 4b can be found in Supplementary Table 1.
Supplementary Figure 5 mTORC1 inhibition suppresses the PSR, related to Fig. 6.
HEK293T cells were treated with the mTORC1 inhibitor rapamycin (100 nM) or AZD8055 (1 μM) overnight. Translation rates of HSF1 were measured by puromycin labelling as described in Fig. 5b (mean of 6 wells of cells per group per experiment, and this experiment was repeated twice). Source data for Supplementary Fig. 5 can be found in Supplementary Table 1.
Supplementary Figure 6 HSF1 is a positive regulator of cell, organ, and body sizes, related to Fig. 7.
(a) and (b) HEK293T cells were co-transfected with indicated shRNAs and siRNAs. HSF1 and JNK1/2 levels were detected by immunoblotting (a), and protein synthesis rates were measured by puromycin labelling (b). (c) Livers of female mice were weighed at 6 weeks of age (mean ± s.d., n = 6, 7, or 8 mice per genotype, One-way ANOVA). (d) HSP90α, HSP72, and HSP25 proteins were detected by immunoblotting in mouse livers. (e) and (f) Kidneys of male mice were weighed at 6 weeks of age (mean ± s.d., n = 4 mice per genotype, One-way ANOVA). (g) Digital photos of female mice at 6 weeks of age were taken and body surface areas were quantitated by ImageJ software (mean ± s.d., n = 3 mice per genotype, One-way ANOVA). (h) and (i) Body weights were measured in female mice at 6 weeks of age (mean ± s.d., n = 6, 13, 16, or 34 mice per genotype, One-way ANOVA). (j) and (k) Whole-body composition was measured in female mice at 6 weeks of age (mean ± s.d., n = 6, 7, or 18 mice per genotype, One-way ANOVA). Statistical significance: ∗∗P < 0.01; ∗∗∗P < 0.001. Uncropped images of blots are shown in Supplementary Fig. 8. Statistics source data for Supplementary Fig. 6f, g can be found in Supplementary Table 1.
Supplementary Figure 7 HSF1-JNK interactions regulate liver growth and proliferation, related to Fig. 8.
(a) and (b) Liver weights and lean mass of 6-week-old female mice were measured (mean ± s.d., n = number of mice per genotype shown in the graphs, One-way ANOVA). (c) Quantitation of mTORC1 signalling in mouse liver tissues shown in Fig. 7e (mean ± s.d., n = 3 independent experiments, One-way ANOVA). (d) Representative flow cytometry dot plots of co-stained liver cells with PI and anti-BrdU antibodies. (e) and (f) MCM2 and c-MYC levels were detected in mouse liver tissues by immunoblotting. (g) Caspase 3 activities were quantitated in liver tissues of 6-week-old male mice using DEVD-R110 as the substrate (mean ± s.d., n = 3 mice per genotype, One-way ANOVA). This experiment was repeated twice. (h) Hepatocyte-specific deletion of Hsf1 was verified by immunostaining of frozen sections of mouse liver tissues using rat monoclonal anti-HSF1 antibody cocktail (1:100) and CF594 anti-rat IgG secondary antibody conjugates. Arrows indicate hepatocytes and arrowheads indicate non-parenchymal cells. Of note, HSF1 expression was still detected in non-parenchymal cells following Cre-mediated recombination. Scale bars: 50 μm. Representative images from 2 independent experiments. (i) HgfmRNA levels were quantitated by qRT-PCR in primary Jnk1MEFs (mean of 3 wells of cells per genotype per experiment, and this experiment was repeated twice). (j) Hsp mRNA levels were quantitated by qRT-PCR in primary Jnk1 MEFs (mean, of 3 wells of cells per genotype per experiment, and this experiment was repeated twice). (k) HgfmRNA levels were quantitated by qRT-PCR in mouse liver tissues (mean ± s.d., n = 3 mice per genotype, One-way ANOVA). This experiment was repeated twice. (l) and (m) Phosphorylation of c-MET was detected by immunoblotting in mouse liver tissues. Statistical significance: ∗P < 0.05; ∗∗P < 0.01; ∗∗∗P < 0.001; n.s.: not significant. Uncropped images of blots are shown in Supplementary Fig. 8. Statistics source data for Supplementary Fig. 7c, g, i–k can be found in Supplementary Table 1.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2709 kb)
Supplementary Table 1
Supplementary Information (XLSX 65 kb)
Supplementary Table 2
Supplementary Information (XLSX 32 kb)
Supplementary Table 3
Supplementary Information (XLSX 21 kb)
Rights and permissions
About this article
Cite this article
Su, KH., Cao, J., Tang, Z. et al. HSF1 critically attunes proteotoxic stress sensing by mTORC1 to combat stress and promote growth. Nat Cell Biol 18, 527–539 (2016). https://doi.org/10.1038/ncb3335
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3335
This article is cited by
-
A puromycin-dependent activity-based sensing probe for histochemical staining of hydrogen peroxide in cells and animal tissues
Nature Protocols (2022)
-
HSF1 is a driver of leukemia stem cell self-renewal in acute myeloid leukemia
Nature Communications (2022)
-
The regulation of protein translation and its implications for cancer
Signal Transduction and Targeted Therapy (2021)
-
The proteostasis guardian HSF1 directs the transcription of its paralog and interactor HSF2 during proteasome dysfunction
Cellular and Molecular Life Sciences (2021)
-
Broad Spectrum Mixed Lineage Kinase Type 3 Inhibition and HIV-1 Persistence in Macrophages
Journal of Neuroimmune Pharmacology (2019)