Abstract
The thermophilic sulfate-reducing archaeon Archaeoglobus fulgidus strain VC-16 (DSM 4304), which is known to oxidize fatty acids and n-alkenes, was shown to oxidize saturated hydrocarbons (n-alkanes in the range C10–C21) with thiosulfate or sulfate as a terminal electron acceptor. The amount of n-hexadecane degradation observed was in stoichiometric agreement with the theoretically expected amount of thiosulfate reduction. One of the pathways used by anaerobic microorganisms to activate alkanes is addition to fumarate that involves alkylsuccinate synthase as a key enzyme. A search for genes encoding homologous enzymes in A. fulgidus identified the pflD gene (locus-tag AF1449) that was previously annotated as a pyruvate formate lyase. A phylogenetic analysis revealed that this gene is of bacterial origin and was likely acquired by A. fulgidus from a bacterial donor through a horizontal gene transfer. Based on three-dimensional modeling of the corresponding protein and molecular dynamic simulations, we hypothesize an alkylsuccinate synthase activity for this gene product. The pflD gene expression was upregulated during the growth of A. fulgidus on an n-alkane (C16) compared with growth on a fatty acid. Our results suggest that anaerobic alkane degradation in A. fulgidus may involve the gene pflD in alkane activation through addition to fumarate. These findings highlight the possible importance of hydrocarbon oxidation at high temperatures by A. fulgidus in hydrothermal vents and the deep biosphere.
Similar content being viewed by others
Introduction
The anaerobic oxidation of hydrocarbons (for example, n-alkanes, n-alkenes and mono- and poly-aromatic hydrocarbons) is performed by various mesophilic microorganisms that belong to the domain Bacteria (Widdel et al., 2010). A limited number of studies have also reported the ability of thermophilic bacteria to oxidize hydrocarbons. The sulfate-reducing Desulfothermus naphtae strain TD3, which oxidizes n-alkanes at 60 °C, represents the only thermophilic anaerobic hydrocarbon-degrading microorganism isolated thus far (Rueter et al., 1994). In 1997, Chen and Taylor generated an enrichment culture that was able to oxidize monoaromatic hydrocarbons coupled to sulfate-reduction at 50 °C (Chen and Taylor, 1997). More recently, an enrichment culture dominated by Desulfotomaculum-like sulfate-reducing bacteria was shown to oxidize propane anaerobically at 60 °C (Kniemeyer et al., 2007).
Regarding the domain Archaea, three anaerobic species capable of growing on hydrocarbons other than methane have been reported to date. All of these organisms are hyperthermophiles. Although sequential transfers of Thermococcus sibiricus (order Thermococcales) on medium with hexadecane as substrate suggest that this archaeon can potentially metabolize alkanes (Mardanov et al., 2009), there is evidence that two members of the order Archaeoglobales can oxidize such substrates. Ferroglobus placidus oxidizes benzene at 85 °C with Fe(III) as an electron acceptor (Holmes et al., 2011) and Archaeoglobus fulgidus was shown to use unsaturated aliphatic hydrocarbons (n-alkenes) as electron donors at 70 °C with thiosulfate or sulfate as terminal electron acceptors (Khelifi et al., 2010). This latter sulfate-reducing archaeon was first isolated from a shallow submarine hot vent in the Mediterranean Sea (Stetter et al., 1987). Thereafter, this microorganism was recovered from a volcanic environment (Zellner et al., 1989), various hot oil field waters in the North Sea (Stetter et al., 1993; Beeder et al., 1994), a continental oil reservoir (L’Haridon et al., 1995) and from a deep aquifer basin in France (Fardeau et al., 2009). Although enrichment cultures containing A. fulgidus were shown to grow on crude oil as substrate (Stetter et al., 1993), the involvement of this organism in hydrocarbon oxidation has not been clearly demonstrated. Thus, A. fulgidus may be of geomicrobiological significance in shallow and deep hot ecosystems, and this archaeon is potentially able to oxidize hydrocarbons other than alkenes (Khelifi et al., 2010).
To date, several pathways for the anaerobic oxidation of hydrocarbons have been described (for reviews, see Boll et al., 2002; Heider, 2007; Grossi et al., 2008; Boll and Heider, 2010; Zedelius et al., 2011). One of the best documented pathways consists of the addition of the hydrocarbon substrate to fumarate. This pathway involves a benzylsuccinate synthase (Beller and Spormann, 1998; Coschigano et al., 1998; Leuthner et al., 1998) or an alkylsuccinate synthase (Callaghan et al., 2008; Grundmann et al., 2008) for aromatic or saturated hydrocarbons, respectively. Both enzymes belong to the class of glycyl radical enzymes. In the case of aliphatic hydrocarbons, this mechanism of oxidation usually involves activation at the subterminal carbon of the alkyl chain and the subsequent addition of the activated alkane to a molecule of fumarate, yielding alkyl-substituted succinates (Kropp et al., 2000; for review see Callaghan, 2013). Activation at the terminal carbon atom of the hydrocarbon has also been observed for propane (Kniemeyer et al., 2007). Long-chain alkane activation by addition to fumarate was shown to occur for some mesophilic denitrifying (Rabus et al., 2001) or sulfate-reducing (Cravo-Laureau et al., 2005; Callaghan et al., 2006) bacterial strains and some mesophilic enrichment cultures (Coates et al., 1997; Kropp et al., 2000).
Here, we provide evidence of the ability of A. fulgidus to oxidize linear saturated aliphatic hydrocarbons (n-alkanes). First results suggesting involvement of an alkylsuccinate synthase in this process need further exploration.
Materials and methods
Chemicals
Fatty acids and saturated aliphatic hydrocarbons (C10-C21 n-alkanes) were purchased from Sigma-Aldrich (Saint-Quentin Fallavier, France).
Growth of A. fulgidus
The sulfate-reducing archaeon A. fulgidus strain VC-16 (DSM 4304), which was isolated from a marine hydrothermal system (Stetter et al., 1987), was obtained from the Deutsche Sammlung von Mikroorganismen und Zellkulturen (Braunschweig, Germany). Strain VC-16 was selected because it is the type strain of this organism and its genome has been sequenced (Klenk et al., 1997).
A. fulgidus was cultivated in anaerobic bicarbonate-buffered mineral medium reduced with sodium sulfide and L-cysteine hydrochloride (Khelifi et al., 2010) and containing 0.1 g l−1 yeast extract and 10 ml l−1 of a vitamin solution (Wolin et al., 1963) as growth factors. Either Na2S2O3 (15 mM) or Na2SO4 (15 mM) was added as an electron acceptor. The carbon and energy source was either sodium octanoate or n-alkanes, as stated in the text. The medium was dispensed into Bellco tubes (10 ml) or bottles (450 ml) under N2:CO2 (80:20) and sealed with butyl rubber stoppers.
Cultures of A. fulgidus with hydrocarbon as a substrate were inoculated (10%, vol/vol) using unless stated a preculture grown on sodium octanoate (2 mM). Before use, the inoculum was washed twice with substrate-free medium and concentrated four times. Cultures grown in tubes or bottles were supplemented with 0.1% (vol/vol) of C10, C12, C14, C15, C16, C20 or C21 n-alkane, corresponding to a concentration of 5.1, 4.4, 3.8, 3.6, 3.4, 2.8 or 2.7 mM, respectively. All of the tested hydrocarbons were added in undiluted form. For quantitative biodegradation experiments, 5 μl of n-hexadecane was added per tube (corresponding to 17 μmol per culture of 11.6 ml) yielding a nominal concentration of 1.46 mM. Triplicate series of assays, hydrocarbon-free controls (inoculated cultures without hydrocarbon) and abiotic controls (noninoculated cultures with hydrocarbon) were systematically performed and incubated at 70 °C in the dark. Tubes containing hydrocarbons were always incubated upside down, and bottles were incubated horizontally to avoid contact between the hydrocarbons and butyl rubber stoppers.
Analytical techniques
Growth of A. fulgidus was monitored by measuring dissolved sulfide production (Cline, 1969) and electron acceptor consumption (thiosulfate or sulfate). Thiosulfate and sulfate concentrations were quantified by ion chromatography (761 Compact IC Metrohm, Metrohm, Villebon-sur-Yvette, France) equipped with a Metrosep Anion Supp1 column (Metrohm). Acetate concentration was determined by HPLC (Thermo Scientific, Villebon-sur-Yvette, France) equipped with an Aminex HPX-87H-300 × 7.8 column (Bio-Rad, Marnes-la-Coquette, France).
The hexadecane concentration was quantified by gas chromatography after extraction of the cultures with heptane following the addition of pristane as an internal standard. Gas chromatography (Perichrom PR2100, Alpha MOS, Toulouse, France) was equipped with a flame ionization detector (250 °C) and capillary column (25 m × 0.53 mm × 1 μm) containing a CP-Sil 5CB stationary phase. The oven temperature was initially held at 100 °C for 10 min and increased to 170 °C at 4 °C min−1. The recovery of the alkane during extraction was ∼98%.
RNA extraction and cDNA synthesis
Total RNAs were extracted from cultures (450 ml) of A. fulgidus grown with either n-hexadecane (C16) or sodium octanoate (C8) as a growth substrate. To obtain cells in comparable physiological states, cells were harvested after 5 days and 7 weeks of incubation with octanoate and hexadecane as carbon sources, respectively. Cells were harvested by centrifugation (20 min, 5800 g, 4 °C), and pellets were washed with 2 ml of RNase-free 10 mM Tris-HCl (pH 8). Cells were pelleted again (2 min, 10 900 g, 4 °C) and stored at −80 °C until use. RNA extractions were carried out with the High Pure RNA isolation kit (Roche Diagnostics, Meylan, France) according to the manufacturer’s instructions with the following modifications. Cells were suspended in 100 μl of buffer (Tris HCl 10 mM, EDTA 1 mM (pH 8) RNase free), and 4 μl lysozyme (40 mg ml−1) was added before incubation (10 min, 37 °C). Cell suspensions were then subjected to a thermal shock (frozen in liquid nitrogen and then incubated at 60 °C for 3 min). Then, 400 μl of lysis buffer (4.5 M guanidine-HCl, 50 mM Tris-HCl, 30% Triton, pH 6.6) was added. The sample was transferred to a microcentrifuge tube to isolate total RNA. After centrifugation (15 s, 6700 g), the flow-through liquid was discarded. Cells were resuspended in 75 μl of DNase buffer (1 M NaCl, 20 mM Tris-HCl, 10 mM MnCl2, pH 7), and 5 μl of DNase (90 U) was added before incubation at room temperature for 30 min. Successive washes with 500 μl of wash solutions 1 and 2 (High Pure RNA isolation kit, Roche Diagnostics) were performed twice to remove any residual traces of DNA. Finally, RNA was eluted with 50 μl of elution buffer.
RNA was quantified at 260 nm with a Nanodrop (Thermo Scientific). RNA quality was checked by agarose gel electrophoresis (1%, 40 mM Tris-acetate buffer, 1 mM EDTA (pH 8.3)). The absence of genomic DNA contamination was confirmed by PCR using specific primers designed to amplify the A. fulgidus pflD gene (locus Tag AF1449; Supplementary Table 1).
For complementary DNA (cDNA) synthesis, 10 μg of total RNA and 3 μg of random primers (Invitrogen, Cergy Pontoise, France) were mixed, heated to 70 °C for 3 min and placed on ice. The cDNA synthesis mix (50 mM Tris-HCl (pH 8.3), 40 mM KCl, 6 mM MgCl2, 10 mM DTT, 0.3 mM dNTPs) was then added. The reaction mix (30 μl) was incubated for 5 min at 25 °C, and 300 units of Superscript II reverse transcriptase (Invitrogen) was then added. The final reaction mix was incubated for 5 min at 25 °C, followed by 1 h at 42 °C and another 15 min at 70 °C, and the volume was adjusted to 100 μl with ultrapure water.
Two cultures grown on n-hexadecane and two cultures grown on sodium octanoate were used to prepare cDNAs that were preserved at −80 °C for further use.
Relative quantification of the cDNAs was carried out with a LightCycler System using the LightCycler FastStart DNA Masterplus SYBR Green I kit (Roche Diagnostics) according to the manufacturer’s instructions. The LightCycler was programmed for an initial step at 95 °C for 480 s, followed by 45 thermal cycles at 95 °C for 15 s, 57 °C for 15 s and 72 °C for 20 s. The specificities of accumulated products were verified using a melting curve analysis. Non-degenerate primer pair AF1449-forward/reverse was used to specifically amplify the pflD transcript (Supplementary Table 1). The relative expression software tool (REST) was used to calculate the relative expression of each gene at each condition (Pfaffl, 2001) using the 16S RNA gene (rrs, locus tag AFrRNA02) as a reference for normalization (primer pair AF16S-forward/reverse, Supplementary Table 1). Quantification on each cDNA sample was performed in triplicate experiments.
Genome analysis and phylogenetic studies
Complete proteomes from Archaea and Bacteria available at the National Center for Biotechnology Information (NCBI) in February 2013 were downloaded and used to build a local database. AF1449 (PflD) and AF1450 (PflC) sequences from A. fulgidus (776 and 302 amino acids in length) were used as seeds to query this local database with the BLASTP program (Altschul et al., 1997). Because our goal was to determine the evolutionary origin of the PflD and PflC A. fulgidus sequences and not to perform the exhaustive analysis of the entire glycyl radical enzyme superfamily, we retrieved only the homologs displaying an e-value of <10−10. For AF1449, we also discarded the few sequences with an atypically large or small size, keeping only the homologs with sizes ranging from 500 to 1000 amino acids. We also queried the protein RefSeq database at the NCBI using BLASTP and the same parameters to query the few archaeal complete genomes that came out during spring 2013. This allowed us to detect AF1449 and AF1450 homologs in Archaeoglobus sulfaticallidus PM70-1, T. litoralis DSM 547 and M. formicicum DSM 3637. In total, we retrieved 1377 and 1416 homologs of AF1449 and AF1450, respectively. These sequences were aligned using MAFFT version 7 (Katoh and Standley, 2013) with the default parameters. The resulting alignments were visually inspected using SEAVIEW version 4 (Gouy et al., 2010) and trimmed using BMGE (BLOSUM30 option) (Criscuolo and Gribaldo, 2010). Based on preliminary phylogenetic analyses, the closest homologs of AF1449 and AF1450 were selected for additional phylogenetic analyses. To avoid the overrepresentation of a few species (for example, more than 50 Escherichia coli genomes were completely sequenced at the time of analysis), we kept only one representative strain per species. In total, we selected 229 AF1449 and 228 AF1450 homologs for the final phylogenetic analyses. The corresponding sequences were realigned using MAFFT with the linsi option that allows constructing accurate alignments. The resulting alignments were inspected and trimmed as described above.
Preliminary phylogenetic analyses were performed with FastTree version 2.1.4 (options-cat 4–gamma) (Price et al., 2010). Local supports of the resulting tree branches were computed using the Shimodaira–Hasegawa test. Maximum likelihood (ML) trees were inferred with PhyML version 3.1 (Guindon et al., 2010) with the LG model (Le and Gascuel, 2008). We included a γ-distribution with four categories (Γ4) to take into account the heterogeneity of evolutionary rates among sites. The α-parameter was estimated by PhyML. The branch robustness of ML trees was estimated using the nonparametric approach implemented in PhyML (100 replicates of the original alignments). Bayesian inferences were performed with MrBayes version 3.1.2 (Ronquist et al., 2012). MrBayes was run with a mixed substitution model and a Γ4 distribution. Four chains were run in parallel for 1 000 000 generations. The first 2000 generations were discarded as burn-in. The remaining trees were sampled every 100 generations to build consensus trees.
Building of (1-methyl-pentadecyl)succinates
Models were built with the MarvinSketch chemical editor from the Chemaxon package (http://www.chemaxon.com/marvin/sketch/index.php). The compound (1-methyl-pentadecyl)succinate contains two asymmetric carbons; therefore, four enantiomers were generated, as shown in Supplementary Figure S1.
Detection of cavities within the protein PflD (AF1449)
Three programs were used to detect and analyze pockets and cavities in the three-dimensional structure of PflD. Cavities on the surface of the protein (PDB code 2F3O; Lehtiö et al., 2006) were detected with CASTp (http://sts.bioengr.uic.edu/castp/calculation.php; Liang et al., 1998). A total of 131 pockets were detected on the surface of AF1449. The deepest cavity exhibited a molecular volume of 478.8 Å3. CAVER 3.0 (http://www.caver.cz/) was used to calculate putative tunnels and channels in AF1449 (Petrek et al., 2006; Chovancova et al., 2012). A tunnel corresponding to the deepest cavity identified with CASTp was detected using CAVER (Supplementary Figures S2 and S3). The program Hollow 1.2 was used to detect and generate high-resolution representations of the major cavity (Ho and Gruswitz, 2008). This program can be freely downloaded at http://hollow.sourceforge.net/.
Modeling of alkyl ligands in the binding site cavity
The four enantiomers of (1-methyl-pentadecyl)succinate were positioned manually and individually in the deepest detected PflD cavity with CASTp. The ensemble was then minimized in explicit water box with periodic conditions with the CHARMM package version 33b1 and CHARMM force field version 22 with CMAP correction (MacKerell et al., 2004; Brooks et al., 2009). The final system containing 35 477 water molecules, 50 sodium ions and 26 chloride ions was heated at 300 K and equilibrated for 1 ns.
Targeted molecular dynamics
The exit of (1-methyl-pentadecyl)succinate derivatives from the catalytic site cavity was modeled using interactive molecular dynamics simulations with NAMD implemented in the VMD package (http://www.ks.uiuc.edu/Research/vmd/imd/). Typically, a constant force directed toward the outside was applied to the whole ligand during the 15 ns MD simulation.
Results
Growth of A. fulgidus on n-alkanes
Because A. fulgidus is known to oxidize a wider range of substrates when using thiosulfate compared with sulfate as a terminal electron acceptor (Stetter, 1988), and thiosulfate reduction is a more thermodynamically favorable reaction than sulfate reduction for oxidizing substrates (Thauer et al., 1977), thiosulfate was used in our growth experiments.
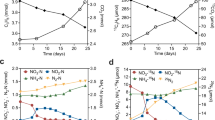
Using cells that had been pregrown on octanoate as an inoculum, cultures of A. fulgidus strain VC-16 (DSM 4304) with n-hexadecane as substrate presented a significant thiosulfate-reducing activity after 8 weeks of incubation. The soluble sulfide production reached 4.1±0.3 mM in the A. fulgidus cultures with hexadecane and only 1.3±0.1 mM in inoculated hexadecane-free controls. Concomitantly, whereas 3.2±0.1 mM of thiosulfate was consumed in the A. fulgidus hexadecane cultures, only 0.7±0.1 mM of thiosulfate was consumed in the hexadecane-free controls (Figure 1a). The ratio obtained between the thiosulfate consumed and sulfide produced is far from that expected (1:2) because only soluble sulfide was measured. Depending on the pH of the culture, soluble sulfide represents only 55–58% of the total sulfide, explaining the high ratio of thiosulfate consumption vs soluble sulfide production. Furthermore, significant differences in soluble sulfide production between the A. fulgidus cultures with hexadecane and alkane-free controls were observed after only 3 weeks of incubation (Supplementary Figure S4) when using cells pregrown on hexadecane as the inoculum.
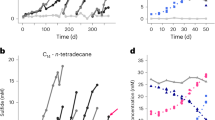
(a) Thiosulfate consumed (gray bars) and soluble sulfide produced (open bars) in cultures of Archaeoglobus fulgidus strain VC-16 after 8 weeks of incubation at 70 °C with n-hexadecane. (b) Thiosulfate consumed in cultures of A. fulgidus strain VC-16 after 16 weeks of incubation at 70 °C with seven different n-alkanes. In both experiments, cultures were inoculated with octanoate-grown cells. Hexadecane was added at a concentration of 3.4 mM, whereas other alkanes were added at similar concentration of carbon. Controls, hydrocarbon free-cultures. The data represent the mean values of triplicate cultures±s.d.
Quantitative growth experiments were carried out with n-hexadecane. After 10 weeks of incubation, after subtracting consumption in biotic and abiotic controls, 3.79±0.99 μmol of hexadecane were oxidized and 54.55±4.72 μmol of thiosulfate were consumed for hexadecane oxidation by the cells (Table 1). Thiosulfate-reducing activity was thus linked to the presence of the alkane in the growth medium.
The theoretical equations for complete hexadecane oxidation (Equation 1) or partial oxidation to acetate (Equation 2) linked to thiosulfate reduction are as follows:


Both the ratios of thiosulfate consumed to alkane oxidized (14.4) and the fact that acetate was not detected in the cultures indicated that A. fulgidus performed a complete oxidation of hexadecane to CO2. The average experimental ratio was close to the theoretical ratio (12.25, Equation 1). However, although a ratio slightly lower than 12.25 (Equation 1) was expected because a small fraction of alkane was likely used in anabolic reactions, the experimental ratio was even higher.
The capacity of hydrocarbon oxidation by A. fulgidus was further tested with other n-alkanes ranging from C10 to C21 (Figure 1b and Supplementary Figure S4). After 16 weeks of incubation, the thiosulfate reduction activity was clearly higher in cultures performed in the presence of n-alkanes compared with hydrocarbon-free controls (Figure 1b). When normalized by time, thiosulfate consumed on hexadecane (Figure 1b) was proportionally higher after 16 weeks of incubation than previously observed after only 8 weeks of incubation (Figure 1a), indicating growth of the population. These results demonstrated that A. fulgidus strain VC-16 was able to grow on the seven different n-alkanes tested using thiosulfate as a terminal electron acceptor. When sulfate was used instead of thiosulfate, and considering the hydrocarbon free controls, 5.8±1.1 mM sulfate were consumed during 7 months of incubation, showing that this strain was also able to oxidize n-hexadecane in the presence of sulfate as electron acceptor.
What is the alkane degradation pathway?
Aerobic microorganisms that degrade alkanes have been studied for a long time; the alkane-activating enzymes overcome the inert nature of the substrate by inserting a reactive oxygen species derived from molecular oxygen. The resulting alcohols are then channeled via fatty acids into β-oxidation (Rojo, 2009). Under anaerobic conditions, one possibility for n-alkane activation is to occur via the addition to fumarate at a secondary carbon atom, resulting in a substituted succinate (Kropp et al., 2000). This reaction is expected to be similar to the anaerobic activation of toluene that leads to benzylsuccinate via a benzylsuccinate synthase (Leuthner et al., 1998). Genes encoding alkylsuccinate synthases have been identified in Desulfatibacillum alkenivorans AK-01 and Desulfoglaeba alkanexedens (Callaghan et al., 2008; Callaghan et al., 2010) and these genes have been annotated as assA genes. Similar genes have been annotated in Azoarcus sp. HxN1 (Grundmann et al., 2008) and Aromatoleum sp. OcN1 (Zedelius et al., 2011) and annotated as masD genes. The detection of assA genes in hydrocarbon-affected environments and enriched cultures has revealed a considerable diversity of these genes (Callaghan et al., 2010).
A BLASTP search (http://blast.ncbi.nlm.nih.gov) against the A. fulgidus genome using the α-subunit of the alkylsuccinate synthase 1 (AssA1, ABH11460) from D. alkenivorans strain AK-01 (Callaghan et al., 2012) identified the product of the gene pflD (locus tag AF1149) as the best hit, with a maximum of identity of 30% (max score 290; e-value: 8e−86). Using AssA2 (ABH11461) from the same strain provided similar results (30% of identity; max score 296; e-value: 2e−88). The same search using the benzylsuccinate synthase (BssA, CAA05052) from Thauera aromatica K172 (Leuthner et al., 1998) as a query also returned the product of the locus tag AF1449 as the best hit (27% of identity; max score 238; e-value: 1e−67). A similar search using the pyruvate formate lyase Pfl1 from E. coli (accession number NP_415423) also returned the locus tag AF1449 as best hit but with a lower score (identity 23%; max score 140; e-value: 6e−37).
The complete sequence alignment with alkyl- and benzylsuccinate synthases, as well as some pyruvate formate lyases (PFLs), confirmed the higher similarity of A. fulgidus PflD (locus tag AF1449) with AssA and BssA, compared with PFL sequences (Figure 2). In particular, the multiple sequence alignment revealed several conserved deletion regions that were specific either to PFLs or to AssA/BssA. Furthermore, enzymes acting as pyruvate formate lyases, as Pfl1 from E. coli, display two adjacent conserved cysteine residues, whereas at this site alkyl- and benzylsuccinate synthases like A. fulgidus PflD contain only one conserved cysteine that accepts the radical from the glycyl residue and initiates catalysis as a thiyl radical (Knappe et al, 1993; Leuthner et al., 1998; Callaghan et al. 2008) (Figure 2, box).
Multiple sequence alignment of A. fulgidus PflD with known alkylsuccinate synthases, benzylsuccinate synthases and PFLs. The N-terminal and C-terminal regions are not shown because of their extreme variability. Conserved residues are shaded in colors. Box corresponds to the sequence region around the catalytic cysteine accepting the radical from the glycyl residue (Knappe et al., 1993). ASSA_Dal, Desulfatibacillum alkenivorans AK-01 (YP_002430896 and YP_002431360); Afulgidu, Archaeoglobus fulgidus DSM 4304 (NP_070278); Asulfati, Archaeoglobus sulfaticallidus PM70 1 (AGK61141); BSSA_Tsp, Thauera sp. DNT-1 (BAC05501); BSSA_Msp, Magnetospirillum sp. TS-6 (BAD42366); BSSA_Asp, Azoarcus sp. T (AAK50372); BSSA_Tar, Thauera aromatica (AAC38454 and CAA05052); BSSA_Dto, Desulfobacula toluolica Tol2 (YP_006759014, YP_006759359, and YP_006760426); BSSA_Aar, Aromatoleum aromaticum EbN1 (YP_158060); BSSA_Gda, Geobacter daltonii FRC-32 (YP_002537892 and YP_002537902); BSSA_Gme, Geobacter metallireducens GS-15 (YP_006720507); PFL1_Eco, Escherichia coli K12 (NP_415423); PFL1_Hin, Haemophilus influenzae 86-028NP (AAX87237); PFL1_Cpa, Clostridium pasteurianum (CAA63748); Tsibiric, Thermococcus sibiricus MM 739 (YP_002994045).
In the A. fulgidus genome, the pflD gene (locus tag AF1449) is immediately followed by a gene annotated as pflC (locus tag AF1450) that encodes a pyruvate formate lyase-activating enzyme (Klenk et al., 1997). Using PflC as seed, a BLASTP search against the genome of D. alkenivorans strain AK-01 identified AssD1 (YP_002430893), AssD2 (YP_002431363) and AssD2’ (YP_002429341) from the Ass1 and Ass2 operon (% of identity: 41%, 41% and 37%; max score: 251, 242 and 171; e-values: e−82, 2e−78 and 3e−51, respectively). This amino acid sequence also exhibited 38% identity (max score 224; e-value: 8e−73) with the BssD (CAA05050) subunit of the benzylsuccinate synthase from T. aromatica K172 but only 29% of identity (max score 114; e-value: 3e−30) with the PflA-activating enzyme of E. coli (NP_415422). The N-terminus of A. fulgidus PflC contained a CX3CX2C sequence motif (box 1, Figure 3) coordinating a [Fe4S4] cluster essential for SAM-radical enzymes (Sofia et al., 2001). This region of the protein also contained two cysteine-rich regions (box 2, Figure 3) that would be involved in [FeS] cluster binding, a feature that is unique to alkyl- and benzylsuccinate synthases and not found in related activating enzymes of PFLs and anaerobic ribonucleotide reductase (Leuthner et al., 1998; Callaghan et al. 2008). Altogether, these data suggest that PflD and PflC sequences from A. fulgidus are more similar to those of alkyl- or benzylsuccinate synthases and activating enzymes, respectively, than to enzymes acting as pyruvate formate lyase as E. coli Pfl1. To go further, an exhaustive phylogenetic analysis of the catalytic and activating enzymes was performed.
Multiple sequence alignment of A. fulgidus PflC with known alkylsuccinate synthases, benzylsuccinate synthases activating proteins and PflA. The N-terminal and C-terminal regions are not shown because of their extreme variability. Conserved residues are shaded in colors. Boxes 1 and 2 correspond to the cysteine-rich regions involved in FeS clusters binding. ASSD_Dal: Desulfatibacillum alkenivorans AK-01 (YP_002430893, YP_002431363 and YP_002431362); Afulgidu, Archaeoglobus fulgidus DSM 4304 (NP_070279); Asulfati, Archaeoglobus sulfaticallidus PM70 1 (AGK60811); BSSD_Tsp, Thauera sp. DNT-1 (BAC05499); BSSD_Msp, Magnetospirillum sp. TS-6 (BAD42364); BSSD_Asp, Azoarcus sp. T (AAK50370); BSSD_Tar, Thauera aromatica (AAC38452 and CAA05050); BSSD_Dto, Desulfobacula toluolica Tol2 (YP_006759016, YP_006759355 and YP_006759356); BSSD_Aar, Aromatoleum aromaticum EbN1 (YP_158058), BSSD_Gda: Geobacter daltonii FRC-32 (YP_002537894 and YP_002537904); BSSD_Gme: Geobacter metallireducens GS-15 (YP_006720509); PFLA_Eco, Escherichia coli K12 (NP_415422); PFLA_Hin, Haemophilus influenzae 86-028NP (YP_247896); PFLA_Cpa, Clostridium pasteurianum (CAA63749); Tsibiric, Thermococcus sibiricus MM 739 (YP_002994043).
Phylogeny of A. fulgidus pflD and pflC gene products
Our survey of complete archaeal and bacterial genomes revealed numerous homologs of the PflD (AF1449) and PflC (AF1450) proteins, among which 1377 and 1416 sequences, respectively, matched our search criteria (see Materials and methods). Nearly all of these homologs belonged to Bacteria (mainly Firmicutes, Deltaproteobacteria, Gammaproteobacteria and Actinobacteria; Supplementary Figure S5). In sharp contrast, only 10 archaeal homologs were identified, although more than 180 complete archaeal genomes were available at the time of the analysis. In addition to A. fulgidus strain VC-16, both proteins were present in the Korarchaeon (Candidatus Korarchaeum cryptofilum OPF8) and in eight Euryarchaeota: one additional member of Archaeoglobales (Archaeoglobus sulfaticallidus PM70-1), three Thermococcales (Thermococcus litoralis DSM 5473, Thermococcus kodakarensis KOD1 and Thermococcus sibiricus MM 739), two Methanobacteriales (Methanobacterium formicicum DSM 3637 and Methanothermobacter thermautotrophicus str. Delta H) and two members of the DHVE2 cluster (Aciduliprofundum boonei T469 and Aciduliprofundum sp. MAR08-339). Preliminary phylogenetic analyses showed that the PflD and PflC trees are consistent (Supplementary Figure S5), although the latter is less resolved (lower Shimodaira–Hasegawa-like values) than the former, which is not surprising given the very restricted number (143 compared with 542) of amino acid positions that could be kept for the phylogenetic analysis. This consistency indicates that the two corresponding genes have undergone roughly similar evolutionary histories (that is, they have been vertically or horizontally transmitted together during their evolutionary history). This is not surprising given that these genes are functionally linked and located in close proximity in most genomes. In both trees, bacterial and archaeal sequences are intermixed, meaning that they do not form two separated groups, as expected if the two genes were present in the common ancestor of Bacteria and Archaea and vertically transmitted during the diversification of these two domains (Supplementary Figure S5). This pattern suggests that horizontal gene transfers (HGTs) occurred between Bacteria and Archaea domains during their diversification. The very limited number of homologs detected in Archaea and their scattering among the bacterial sequences suggests that several HGTs occurred from Bacteria to Archaea. HGTs also occurred among Bacteria, given that the phylogenies of PflC and PflD are inconsistent with the current bacterial taxonomy (for instance, the monophyly of each bacterial phyla is not recovered).
A close examination of the PflD tree showed a deep and well-supported split (Supplementary Figure S5). Interestingly, the archaeal sequences (including PflD from A. fulgidus) belonged to the same subtree than AssA and BssA sequences, whereas the Pfl1 sequences from E. coli K12 and H. influenzae belonged to the other. The PflC tree presented a similar pattern. This indicates that A. fulgidus PflD and PflC sequences, as well as the other archaeal sequences, are more closely related to AssA/BssA and AssD/BssD than to Pfl1.
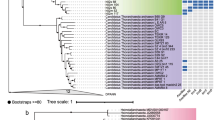
To obtain a clearer picture of the evolutionary origin of A. fulgidus sequences, we performed a second phylogenetic analyses focused on the A. fulgidus PflD and PflC sequences and their closest homologs (Figure 4 and Supplementary Figures S6 and S7). Bacterial and archaeal sequences appeared again intermixed, confirming that archaeal sequences were likely acquired through HGTs from Bacteria. More precisely, at least three independent HGTs occurred from different bacterial donors. The first was an HGT to the common ancestor of A. fulgidus strain VC-16 and A. sulfaticallidus PM70-1. The second was an HGT to the ancestor shared by M. formicicum DSM 3637 and M. thermautotrophicus str. Delta H. The third was an HGT to Candidatus Korarchaeum cryptofilum OPF8, to one of the three Thermococcales (T. litoralis DSM 5473, T. kodakarensis KOD1 or T. sibiricus MM 739), or to the ancestor of A. boonei T469 and Aciduliprofundum sp. MAR08-339. Additional secondary HGTs are likely to explain the close evolutionary link between the sequences carried by these unrelated archaeal lineages (Brochier-Armanet et al., 2011).
Maximum-likelihood tree of the homologs belonging to the same subtree than A. fulgidus PflD (229 sequences, 532 positions) (see Supplementary Figure S6). The colors correspond to major taxonomic groups (red: Archaea, light blue: Gammaproteobacteria, green: Deltaproteobacteria, pink: Firmicutes, yellow: Fusobacteria, dark blue: Bacteroidetes/Chlorobi, black: other bacterial phyla). For clarity, some well-supported clusters were collapsed. The complete tree is shown as Supplementary Figure S7. Numbers at the branches correspond to bootstrap values (for clarity, only values >50% are shown). The scale bar represents the average number of substitutions per site.
Quantification of functional gene transcription
The expression level of pflD in cells grown with either n-hexadecane or octanoate (considered here as a control) was evaluated in the presence of thiosulfate as a terminal electron acceptor. The expression ratio of this gene following growth on these two substrates as measured by quantitative reverse transcriptase-PCR revealed that the amount of pflD transcript was 127.1±14.8-fold higher when cells were cultured on n-hexadecane than when cultured on octanoate. The pflD gene was thus upregulated when cells were grown in the presence of n-hexadecane, suggesting that the encoded protein played an important function in the alkane oxidation process.
Structural analysis of PflD (AF1449) and molecular dynamic simulations
The three-dimensional structure of PflD (AF1449) has been previously obtained (Lehtiö et al., 2006), and the authors concluded that this enzyme was not a pyruvate formate lyase because it did not possess the cysteine pair found in PFLs but did have the arginine residues that were believed to be required for catalysis in PFLs. It was further hypothesized that this enzyme was not a benzylsuccinate synthase as the active site was too small to accommodate the alkyl-substituted monoaromatic hydrocarbon substrate (Lehtiö et al., 2006).
We examined whether this protein was able to accommodate n-alkanes. A number of tools have been developed to detect and analyze cavities and tunnels in protein surfaces (Brezovsky et al., 2012). Three software programs were applied to the available X-ray structure of PflD and revealed a large number of cavities at the protein surface. The deepest cavity formed a tunnel that was sufficiently large and deep to accommodate a hexadecane molecule. This tunnel was surrounded by hydrophobic residues and exhibited a total molecular volume of ∼480 Å3.
In the X-ray structure, this cavity was occupied by an electron density that was previously modeled as triethylene glycol and glycerol for the tunnel and active site, respectively (Lehtiö et al., 2006). Because of the low resolution of the X-ray structure (2.9 Å), the authors noted that the tunnel could be occupied by an unknown hydrophobic co-substrate molecule bound to the enzyme. We looked for the absence of steric hindrance between PflD and the alkylsuccinate that was formed by addition of the sub-terminal carbon atom (C-2) of hexadecane to fumarate. This alkylsuccinate contains two asymmetric carbons and thus can exist as four possible stereoisomers (Supplementary Figure S1). These four isomers were modeled and docked into the protein structure (Figure 5 and Supplementary Figures S2 and S3). Each molecule had an estimated molecular volume of ∼370 Å3; therefore, all four stereoisomers fit within the protein tunnel (Figure 5). Targeted molecular dynamic simulations further showed that the four stereoisomers of (1-methyl-pentadecyl)succinate could exit the protein tunnel. Molecular modeling thus indicates that n-hexadecane could be able to penetrate through the protein tunnel to reach the catalytic site. Following addition to fumarate, the resulting alkylsuccinate could exit the enzyme. A similar analysis on two other PFLs with a known three-dimensional structure (pyruvate formate lyase 1 from E. coli (PDB code 3PFL) and B12-independent glycerol dehydratase from Clostridium butyricum (PDB code 1R9E)) did not reveal the presence of a deep hydrophobic cavity, needed to accommodate a long-chain alkylsuccinate (data not shown). Using a mesophilic nitrate-reducing bacterial strain, Jarling et al. (2012) demonstrated that the alkylsuccinate synthase has an imperfect stereoselectivity and yields only two of the four possible stereoisomers. Our simulations indicate that all four isomers may be produced by A. fulgidus, but further studies are needed to determine which stereoisomers are effectively produced.
Sliced view of the three-dimensional (3D) structure of A. fulgidus PflD (PDB code 2F3O). The deepest cavity at the surface of PflD can accommodate long alkyl chains, such as hexadecane or (1-methyl-pentadecyl)succinate (the (2R, 2’R) isomer is shown here). The insert shows a top view of the protein cavity that is sufficiently large for the entry of hexadecane and exit of (1-methylpentadecyl)succinate. The deepest cavity detected with the Hollow program (http://hollow.sourceforge.net/) is highlighted with red wireframe. The figure was prepared with the PyMOL software (DeLano, 2005).
Discussion
We recently reported that A. fulgidus strain VC-16 was able to grow on a wide range of unsaturated hydrocarbons (C12-C21 n-alk-1-enes) and fatty acids (butyrate, valerate, octanoate, nonanoate, palmitate and stearate) when cultures were incubated for 1 month at 70 °C under anaerobic conditions in the presence of thiosulfate as the terminal electron acceptor (Khelifi et al., 2010). Attempts to grow strain VC-16 on some saturated aliphatic hydrocarbons (dodecane, hexadecane and pristane) were also investigated, but none of these alkanes induced growth after 1 month of incubation (Khelifi et al., 2010).
In the present study, cultures of A. fulgidus with C10–C21 n-alkanes were incubated for several months at 70 °C in the presence of thiosulfate, and significant differences in thiosulfate consumption between cultures of A. fulgidus and controls were observed after only 2 months of incubation. The relative increase in sulfide production with time (Figure 1) suggested growth of A. fulgidus on these saturated hydrocarbons.
Interestingly, Mbadinga et al. (2012) recently showed that the archaeal community of a thermophilic methanogenic enrichment culture oxidizing long-chain alkanes in the absence of sulfate as terminal electron acceptor was dominated by members of the order Archaeoglobales. Although the dominant phylotype was moderately related to cultured organisms (90–93% similarities), these data together with our work suggest that some Archaeoglobales-related species, such as A. fulgidus or relatives (for example, A. sulfaticallidus PM70-1), are involved in the anaerobic oxidation of long-chain hydrocarbons. This further highlights the ecological role that could be occupied by A. fulgidus and relatives in extreme environments (for example, shallow hydrothermal vents, oil reservoirs and so on) where hydrocarbons are available (Konn et al., 2009) either in the presence of sulfate as terminal chemical or of hydrogenotrophic methanogens as biological electron acceptor. This role is of particular interest in oil reservoirs, where hydrogen-scavenging methanogens have been demonstrated to actively participate to methanogenesis from crude oil (Jones et al., 2008) but where microorganisms involved in the first steps of hydrocarbon oxidation have not been identified. A. fulgidus, which has been isolated many times from deep hot oil reservoirs (Stetter et al., 1993; Beeder et al., 1994; L’Haridon et al., 1995), may be one of the organisms involved in alkane oxidation. Further studies are required to know whether this archaeon is able to interact with hydrogenotrophic methanogens or use sulfate for oxidizing hydrocarbons in situ that could be of geomicrobiological significance in deep hot oil reservoirs, where the presence of methanogens or sulfate as possible terminal electron acceptors are common features (Magot et al., 2000; Ollivier and Alazard, 2010). In the case of hydrocarbon oxidation via sulfate reduction by A. fulgidus, the sulfide produced has to be taken into account regarding its pernicious effects (souring and biocorrosion) in the oil industry (Magot et al., 2000).
Within the genome of A. fulgidus strain VC-16, we were able to detect a gene that was strongly upregulated in the presence of hexadecane, and that resembles to the catalytic subunit of a bacterial alkylsuccinate synthase gene. This gene, named pflD (locus tag AF1449), was previously annotated as a pyruvate formate lyase family encoding gene, a group of glycyl radical enzymes (Klenk et al., 1997). An analysis of the three-dimensional structure of PflD (Lehtiö et al., 2006) indicated that the linear hexadecane and the four possible (1-methyl-pentadecyl)succinates had dimensions compatible with the tunnel leading to the active site and with the active site itself (Figure 5).
A full biochemical and enzymatic characterization of the A. fulgidus PflD is still needed to provide direct evidence for the alkylsuccinate synthase functionality in A. fulgidus. However, our observations support the hypothesis that PflD may act as an alkylsuccinate synthase and that this enzyme would be responsible for the first step of n-alkane oxidation in A. fulgidus strain VC-16. The hypothesis could be extended to A. sulfaticallidus PM70-1 because of the close relationships between the PflC and PflD sequences found in this archaeon and A. fulgidus (Figure 4). Recently, the hyperthermophilic archaeon Thermococcus sibiricus MM 739, which was isolated from a well of never-flooded oil-bearing Jurassic horizon of a high-temperature oil reservoir (Western Siberia) (Miroshnichenko et al., 2001), was also shown to grow on hexadecane but not on toluene (Mardanov et al., 2009). Although the process of anaerobic alkane oxidation has not been fully elucidated by this study, it is noteworthy that a gene (Tsib_0631) homologous to pflD has been highlighted in the genome of this archaeon (Mardanov et al., 2009). Our further investigations suggest that the product of this gene could be more closely related to A. fulgidus and AssA/BssA than to E. coli or H. influenzae PFL.
Based on the biochemical and genomic data, the benzylsuccinate synthase from T. aromatica has been proposed to be a heterotrimer composed of a large catalytic subunit (α) and two small subunits (β and γ), the functions of which are still unknown (Leuthner et al., 1998). From genomic analysis, the same organization has been proposed for the alkylsuccinate synthase Mas from Azoarcus sp. HxN1 (Grundmann et al., 2008), whereas for alkylsuccinate synthase Ass from D. alkenivorans AK-01, this trimeric structure is unclear (Callaghan et al., 2008). In A. fulgidus, we were unable to identify homologs of the masC/bssC or masE/bssB genes that would encode the β and γ small subunits of the corresponding trimeric enzymes. It is to be noted that PflD of A. fulgidus, which is encoded by the pflD gene, is likely a homotetrameric enzyme, as shown from previous three-dimensional analysis (Lehtiö et al., 2006). In addition to the genes encoding the catalytic and activating enzymes, the gene clusters encoding alkyl- or benzylsuccinate synthases also include at least one gene encoding proteins involved in the β-oxidation of fatty acids (Callaghan et al., 2008; Grundmann et al., 2008). In the case of A. fulgidus, the pflD gene cluster does not include a gene involved in the β-oxidation pathway. However, the genome of this archaeon shows that genes of the β-oxidation pathways are highly redundant with the presence of 57 β-oxidation genes, suggesting that this organism may degrade a variety of hydrocarbons and organic acids (Klenk et al., 1997).
Our present work further supports the hypothesis that hydrocarbon activation in A. fulgidus VC-16 may occur through addition to fumarate, but this does not preclude the possibility that this archaeon could operate other alkane activation mechanisms as already reported in case of the denitrifying strain EbN1 oxidizing monoaromatic hydrocarbons (Champion et al., 1999).
Accession codes
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402.
Beeder J, Nilsen RK, Rosnes JT, Torsvik T, Lien T . (1994). Archaeoglobus fulgidus isolated from hot North Sea oil field waters. Appl Environ Microbiol 60: 1227–1231.
Beller HR, Spormann AM . (1998). Analysis of the novel benzylsuccinate synthase reaction for anaerobic toluene activation based on structural studies of the product. J Bacteriol 180: 5454–5457.
Boll M, Fuchs G, Heider J . (2002). Anaerobic oxidation of aromatic compounds and hydrocarbons. Curr Opin Chem Biol 6: 604–611.
Boll M, Heider J . (2010). Anaerobic degradation of hydrocarbons: Mechanisms of C-H-bond activation in the absence of oxygen. In: Timmis KN, (ed). Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, Heidelberg, pp 1011–1024.
Brezovsky J, Chovancova E, Gora A, Pavelka A, Biedermannova L, Damborsky J . (2012). Software tools for identification, visualization and analysis of protein tunnels and channels. Biotechnol Adv 31: 38–49.
Brochier-Armanet C, Forterre P, Gribaldo S . (2011). Phylogeny and evolution of the Archaea: one hundred genomes later. Curr Opin Microbiol 14: 274–281.
Brooks BR, Brooks CL 3rd, Mackerell AD Jr, Nilsson L, Petrella RJ, Roux B et al. (2009). CHARMM: the biomolecular simulation program. J Comput Chem 30: 1545–1614.
Callaghan AV . (2013). Enzymes involved in the anaerobic oxidation of n-alkanes: from methane to long-chain paraffins. Front Microbiol 4: 89.
Callaghan AV, Davidova IA, Savage-Ashlock K, Parisi VA, Gieg LM, Suflita JM et al. (2010). Diversity of benzyl- and alkylsuccinate synthase genes in hydrocarbon-impacted environments and enrichment cultures. Environ Sci Technol 44: 7287–7294.
Callaghan AV, Gieg LM, Kropp KG, Suflita JM, Young LY . (2006). Comparison of mechanisms of alkane metabolism under sulfate-reducing conditions among two bacterial isolates and a bacterial consortium. Appl Environ Microbiol 72: 4274–4282.
Callaghan AV, Morris BEL, Pereira IAC, McInerney MJ, Austin RN, Groves JT et al. (2012). The genome sequence of Desulfatibacillum alkenivorans AK-01: a blueprint for anaerobic alkane oxidation. Environ Microbiol 14: 101–113.
Callaghan AV, Wawrik B, Chadhain SMN, Young LY, Zylstra GJ . (2008). Anaerobic alkane-degrading strain AK-01 contains two alkylsuccinate synthase genes. Biochem Biophys Res Commun 366: 142–148.
Champion KM, Zengler K, Rabus R . (1999). Anaerobic degradation of ethylbenzene and toluene in denitrifying strain EbN1 proceeds via independent substrate-induced pathways. J Molec Microbiol Biotechnol 1: 157–164.
Chen C-I, Taylor RT . (1997). Thermophilic biodegradation of BTEX by two consortia of anaerobic bacteria. Appl Microbiol Biotechnol 48: 121–128.
Chovancova E, Pavelka A, Benes P, Strnad O, Brezovsky J, Kozlikova B et al. (2012). CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput Biol 8: e1002708.
Cline JD . (1969). Spectrophotometric determination of hydrogen sulfide in natural waters. Limnol Oceanogr 14: 454–458.
Coates JD, Woodward J, Philip P, Lovley DR . (1997). Anaerobic degradation of polycyclic aromatic hydrocarbons and alkanes in petroleum-contaminated marine harbor sediments. Appl Environ Microbiol 63: 3589–3593.
Coschigano PW, Wehrman TS, Young LY . (1998). Identification and analysis of genes involved in anaerobic toluene metabolism by strain T1: putative role of a glycine free radical. Appl Environ Microbiol 64: 1650–1656.
Cravo-Laureau C, Grossi V, Raphel D, Matheron R, Hirschler-Réa A . (2005). Anaerobic n-alkane metabolism by sulfate-reducing bacterium, Desulfatibacillum aliphaticivorans strain CV2803T. Appl Environ Microbiol 71: 3458–3467.
Criscuolo A, Gribaldo S . (2010). BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol Biol 10: 210.
DeLano WL . (2005). The case for open-source software in drug discovery. Drug Discov Today 10: 213–217.
Fardeau ML, Goulhen F, Bruschi M, Khelifi N, Cayol JL, Ignatiadis I et al. (2009). Archaeoglobus fulgidus and Thermotoga elfii, thermophilic isolates from deep geothermal water of the Paris basin. Geomicrobiol J 26: 119–130.
Gouy M, Guindon S, Gascuel O . (2010). SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27: 221–224.
Grossi V, Cravo-Laureau C, Guyoneaud R, Ranchou-Peyruse A, Hirschler-Réa A . (2008). Metabolism of n-alkanes and n-alkenes by anaerobic bacteria: a summary. Org Geochem 39: 1197–1203.
Grundmann O, Behrends A, Rabus R, Amann J, Halder T, Heider J et al. (2008). Genes encoding the candidate enzyme for anaerobic activation of n-alkanes in the denitrifying bacterium, strain HxN1. Environ Microbiol 10: 376–385.
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O . (2010). New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59: 307–321.
Heider J . (2007). Adding handles to unhandy substrates: anaerobic hydrocarbon activation mechanisms. Curr Opin Chem Biol 11: 188–194.
Ho BK, Gruswitz F . (2008). HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct Biol 8: 49.
Holmes DE, Risso C, Smith JA, Lovley DR . (2011). Anaerobic oxidation of benzene by the hyperthermophilic archaeon Ferroglobus placidus. Appl Environ Microbiol 77: 5926–5933.
Jarling R, Sadeghi M, Drozdowska M, Lahme S, Buckel W, Rabus R et al. (2012). Stereochemical investigations reveal the mechanism of the bacterial activation of n-alkanes without oxygen. Angew Chem Int Ed 51: 1334–1338.
Jones DM, Head IM, Gray ND, Adams JJ, Rowan AK, Aitken CM et al. (2008). Crude-oil biodegradation via methanogenesis in subsurface petroleum reservoirs. Nature 451: 176–180.
Katoh K, Standley DM . (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30: 772–780.
Khelifi N, Grossi V, Dolla A, Hamdi M, Tholozan J-L, Ollivier B et al. (2010). Anaerobic oxidation of fatty acids and alkenes by the hyperthemophilic sulfate-reducing archaeon, Archaeoglobus fulgidus. Appl Environ Microbiol 76: 3057–3060.
Klenk H-P, Clayton RA, Tomb J-F, White O, Nelson KE, Ketchum KA et al. (1997). The complete genome sequence of the hyperthermophilic, sulphate-reducing archaeon Archaeoglobus fulgidus. Nature 390: 364–370.
Knappe J, Elbert S, Frey M, Wagner AFV . (1993). Pyruvate formate lyase mechanism involving the protein-based glycyl radical. Biochem Soc Trans 21: 731–734.
Kniemeyer O, Musat F, Sievert SM, Knittel K, Wilkes H, Blumenberg M et al. (2007). Anaerobic oxidation of short-chain hydrocarbons by marine sulphate-reducing bacteria. Nature 449: 898–901.
Konn C, Charlou JL, Donval JP, Holm NG, Dehairs F, Bouillon S . (2009). Hydrocarbons and oxidized organic compounds in hydrothermal fluids from Rainbow and Lost City ultramafic-hosted vents. Chem Geol 258: 299–314.
Kropp KG, Davidova IA, Suflita JM . (2000). Anaerobic oxidation of n-dodecane by an addition reaction in a sulfate-reducing bacterial enrichment culture. Appl Environ Microbiol 66: 5393–5398.
L’Haridon S, Reysenbacht AL, Glenat P, Prieur D, Jeanthon C . (1995). Hot subterranean biosphere in a continental oil reservoir. Nature 377: 223–224.
Le SQ, Gascuel O . (2008). An improved general amino acid replacement matrix. Mol Biol Evol 25: 1307–1320.
Lehtiö L, Grossmann JG, Kokona B, Fairman R, Goldman A . (2006). Crystal structure of a glycyl radical enzyme from Archaeoglobus fulgidus. J Mol Biol 357: 221–235.
Leuthner B, Leutwein C, Schulz H, Hörth P, Haehnel W, Schiltz E et al. (1998). Biochemical and genetic characterization of benzylsuccinate synthase from Thauera aromatica: a new glycyl radical enzyme catalysing the first step in anaerobic toluene metabolism. Mol Microbiol 28: 615–628.
Liang J, Edelsbrunner H, Woodward C . (1998). Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci 7: 1884–1897.
MacKerell AD, Feig M, Brooks CL . (2004). Improved treatment of the protein backbone in empirical force fields. J Amer Chem Soc 126: 698–699.
Magot M, Ollivier B, Patel BKC . (2000). Microbiology of petroleum reservoirs. Antonie van Leeuw 77: 103–116.
Mardanov AV, Ravin NV, Svetlitchnyi VA, Beletsky AV, Miroshnichenko ML, Bonch-Osmolovskaya EA et al. (2009). Metabolic versatility and indigenous origin of the archaeon Thermococcus sibiricus, isolated from a Siberian oil reservoir, as revealed by genome analysis. Appl Environ Microbiol 75: 4580–4588.
Mbadinga SM, Li K-P, Zhou L, Wang L-Y, Yang S-Z, Liu J-F et al. (2012). Analysis of alkane-dependent methanogenic community derived from production water of a high-temperature petroleum reservoir. Appl Microbiol Biotechnol 96: 1–12.
Miroshnichenko M, Hippe H, Stackebrandt E, Kostrikina N, Chernyh N, Jeanthon C et al. (2001). Isolation and characterization of Thermococcus sibiricus sp. nov. from a Western Siberia high-temperature oil reservoir. Extremophiles 5: 85–91.
Ollivier B, Alazard D . (2010). The oil reservoir ecosystem. In: Timmis KN (ed). Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, Heidelberg, pp 2261–2269.
Petrek M, Otyepka M, Banás P, Kosinová P, Koca J, Damborský J . (2006). CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinformatics 7: 316.
Pfaffl MW . (2001). A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29: 2002–2007.
Price MN, Dehal PS, Arkin AP . (2010). FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS One 5: e9490.
Rabus R, Wilkes H, Behrends A, Armstroff A, Fischer T, Pierik AJ et al. (2001). Anaerobic initial reaction of n-alkanes in denitrifying bacterium: evidence for (1-methylpentyl)succinate as initial product and for involvement of an organic radical in n-hexane metabolism. J Bacteriol 183: 1707–1715.
Rojo F . (2009). Degradation of alkanes by bacteria. Environ Microbiol 11: 2477–2490.
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S et al. (2012). MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61: 539–542.
Rueter P, Rabus R, Wilkes H, Aeckersberg F, Rainey FA, Jannash HW et al. (1994). Anaerobic oxidation of hydrocarbons in crude oil by new types of sulphate-reducing bacteria. Nature 372: 455–457.
Sofia HJ, Chen G, Hetzler BG, Reyes-Spindola JF, Miller NE . (2001). Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res 29: 1097–1106.
Stetter KO . (1988). Archaeoglobus fulgidus gen. nov., sp. nov.: a new taxon of extremely thermophilic archeabacteria. System Appl Microbiol 10: 172–173.
Stetter KO, Huber R, Blochl E, Kurr M, Eden RD, Fielder M et al. (1993). Hyperthermophilic archaea are thriving in deep North Sea and Alaskan oil reservoirs. Nature 365: 743–745.
Stetter KO, Lauerer G, Thomm M, Neuner A . (1987). Isolation of extremely thermophilic sulfate reducers - evidence for a novel branch of Archaebacteria. Science 236: 822–824.
Thauer RK, Jungermann K, Decker K . (1977). Energy conservation in chemotrophic anaerobic bacteria. Bacteriol Rev 41: 100–180.
Widdel F, Knittel K, Galushko A . (2010). Anaerobic hydrocarbon-degrading microorganisms: an overview. In: Timmis KN (ed). Handbook of Hydrocarbon and Lipid Microbiology. Springer-Verlag: Berlin, Heidelberg, pp 1998–2021.
Wolin EA, Wolin MJ, Wolfe RS . (1963). Formation of methane by bacterial extracts. J Biol Chem 238: 2882–2886.
Zedelius J, Rabus R, Grundmann O, Werner I, Brodkorb D, Schreiber F et al. (2011). Alkane degradation under anoxic conditions by a nitrate-reducing bacterium with possible involvement of the electron acceptor in substrate activation. Environ Microbiol Rep 3: 125–135.
Zellner G, Stackebrandt E, Kneifel H, Messner P, Sleytr UB, Conway De Macario E et al. (1989). Isolation and characterization of a thermophilic, sulfate reducing Archaebacterium, Archaeoglobus fulgidus strain Z. System Appl Microbiol 11: 151–160.
Acknowledgements
This work was partially supported by a French CNRS-INSU grant to V. Grossi (BIOHYDEX EC2CO-Microbien project). C. Brochier-Armanet is supported by the Investissement d'Avenir grant Ancestrome (ANR-10-BINF-01-01) and is a member of the Institut Universitaire de France. We would like to thank the anonymous reviewers for allowing an opportunity to improve this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Khelifi, N., Amin Ali, O., Roche, P. et al. Anaerobic oxidation of long-chain n-alkanes by the hyperthermophilic sulfate-reducing archaeon, Archaeoglobus fulgidus. ISME J 8, 2153–2166 (2014). https://doi.org/10.1038/ismej.2014.58
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.58
Keywords
This article is cited by
-
Newly discovered Asgard archaea Hermodarchaeota potentially degrade alkanes and aromatics via alkyl/benzyl-succinate synthase and benzoyl-CoA pathway
The ISME Journal (2021)
-
A review on microbial diversity and genetic markers involved in methanogenic degradation of hydrocarbons: futuristic prospects of biofuel recovery from contaminated regions
Environmental Science and Pollution Research (2021)
-
Long-chain n-alkane biodegradation coupling to methane production in an enriched culture from production water of a high-temperature oil reservoir
AMB Express (2020)
-
Deciphering the archaeal communities in tree rhizosphere of the Qinghai-Tibetan plateau
BMC Microbiology (2020)
-
Novel clostridial lineages recovered from metagenomes of a hot oil reservoir
Scientific Reports (2020)