Abstract
Fabry disease is an inborn error of glycolipid catabolism resulting from lesions in the gene encoding α-galactosidase (GLA). To elucidate the basis of Fabry disease, we constructed structural models of mutant GLAs responsible for the disease and calculated indexes, i.e., the numbers of atoms affected in the main chain and side chain of each mutant GLA, the root-mean-square distance values, and the solvent-accessible surface-area values, based on 212 Fabry amino acid substitutions previously reported (196 classic and 16 variant). As two therapeutic options, enzyme replacement and enzyme enhancement, are now available for this disease, proper prediction of the natural outcome and therapeutic efficiency based on the molecular evidence for individual cases are critical for patients’ quality of life. Our results revealed that structural changes in the classic Fabry group were generally large and tended to be in the core region of a protein or located in the functionally important region, including the active-site pocket. On the other hand, structural changes in the variant Fabry group were small or localized on the surface of the molecule far away from the active site. We focused on structural changes due to amino acid substitutions for which substrate analogues are effective for improving the stability or transportation of mutant GLAs, and the results of the study revealed that they are small or localized on the molecular surface, regardless of the phenotype. Coloring of affected atoms based on distances between wild type and mutant ones clearly showed the characteristic structural changes in the GLA protein geographically and subquantitatively. Structural investigation is useful for elucidation of the basis of Fabry disease and predicting disease outcome.
Similar content being viewed by others
Introduction
Lysosomal α-galactosidase (GLA, EC3.2.1.22) catalyzes hydrolysis of terminal α-d-galactosyl residues of glycoconjugates, predominantly globotriaosylceramide (GL-3), in lysosomes. The enzyme is encoded by the GLA gene on the long arm of the X-chromosome and is synthesized on endoplasmic reticulum (ER)-bound ribosomes as a precursor form, which consists of 429 amino acid residues. Then, the enzyme is translocated into the lumen of the ER, with subsequent cleavage of the signal peptide consisting of 31 residues. Then, the enzyme is modified in the ER by the addition of N-linked oligosaccharides. The oligosaccharides are then trimmed in the ER, and the enzyme is transferred to the Golgi apparatus, where further modification of sugar chains and the addition of mannose 6-phosphate residues occur. The enzyme, having mannose 6-phosphate residues at the nonreducing ends of sugar chains, is transported to endosomes via mannose 6-phosphate receptors. Subsequently, the enzyme is transported to lysosomes, where it exerts its function as a mature form consisting of 398 residues. The native GLA from humans is thought to have a homodimeric structure.
A genetic defect causes progressive accumulation of GL-3, which results in Fabry disease (MIM 301500) (Desnick et al. 2001). This disease exhibits a wide clinical spectrum. Patients with the classic form having no GLA activity develop systemic manifestations, including pain in peripheral extremities, hypohidrosis, angiokeratoma, corneal clouding, renal failure, and cardio- and cerebrovascular disorders. On the other hand, there are also variant Fabry-disease patients with residual GLA activity and milder clinical manifestations, sometimes limited to heart disorders. So far, more than 500 genetic mutations causing Fabry disease have been reported (Desnick et al. 2001). Among them, gross alterations of the GLA gene have been identified in patients with the classic form, but missense mutations comprising the majority of mutations have been found in both classic and variant forms.
Recombinant GLAs produced in Chinese hamster ovary cells and human fibroblasts have been developed and are clinically available for enzyme replacement therapy for Fabry disease (Eng et al. 2001a, b; Schiffmann et al. 2000). Recently, another potential approach for treating Fabry disease was developed, and a clinical trial has been performed. This enzyme enhancement therapy is based on the ability of substrate analogues including galactose and 1-deoxygalactonojirimycin to improve the stability or transportation of mutant GLAs in cells, but the therapy is only efficient in a limited group of patients having specific missense mutations (Frustaci et al. 2001; Yam et al. 2006; Fan and Ishii 2007). As a high incidence of variant Fabry disease has been revealed by newborn screening (Spada et al. 2006), prediction of the clinical outcome of the disease is becoming more and more important to determine a proper schedule for treating the disease.
Previously, we built structural models of mutant GLAs resulting from 161 missense mutations by means of homology modeling with SYBYL/BIOPOLYMER (TRIPOS, St Louis, MO, USA) and examined the correlation between structural changes in GLAs and clinical and biochemical phenotypes (Matsuzawa et al. 2005).
Recently, we developed a structural analysis system for mutant proteins involving molecular modeling software, TINKER, developed by Ponder et al. (Department of Biochemistry and Molecular Biophysics, Washington University) (Ren and Ponder 2003), which is available worldwide. We applied it to investigations on lysosomal diseases including mucopolysaccharidosis type 6 (Saito et al. 2008), mucopolysaccharidosis type 1 (Sugawara et al. 2008), and Tay-Sachs disease (Ohno et al. 2008). We believe that the standardization of a structural analysis method will enable us to compare the results for different genetic disorders, which will provide us with a deeper insight into the basis of genetic disorders. Furthermore, because TINKER is free software, other researchers can easily conduct follow-up studies.
In this study, we conducted further structural investigation of Fabry disease using the same structural analysis system. We increased the number of Fabry patients for the analysis and examined structural changes in GLAs due to 212 amino acid substitutions by determining the number of atoms affected, the root-mean-square distance (RMSD), and the solvent-accessible surface area (ASA). Then, we paid attention to mutant GLAs for which substrate analogues are effective for stabilization or transportation to lysosomes and characterized their structural changes by coloring the affected atoms.
Materials and methods
Amino acid substitutions causing classic and variant Fabry disease
In this study, we analyzed 212 missense mutations (196 classic and 16 variant) responsible for Fabry disease. Amino acid substitutions, phenotypes, and references are summarized in Table 1.
Development of a structural analysis system for mutant proteins
We developed a structural analysis system for mutant proteins to examine their structural changes responsible for genetic diseases from various viewpoints. This system comprises six stages: (1) modeling mutant proteins, (2) determining the number of atoms affected by amino acid substitutions, (3) determining the RMSD values of all atoms in the mutant proteins, (4) determining ASA values of amino acid residues in the mutant proteins, (5) statistical analysis, and (6) coloring the atoms affected in the mutant proteins based on the differences between wild-type and mutant ones. Then, we applied the system to elucidation of the basis of Fabry disease.
Structural modeling of mutant GLAs responsible for Fabry disease and determination of the number of atoms affected by amino acid substitutions
Structural modeling of mutant GLAs was performed using molecular modeling software TINKER (Kundrot et al. 1991; Dudek and Ponder 1995; Kong and Ponder 1997; Pappu et al. 1998; Ren and Ponder 2003). The crystal structure of human GLA (Garman and Garboczi 2004) (PDB: 1R46) was used as a template, and energy minimization was performed. The root-mean-square gradient value was set at 0.05 kcal/mol Å. Each mutant model was then superimposed on the wild-type GLA structure based on Cα atoms by the least-square-mean fitting method (Kabsch 1976, 1978; Sakuraba et al. 2000, 2004). In this study, we defined that the structure was affected by an amino acid substitution when the position of an atom in a mutant differed from that in the wild type by more than the cutoff distance (0.15 Å) based on total RMSD, as described previously (Matsuzawa et al. 2005). Then, we determined the numbers of atoms affected in GLA main chain and side chain.
Determination of RMSD values of all atoms in mutant GLAs
RMSD values of all atoms in mutant GLAs were determined according to Weiner’s method (Weiner et al. 1984) to predict the degrees of GLA structural changes, and the average RMSD values for the classic and variant Fabry groups were determined and compared with each other, as described previously (Sugawara et al. 2008).
Determination of ASA values of amino acid residues in mutant GLAs
To predict the position of a substituted amino acid residue in the GLA molecule, the ASA value of each residue in the wild-type GLA was calculated using ACCESS (McDonald and Thornton 1994). The average ASA values of the residues for which a substitution had been found in the classic and variant Fabry groups were determined and compared with each other, as described previously (Saito et al. 2008; Sugawara et al. 2008).
Statistical analysis
Statistical analysis to determine differences in the numbers of atoms affected, RMSD values, and ASA values between classic and variant Fabry groups was performed using the F test and then Welch’s t test, it being taken that there was a significant difference if P < 0.05.
Coloring the atoms affected in mutant GLAs for which substrate analogues are effective
To determine the influence of amino acid substitutions geographically and semiquantitatively, coloring the affected atoms in the three-dimensional structure of GLA based on the distances between the wild-type and mutant ones was performed, followed by determination of the numbers of affected atoms, RMSD values, and ASA values. We analyzed amino acid substitutions including E59K, E66Q, M72V, I91T, A97V, R112H, F113L, A156V, L166V, N215S, G260A, Q279E, M296I, M296V, R301Q, R356W, and G373S, for which substrate analogues are effective for stabilization or transportation of mutant enzymes to lysosomes (Okumiya et al. 1995b; Yam et al. 2006; Ishii et al. 2007).
Results
Localization of amino acid substitutions responsible for Fabry disease
According to the crystallographic structure of human GLA (Garman and Garboczi 2004), the enzyme unit comprises two domains: an N-terminal (β/α)8-barrel domain and a C-terminal antiparallel β-sheet domain. The active-site pocket is localized in the C-terminal of the β-sheet of the N-terminal domain. We determined the locations of residues of which amino acid substitutions have been identified in Fabry disease patients in the homodimeric enzyme structure (Fig. 1a). Then, we localized the residues of which amino acid substitutions are responsible for classic (Fig. 1b, c) and variant (Fig. 1d, e) Fabry groups in the GLA subunit to compare them with each other. In the classic Fabry group, amino acid substitutions were distributed all over the enzyme protein molecule, including the active-site pocket. On the other hand, in the variant Fabry group, they were located far from the active-site pocket, and most of them were localized on the molecular surface of the protein.
Localization of amino acid substitutions responsible for Fabry disease in the α-galactosidase (GLA) structure. Secondary structures in GLA are shown as a tube drawing. Locations of amino acid substitutions identified in the classic and variant Fabry groups are shown in yellow and green, respectively. N The N-terminal (β/α)8-barrel domain. C The C-terminal antiparallel β-sheet domain. An arrow indicates the active-site pocket. a Homodimeric GLA structure, classic and variant Fabry groups. b Front view of the GLA subunit, classic Fabry group. c Side view of the GLA subunit, classic Fabry group. d Front view of the GLA subunit, variant Fabry group. e Side view of the GLA subunit, variant Fabry group
Numbers of atoms affected by amino acid substitutions responsible for Fabry disease
We built structural models of the mutant GLAs and calculated the number of atoms affected by the amino acid substitution for each mutant model (Table 1), the results being summarized in Fig. 2.
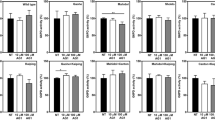
Numbers of atoms in the a main chain and the b side chain of the α-galactosidase (GLA) protein affected by amino acid substitutions. Classic classic Fabry group. Variant variant Fabry group. Mutations located in the active-site pocket are colored red, others being colored black. Boxes indicate mean ± standard errors of mean
The classic Fabry group showed a wide distribution. Averages for the affected atoms in the main chain and side chain were 108 and 130, respectively. In particular, regarding the former, 108 of the 196 classic cases (56%) had 50 atoms or more affected. There were 36 amino acid substitutions located in the active-site pocket, all of which cause the classic Fabry phenotype. These cases are colored red for their differentiation from other cases, which are colored black.
In contrast, the number of affected atoms in the variant Fabry group was low, and the distribution was narrower. These cases are colored black. Averages of the affected atoms in the main chain and the side chain were 18 and 21, respectively. In particular, regarding the main-chain atoms, 14 of the 16 variant Fabry cases (88%) had 49 atoms or less affected.
The F test showed that the distribution exhibited unequal variance (P < 0.05) between the classic and variant Fabry groups, and thus, Welch’s t test was performed. Results revealed that there were significant differences in the numbers of affected atoms in both the main chain and side chain between the two groups (P < 0.05).
RMSD values for amino acid substitutions responsible for Fabry disease
The RMSD values for the classic and variant Fabry groups were determined. Results are shown in Fig. 3. The average RMSD values in the classic and variant Fabry groups were 0.089 and 0.029 Å, respectively. In 115 of the 196 classic Fabry cases (59%), the RMSD value was ≥0.05 Å. On the other hand, it was <0.05 Å in 14 of the 16 variant Fabry cases (88%). Results of the F test followed by Welch’s t test showed that there was a significant difference in the RMSD values between the two groups.
ASA values of amino acid substitutions responsible for Fabry disease
To determine and compare locations of amino acid residues in the GLA molecule associated with the classic (126 residues) and variant (15 residues) Fabry cases, the ASA values of the residues in the wild-type GLA structure were calculated, the results being shown in Fig. 4 (the result for each residue is presented in “Supplementary data No. l”). In the classic Fabry group, the average ASA value for the 126 residues analyzed was 13.3 Å2, 93 of them (74%) being <20 Å2. In the variant Fabry group, the average ASA value for the 15 residues analyzed was 25.1 Å2, eight of them being ≥20 Å2 (53%).
The F test followed by Welch’s t test revealed that P was 0.09. Results suggest that the residues associated with classic Fabry mutations tend to be less solvent-accessible than those associated with variant ones, although this could not be confirmed statistically.
Coloring the affected atoms due to amino acid substitutions for which substrate analogues are effective
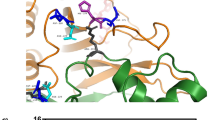
Regarding amino acid substitutions responsible for Fabry disease, we paid attention to 17 mutations for which substrate analogues improved the stability or transportation of mutant GLAs in cells, i.e., E59K (classic), E66Q (variant), M72V (variant), I91T (variant), A97V (classic), R112H (variant), F113L (variant), A156V (classic), L166V (classic), N215S (variant), G260A (classic), Q279E (variant), M296I (variant), M296V (variant), R301Q (variant), R356W (classic), and G373S (classic). Coloring the affected atoms in the three-dimensional GLA structure was performed for these mutations, the results being shown in Fig. 5.
Coloring the atoms in the three-dimensional structure affected by amino acid substitutions for which substrate analogues are effective. The degrees and distributions for E59K, E66Q, M72V, I91T, A97V, R112H, F113L, A156V, L166V, N215S, G260A, Q279E, M296I, M296V, R301Q, R356W, and G373S, for which substrate analogues are effective, are shown. Each atom is colored according to the distance between the atom in the mutant and the corresponding atom in the wild-type structure. The colors of the atoms show the distances as follows: blue <0.15 Å, 0.15 Å ≤ cyan < 0.30 Å, 0.30 Å ≤ green < 0.45 Å, 0.45 Å ≤ yellow < 0.60 Å, 0.60 Å ≤ orange < 0.75 Å, and red ≥ 0.75 Å. Arrows indicate the active-site pocket
Coloring the affected atoms clearly allowed visualization of structural changes. Determining the numbers of atoms affected and RMSD and ASA values confirmed the results. In most mutant GLAs, the predicted structural changes were small (numbers of affected atoms in both the main chain and side chain <50, and RMSD < 0.05: E66Q, M72V, I91T, A97V, F113L, L166V, L215S, G260A, Q279E, M296I, M296V, and G373S), or localized on the molecular surface, although the structural changes were not small (ASA ≥ 20: E59K, R112H, and R301Q), regardless of phenotype. There were only two exceptions, A156V and R356W, and their clinical phenotype was classic. None of the amino acid substitutions for which substrate analogues are effective caused any structural changes in the active site.
Discussion
Considering the results of newborn screening, the incidence of Fabry disease is unexpectedly high (1 in 3,000–4,000 male newborns), especially the variant form (Spada et al. 2006). It is very important to examine the structural changes in the enzyme protein responsible for the different phenotypes to elucidate the basis of Fabry disease and to predict disease outcome. Garman and Garboczi calculated the side-chain-accessible surface area and revealed that the residues involved in Fabry mutations tend to be less solvent-accessible than the typical residues and that most of them lead to disruption of the hydrophobic core of the protein (Garman and Garboczi 2004; Garman 2007). However, there is little structural information on defective GLA proteins, although a large number of gene mutations responsible for Fabry disease have been reported so far.
In this study, we constructed structural models of Fabry mutant GLAs and examined their structural changes from various aspects by determining the number of atoms affected, as well as RMSD and ASA values, using our structural analysis system. The results revealed that structural changes in the classic Fabry group are generally large and tend to be in the core region of the protein. About 85% (116/196) of the amino acid substitutions leading to classic Fabry disease satisfied one of the conditions given below; number of affected atoms in the main chain ≥50, number in the side chain ≥50, RMSD ≥ 0.05 Å, or ASA < 20 Å2. They seriously affected protein folding or intracellular transport, leading to a deficiency of enzyme activity. All amino acid substitutions causing structural changes of the active-site pocket resulted in the classic phenotype. In such cases, structural changes would seriously affect expression of GLA activity, even if the number of influenced atoms is relatively small.
On the other hand, the predicted structural changes in GLA are generally small or localized on the surface of the molecule far away from the active site in the variant Fabry group. In such cases, a small amount of enzyme having GLA activity would be protected from the ER’s quality control system and transported to lysosomes, resulting in residual enzyme activity.
The number of affected atoms calculated using TINKER differs from that calculated using SYBYL/BIOPOLYMER. This is not surprising, because the minimized structure depends on the minimization algorithm, force field, and computational implementation. As shown in supplementary data No. 2 and No. 3, the number of affected atoms calculated using TINKER was generally larger than that using SYBYL/BIOPOLYMER. The mutant model obtained with TINKER was well optimized compared with that with SYBYL/BIOPOLYMER, which indicates that the mutant model constructed in this study is improved compared with the previous one (Matsuzawa et al. 2005). However, the supplementary data suggest that the number of affected atoms calculated using SYBYL/BIOPOLYMER was correlated with that using TINKER. Therefore, the discussion in the previous study is thought to remain correct.
Furthermore, we focused on the structural changes due to amino acid substitutions for which substrate analogues are effective and examined them by determining the affected atoms, RMSD values, and ASA values, followed by coloring the affected atoms. Results revealed that they cause small structural changes in GLA or are localized on the molecular surface, except for a couple of exceptions. None of them affected the active site. These results suggest that binding of a substrate analogue to a mutant enzyme protein in which a small structural change has occurred on the surface of the molecule reduces its folding defect and increases its stability in cells. Previously, we expressed mutant GLAs including M72V, L156V, L166V, Q279E, and R301Q in COS-1 cells and Sf9 cells and examined their biochemical characteristics (Ishii et al. 1993; Okumiya et al. 1995a, 1998; Kase et al. 2000). The expressed products had GLA activity, but they were unstable and easily lost their activity in vitro, suggesting that their structural changes are located far from the active site and that the degree of the changes is not so large. The results are well correlated with those of the structural analysis performed this time.
In conclusion, we investigated the structural changes in GLA responsible for Fabry disease. Results showed a correlation between the three-dimensional structural changes and clinical phenotypes, and they also revealed the characteristics of the structural changes in mutant enzyme proteins for which substrate analogues are effective. Structural investigation is useful for elucidation of the basis of Fabry disease, and it will increase our ability to determine a proper therapeutic schedule for this disease.
References
Ashley GA, Shabbeer J, Yasuda M, Eng CM, Desnick RJ (2001) Fabry disease: twenty novel alpha-galactosidase A mutations causing the classical phenotype. J Hum Genet 46:192–196
Ashton-Prolla P, Tong B, Shabbeer J, Astrin KH, Eng CM, Desnick RJ (2000) Fabry disease: twenty-two novel mutations in the alpha-galactosidase A gene and genotype/phenotype correlations in severely and mildly affected hemizygotes and heterozygotes. J Investig Med 48:227–235
Bernstein HS, Bishop DF, Astrin KH, Kornreich R, Eng CM, Sakuraba H, Desnick RJ (1989) Fabry disease: six gene rearrangements and an exonic point mutation in the alpha-galactosidase gene. J Clin Invest 83:1390–1399
Blanch LC, Meaney C, Morris CP (1996) A sensitive mutation screening strategy for Fabry disease: detection of nine mutations in the alpha-galactosidase A gene. Hum Mutat 8:38–43
Blaydon D, Hill J, Winchester B (2001) Fabry disease: 20 novel GLA mutations in 35 families. Hum Mutat 18:459
Calado J, Dickson J, Rueff J (2004) Human gene mutations. Hum Genet 115:347
Cooper A, Cooper JA, Wraith JE (2000) Human gene mutations. Hum Genet 107:535–536
Davies JP, Winchester BG, Malcolm S (1993) Mutation analysis in patients with the typical form of Anderson-Fabry disease. Hum Mol Genet 2:1051–1053
Davies J, Christomanou H, Winchester B, Malcolm S (1994) Detection of 8 new mutations in the alpha-galactosidase A gene in Fabry disease. Hum Mol Genet 3:667–669
Davies JP, Eng CM, Hill JA, Malcolm S, MacDermot K, Winchester B, Desnick RJ (1996) Fabry disease: fourteen alpha-galactosidase A mutations in unrelated families from the United Kingdom and other European countries. Eur J Hum Genet 4:219–224
Desnick RJ, Ioannou YA, Eng CM (2001) Alpha-galactosidase A deficiency: Fabry disease. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic and molecular bases of inherited disease, 8th edn. McGraw-Hill, New York, pp 3733–3774
Dobrovolny R, Dvorakova L, Ledvinova J, Magage S, Bultas J, Lubanda JC, Elleder M, Karetova D, Pavlikova M, Hrebicek M (2005) Relationship between X-inactivation and clinical involvement in Fabry heterozygotes. Eleven novel mutations in the alpha-galactosidase A gene in the Czech and Slovak population. J Mol Med 83:647–654
Dudek MJ, Ponder JW (1995) Accurate modeling of the intramolecular electrostatic energy of proteins. J Comput Chem 16:791–816
Eng CM, Resnick-Silverman LA, Niehaus DJ, Astrin KH, Desnick RJ (1993) Nature and frequency of mutations in the alpha-galactosidase A gene that cause Fabry disease. Am J Hum Genet 53:1186–1197
Eng CM, Niehaus DJ, Enriquez AL, Burgert TS, Ludman MD, Desnick RJ (1994) Fabry disease: twenty-three mutations including sense and antisense CpG alterations and identification of a deletional hot-spot in the alpha-galactosidase A gene. Hum Mol Genet 3:1795–1799
Eng CM, Ashley GA, Burgert TS, Enriquez AL, D’Souza M, Desnick RJ (1997) Fabry disease: thirty-five mutations in the alpha-galactosidase A gene in patients with classic and variant phenotypes. Mol Med 3:174–182
Eng CM, Banikazemi M, Gordon RE, Goldman M, Phelps R, Kim L, Gass A, Winston J, Dikman S, Fallon JT, Grodie S, Stacy CB, Mehta D, Parsons R, Norton K, O’Callaghan M, Desnick RJ (2001a) A phase 1/2 clinical trial of enzyme replacement in Fabry disease: pharmacokinetic, substrate clearance, and safety studies. Am J Hum Genet 68:711–722
Eng CM, Guffon N, Wilcox WR, Germain DP, Lee P, Waldek S, Caplan L, Linthorst GE, Desnick RJ (2001b) Safety and efficacy of recombinant human alpha-galactosidase A replacement therapy in Fabry’s disease. N Engl J Med 345:9–16
Fan J-Q, Ishii S (2007) Active-site specific chaperone for Fabry disease; Yin and Yang of enzyme inhibitors. FEBS J 274:4962–4971
Frustaci A, Chimenti C, Ricci R, Natale L, Russo MA, Pieroni M, Eng CM, Desnick RJ (2001) Improvement in cardiac function in the cardiac variant of Fabry’s disease with galactose-infusion therapy. N Engl J Med 345:25–32
Garman SC, Garboczi DN (2004) The molecular defect leading to Fabry disease: structure of human alpha-galactosidase. J Mol Biol 337:319–335
Garman SC (2007) Structure-function relationships in alpha-galactosidase A. Acta Paediatr Suppl 96:6–16
Germain D, Biasotto M, Tosi M, Meo T, Kahn A, Poenaru L (1996) Fluorescence-assisted mismatch analysis (FAMA) for exhaustive screening of the alpha-galactosidase A gene and detection of carriers in Fabry disease. Hum Genet 98:719–726
Germain DP, Salard D, Fellmann F, Azibi K, Caillaud C, Bernard MC, Poenaru L (2001) Identification of a novel de novo mutation (G373D) in the alpha-galactosidase A gene (GLA) in a patient affected with Fabry disease. Hum Mutat 17:353
Germain DP, Shabbeer J, Cotigny S, Desnick RJ (2002) Fabry disease: twenty novel alpha-galactosidase A mutations and genotype-phenotype correlations in classical and variant phenotypes. Mol Med 8:306–312
Guffon N, Froissart R, Chevalier-Porst F, Maire I (1998) Mutation analysis in 11 French patients with Fabry disease. Hum Mutat Suppl 1:S288–S290
Iga MI, Okayama A, Matsuyama M, Sasaki T, Murai K, Hashida S, Morishita K, Tsubouchi H (2001) Human gene mutations. Hum Genet 109:126
Ishii S, Sakuraba H, Suzuki Y (1992) Point mutations in the upstream region of the alpha-galactosidase A gene exon 6 in an atypical variant of Fabry disease. Hum Genet 89:29–32
Ishii S, Kase R, Sakuraba H, Suzuki Y (1993) Characterization of mutant alpha-galactosidase gene product for the late-onset cardiac form of Fabry disease. Biochem Biophys Res Commun 197:1585–1589
Ishii S, Chang HH, Kawasaki K, Yasuda K, Wu HL, Garman SC, Fan JQ (2007) Mutant alpha-galactosidase A enzymes identified in Fabry disease patients with residual enzyme activity: biochemical characterization and restoration of normal intracellular processing by 1-deoxygalactonojirimycin. Biochem J 406:285–295
Kabsch W (1976) A solution for the best rotation to relate two sets of vectors. Acta Crystallogr A 32:827
Kabsch W (1978) A discussion of the solution for the best rotation to relate two sets of vectors. Acta Crystallogr A 34:922–923
Kase R, Bierfreund U, Klein A, Kolter T, Utsumi K, Itoh K, Sandhoff K, Sakuraba H (2000) Characterization of two alpha-galactosidase mutants (Q279E and R301Q) found in an atypical variant of Fabry disease. Biochem Biophys Res Commun 1501:227–235
Kimura K, Sato-Matsumura KC, Nakamura H, Onodera Y, Morita K, Enami N, Shougase T, Ohsaki T, Kato M, Takahashi T, Yamaguchi Y, Shimizu H (2002) A novel A97P amino acid substitution in alpha-galactosidase A leads to a classical Fabry disease with cardiac manifestations. Br J Dermatol 147:545–548
Koide T, Ishiura M, Iwai K, Inoue M, Kaneda Y, Okada Y, Uchida T (1990) A case of Fabry’s disease in a patient with no alpha-galactosidase A activity caused by a single amino acid substitution of Pro-40 by Ser. FEBS Lett 259:353–356
Kong MJ, Ponder JW (1997) Reaction field methods for off-center multipoles. J Chem Phys 107:481–492
Kotanko P, Kramar R, Devrnja D, Paschke E, Voigtländer T, Auinger M, Pagliardini S, Spada M, Demmelbauer K, Lorenz M, Hauser AC, Kofler HJ, Lhotta K, Neyer U, Pronai W, Wallner M, Wieser C, Wiesholzer M, Zodl H, Födinger M, Sunder-Plassmann G (2004) Results of a nationwide screening for Anderson-Fabry disease among dialysis patients. J Am Soc Nephrol 15:1323–1329
Kundrot CE, Ponder JW, Richards FM (1991) Algorithms for calculating excluded volume and its derivative as a function of molecular conformation and their use in energy minimization. J Comput Chem 12:402–409
Lai L, O’Meara M, Lien YH (2001) Human gene mutations. Hum Genet 109:469
Lee JK, Kim GH, Kim JS, Kim KK, Lee MC, Yoo HW (2000) Identification of four novel mutations in five unrelated Korean families with Fabry disease. Clin Genet 58:228–233
Matsuzawa F, Aikawa S, Doi H, Okumiya T, Sakuraba H (2005) Fabry disease: correlation between structural changes in alpha-galactosidase, and clinical and biochemical phenotype. Hum Genet 117:317–328
McDonald IK, Thornton JM (1994) Satisfying hydrogen bonding potential in proteins. J Mol Biol 238:777–793
Miyazaki T, Kajita M, Ohmori S, Mizutani N, Niwa T, Murata Y, Seo H (1998) A novel mutation (E358K) in the alpha-galactosidase A gene detected in a Japanese family with Fabry disease. Hum Mutat Suppl 1:S139–S140
Morrone A, Cavicchi C, Bardelli T, Antuzzi D, Parini R, Di Rocco M, Feriozzi S, Gabrielli O, Barone R, Pistone G, Spisni C, Ricci R, Zammarchi E (2003) Fabry disease: molecular studies in Italian patients and X inactivation analysis in manifesting carriers. J Med Genet 40:e103
Nakao S, Takenaka T, Maeda M, Kodama C, Tanaka A, Tahara M, Yoshida A, Kuriyama M, Hayashibe H, Sakuraba H et al (1995) An atypical variant of Fabry’s disease in men with left ventricular hypertrophy. N Engl J Med 333:288–93
Ohno K, Saito S, Sugawara K, Sakuraba H (2008) Structural consequences of amino acid substitutions causing Tay-Sachs disease. Mol Genet Metab (in press)
Okumiya T, Ishii S, Kase R, Kamei S, Sakuraba H, Suzuki Y (1995a) Alpha-galactosidase gene mutations in Fabry disease: heterogeneous expressions of mutant enzyme protein. Hum Genet 95:557–561
Okumiya T, Ishii S, Takenaka T, Kase R, Kamei S, Sakuraba H, Suzuki Y (1995b) Galactose stabilizes various missense mutants of alpha-galactosidase in Fabry disease. Biochem Biophys Res Commun 214:1219–1224
Okumiya T, Kawamura O, Itoh K, Kase R, Ishii S, Kamei S, Sakuraba H (1998) Novel missense mutation (M72 V) of alpha-galactosidase gene and its expression product in an atypical Fabry hemizygote. Hum Mutat Suppl 1:S213–S216
Pappu RV, Hart RW, Ponder JW (1998) Analysis and application of potential energy smoothing for global optimization. J Phy Chem B 102:9725–9742
Ploos van Amstel JK, Jansen RP, de Jong JG, Hamel BC, Wevers RA (1994) Six novel mutations in the alpha-galactosidase A gene in families with Fabry disease. Hum Mol Genet 3:503–505
Ren P, Ponder JW (2003) Polarizable atomic multipole water model for molecular mechanics simulation. J Phy Chem B 107:5933–5947
Rodríguez-Marí A, Coll MJ, Chabás A (2003) Molecular analysis in Fabry disease in Spain: fifteen novel GLA mutations and identification of a homozygous female. Hum Mutat 2:258
Sachdev B, Takenaka T, Teraguchi H, Tei C, Lee P, McKenna WJ, Elliott PM (2002) Prevalence of Anderson- Fabry disease in male patients with late onset hypertrophic cardiomyopathy. Circulation 105:1407–1411
Saito S, Ohno K, Sugawara K, Sakuraba H (2008) Structural and clinical implications of amino acid substitutions in N-acetylgalactosamine-4-sulfatase: Insight into mucopolysaccharidosis type VI. Mol Genet Metab 93:419–425
Sakuraba H, Oshima A, Fukuhara Y, Shimmoto M, Nagao Y, Bishop DF, Desnick RJ, Suzuki Y (1990) Identification of point mutations in the alpha-galactosidase A gene in classical and atypical hemizygotes with Fabry disease. Am J Hum Genet 47:784–789
Sakuraba H, Matsuzawa F, Aikawa S, Doi H, Kotani M, Lin H, Ohno K, Tanaka A, Yamada H, Uyama E (2000) Molecular and structural studies of the GM2 gangliosidosis O variant. J Hum Genet 47:176–183
Sakuraba H, Matsuzawa F, Aikawa S, Doi H, Kotani M, Nakada H, Fukushige T, Kanzaki T (2004) Structural and immunocytochemical studies on alpha-N-acetylgalactosaminidase deficiency (Schindler/Kanzaki disease). J Hum Genet 49:1–8
Schäfer E, Baron K, Widmer U, Deegan P, Neumann HP, Sunder-Plassmann G, Johansson JO, Whybra C, Ries M, Pastores GM, Mehta A, Beck M, Gal A (2005) Thirty-four novel mutations of the GLA gene in 121 patients with Fabry disease. Hum Mutat 25:412
Schiffmann R, Murray GJ, Treco D, Daniel P, Sellos-Moura M, Myers M, Quirk JM, Zirzow GC, Borowski M, Loveday K, Anderson T, Gillespie F, Oliver KL, Jeffries NO, Doo E, Liang TJ, Kreps C, Gunter K, Frei K, Crutchfield K, Selden RF, Brady RO (2000) Infusion of alpha-galactosidase A reduces tissue globotriaosylceramide storage in patients with Fabry disease. Proc Natl Acad Sci USA 97:365–370
Shabbeer J, Yasuda M, Luca E, Desnick RJ (2002) Fabry disease: 45 novel mutations in the alpha-galactosidase A gene causing the classical phenotype. Mol Genet Metab 76:23–30
Shabbeer J, Robinson M, Desnick RJ (2005) Detection of alpha-galactosidase A mutations causing Fabry disease by denaturing high performance liquid chromatography. Hum Mutat 25:299–305
Shabbeer J, Yasuda M, Benson SD, Desnick RJ (2006) Fabry disease: identification of 50 novel alpha-galactosidase A mutations causing the classic phenotype and three-dimensional structural analysis of 29 missense mutations. Hum Genomics 2:297–309
Spada M, Pagliardini S, Yasuda M, Tukel T, Thiagarajan G, Sakuraba H, Ponzon A, Desnick RJ (2006) High incidence of later-onset Fabry disease revealed by newborn screening. Am J Hum Genet 79:31–40
Sugawara K, Saito S, Ohno K, Okuyama T, Sakuraba H (2008) Structural study on mutant alpha-L-iduronidases: insight into mucopolysaccharidosis type I. J Hum Genet 53:467–474
Takata T, Okumiya T, Hayashibe H, Shimmoto M, Kase R, Itoh K, Utsumi K, Kamei S, Sakuraba H (1997) Screening and detection of gene mutations in Japanese patients with Fabry disease by non-radioactive single-stranded conformation polymorphism analysis. Brain Dev 19:111–116
Topaloglu AK, Ashley GA, Tong B, Shabbeer J, Astrin KH, Eng CM, Desnick RJ (1999) Twenty novel mutations in the alpha-galactosidase A gene causing Fabry disease. Mol Med 5:806–811
Verovnik F, Benko D, Vujkovac B, Linthorst GE (2004) Remarkable variability in renal disease in a large Slovenian family with Fabry disease. Eur J Hum Genet 12:678–681
von Scheidt W, Eng CM, Fitzmaurice TF, Erdmann E, Hübner G, Olsen EG, Christomanou H, Kandolf R, Bishop DF, Desnick RJ (1991) An atypical variant of Fabry’s disease with manifestations confined to the myocardium. N Engl J Med 324:395–399
Weiner SJ, Kallman PA, Case DA, Singh UC, Ghio C, Alagona G, Profeta S, Weiner P (1984) A new force field for molecular mechanical simulation of nucleic acids and proteins. J Am Chem Soc 106:765–784
Yam GH-F, Bosshard N, Zuber C, Steinmann B, Roth J (2006) Pharmacological chaperone corrects lysosomal storage in Fabry disease caused by trafficking-incompetent variants. Am J Physiol Cell Physiol 290:C1076–C1082
Yang CC, Lai LW, Whitehair O, Hwu WL, Chiang SC, Lien YH (2003) Two novel mutations in the alpha-galactosidase A gene in Chinese patients with Fabry disease. Clin Genet 63:205–209
Yasuda M, Shabbeer J, Benson SD, Maire I, Burnett RM, Desnick RJ (2003) Fabry disease: characterization of alpha-galactosidase A double mutations and the D313Y plasma enzyme pseudodeficiency allele. Hum Mutat 22:486–492
Acknowledgments
We thank Dr. J. Ponder (Department of Biochemistry and Molecular Biophysics, Washington University) for providing us with the TINKER software. We also thank I.K. McDonald, D. Naylor, D. Jones, J.M. Thornton, S. Hubbard, D.K. Smith, R. Laskowski, and G. Hutchinson for providing us with the HBPLUS. This work was partly supported by grants from the Japan Society for the Promotion of Science; the Ministry of Education, Science, Sports and Culture of Japan; the Ministry of Health and Welfare of Japan; the Japan Science and Technology Agency; and CREST.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclaimers: None.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sugawara, K., Ohno, K., Saito, S. et al. Structural characterization of mutant α-galactosidases causing Fabry disease. J Hum Genet 53, 812–824 (2008). https://doi.org/10.1007/s10038-008-0316-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-008-0316-9
Keywords
This article is cited by
-
High-throughput screening identified disease-causing mutants and functional variants of α-galactosidase A gene in Japanese male hemodialysis patients
Journal of Human Genetics (2012)
-
Fabry-database.org: database of the clinical phenotypes, genotypes and mutant α-galactosidase A structures in Fabry disease
Journal of Human Genetics (2011)
-
Biochemical and structural study on a S529V mutant acid α-glucosidase responsive to pharmacological chaperones
Journal of Human Genetics (2011)
-
Prediction of the clinical phenotype of Fabry disease based on protein sequential and structural information
Journal of Human Genetics (2010)
-
Structural modeling of mutant α-glucosidases resulting in a processing/transport defect in Pompe disease
Journal of Human Genetics (2009)