Abstract
The genomic structures of Oryza sativa (A genome) and O. meyeriana (G genome) were comparatively studied using bicolor genomic in situ hybridization (GISH). GISH was clearly able to discriminate between the chromosomes of O. sativa and O. meyeriana in the interspecific F1 hybrids without blocking DNA, and co-hybridization was hardly detected. The average mitotic chromosome length of O. meyeriana was found to be 1.69 times that of O. sativa. A comparison of 4,6-diamidino-2-phenylindole staining showed that the chromosomes of O. meyeriana were more extensively labelled, suggesting that the G genome is amplified with more repetitive sequences than the A genome. In interphase nuclei, 9-12 chromocenters were normally detected and nearly all the chromocenters constituted the G genome-specific DNA. More and larger chromocenters formed by chromatin compaction corresponding to the G genome were detected in the hybrid compared with its parents. During pachytene of the F1 hybrid, most chromosomes of A and G did not synapse each other except for 1-2 chromosomes paired at the end of their arms. At meiotic metaphase I, three types of chromosomal associations, i.e. O. sativa-O. sativa (A-A), O. sativa-O. meyeriana (A-G) and O. meyeriana-O. meyeriana (G-G), were observed in the F1 hybrid. The A-G chromosome pairing configurations included bivalents and trivalents. The results provided a foundation toward studying genome organization and evolution of O. meyeriana.
Similar content being viewed by others
Introduction
The genus Oryza is composed of two cultivated and more than 20 wild species, and 10 genome sets have been identified including the diploids and tetraploids 1, 2, 3. Wild rice species are extremely valuable gene resources for the genetic improvement of cultivated rice and for the study of genomics 4. A better understanding of the genomic relationships among the species will facilitate effective conservation and efficient utilization of the gene resources in Oryza 5.
Genomic in situ hybridization (GISH) is a powerful method for the characterization of genomes and chromosomes in allopolyploid species 6, 7, 8, hybrid plants 9, 10 and recombinant breeding lines 11, 12. It has played a significant role in understanding genome origin and evolution in the genus Oryza 5, 13, 14, 15, 16, 17. GISH was used to study relationships among the B, C and D genomes, and the result suggested that the D genome was closer to the C than to the B genome 13. The differentiation and relationships among the C, D and E genomes in the O. officinalis complex were detected by multicolor GISH, which indicated that the C, D and E genomes shared a substantial amount of similar sequences, and differentiation between the D and C genomes of O. alta was less than that between the E genome and each of the C and D genomes 5. The different Oryza genomes, A, B and C, involved in rice somatic hybrids, and A and C in hybrid plants, were distinguished using GISH technique 14, 15, 16. Multicolor GISH was also applied to identify the genomic constitution of three tetraploid species (2n=4x=48) in the O. officinalis complex, i.e. O. malampuzhaensis, O. minuta and O. punctata 17.
O. meyeriana is a diploid wild rice species with G genome 2 and distributes in the tropics and subtropics of South and Southeast Asian countries 18. It is one of three native wild rice species in China growing in the southern Yunnan Province and the southwestern Hainan Province 19, 20. A variety of beneficial traits, such as disease and pest resistances, and tolerances to drought and shade stress, have been found in O. meyeriana 18, 21. Efforts have been devoted to understanding the genomic composition 2 and genetic variation 3, 19, 20 of O. meyeriana. However, the cytological features of O. meyeriana and the genomic relationship between A and G genomes have not been thoroughly investigated.
In this study, we applied comparative GISH to distinguish between O. sativa and O. meyeriana chromosomes in their interspecific hybrid, and analyzed the spatial distribution and organization of the two genomes in interphase nuclei. In addition, we also investigated the intergenomic pairing at pachytene and meiotic metaphase I of the hybrid. These data provided a foundation toward studying genome organization and evolution of O. meyeriana.
Materials and methods
Plant materials and probes
O. meyeriana, O. sativa subsp. indica cv IR36 and their interspecific hybrid 22 were the target materials used for chromosome preparation. The 45S rDNA subclone pTa71, which contains the coding sequences for the 45S rRNA genes of wheat 23, and the total genomic DNAs of O. sativa and O. meyeriana were used as probes in fluorescence in situ hybridization (FISH) analysis.
Chromosome preparation
Plants were grown in a greenhouse. Root tips from vigorously growing plants were collected and fixed immediately in ethanol:acetic acid (3:1) at 4 °C overnight. After washing with distilled water, the root tips were treated with a mixture of 2% pectinase (SERVA) and 2% cellulase (SERVA) at 28 °C for 3 h before squashing on slides and drying in a flame 24. For pachytene chromosome preparation, young panicles of plants were harvested and fixed. Microsporocytes at the pachytene stage were macerated in 2% pectinase and 2% cellulase at 28 °C for 2 h and squashed on slides. The slides were kept under −20 °C before FISH.
FISH
Plasmid DNA was isolated following an alkaline lysis method 25. Total genomic DNAs of O. sativa and O. meyeriana were extracted from leaves according to the procedure of Doyle 26. FISH was performed as described previously by Xiong et al. 27. The probes were labelled with digoxigenin-11-dUTP or biotin-11-dUTP, following the protocol for nick translation suggested by the kit maker (Sino-American Biotechnology Company). The biotin-labelled probes were detected in a three-step detection/amplification: streptavidin-Cy3 (Vector Laboratories), biotinylated anti-streptavidin (Vector Laboratories) and streptavidin-Cy3. The digoxigenin-labelled probes were detected with sheep-anti-digoxigenin-FITC (Roche Molecular Biochemicals) and amplified with rabbit-anti-sheep-FITC (Vector Laboratories). For each step of the reaction, slides were incubated at 37 °C for 30 min and washed with phosphate-buffered saline at intervals. Finally the slides were counterstained with 1 μg/mL 4,6-diamidino-2-phenylindole (DAPI) in Vectashield, an antifade solution (Vector Laboratories).
Cytological measurements and analysis
Chromosomes were viewed with a fluorescence microscope system (Olympus BX60 equipped with a cooled CCD camera Sensys 1401E). Gray scale images were captured for each fluorescence channel and then merged with V++ Precision Digital Imaging System software. Digital images were measured using Meta Imaging Series 4.6 software and final images were integrated with Adobe Photoshop v5.1. For comparison of the lengths of the chromosomes between O. meyeriana and O. sativa, chromosomes in five complete mitotic cells of the F1 hybrid were measured and standard deviations were calculated.
Results
GISH analysis of the F1 hybrid of O. sativa × O. meyeriana
The biotin-labelled O. meyeriana genomic DNA as probes without blocking DNA of O. sativa was hybridized to mitotic metaphase chromosomes of the F1 hybrid between O. sativa and O. meyeriana. All the chromosomes of the G genome were labelled with strong red signals and signals were hardly detected on those of the A genome(Figure 1A).
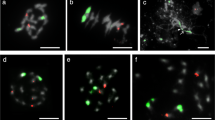
Bicolor GISH with the two genomic probes of A (red) and G (green) at mitotic metaphase of interspecific hybrid. Counterstaining was applied with DAPI (blue). (A) Mitotic chromosomes probed with the genomic DNA of O. meyeriana (red). (B) Discrimination of the two different genomes in mitotic metaphase. (C) B with DAPI-stained chromosomes was converted to a black-and-white image to enhance the visualization of the distribution of euchromatin and heterochromatin in chromosomes of metaphase. (D) Mitotic chromosomes probed with genomic DNA from O. meyeriana (red) and 45S rDNA (green). Arrows indicate the position of NORs.
Bicolor FISH was performed on mitotic chromosomes of the O. sativa × O. meyeriana F1 hybrid (Figure 1B) using the digoxigenin-labelled genomic DNA probes from O. meyeriana and biotin-labelled ones from O. sativa. Two kinds of hybridization signals were mutually exclusive and almost no mixed signals were observed. Twelve chromosomes appeared green and should be from O. meyeriana; the other 12 appeared red and should be from O. sativa. Thus, the A and G genomes were clearly discriminated on the same chromosome preparation.
DAPI binds preferentially to AT-rich regions 28 and is a common dye to stain heterochromatin 29. Scrutiny of a number of cells at metaphase indicated that the chromosomes of O. meyeriana showed heavier DAPI stain than those of O. sativa. To enhance the visualization of distribution of euchromatin and heterochromatin along the mitotic chromosomes, Figure 1B with DAPI-stained chromosomes was converted to a black-and-white image (Figure 1C). For O. sativa, most of the notable heterochromatic regions were observed in the pericentric region and relatively low DAPI staining intensity was distributed in the distal chromosome arms. Unlike O. sativa, the entire chromosomes of O. meyeriana were heavily stained with DAPI.
GISH images clearly demonstrated that the hybrid cells showed an obvious difference in chromosome size between the A and G genomes. The chromosomes from O. meyeriana appeared much larger than those from O. sativa (Figures 1B and C). The chromosomes of the hybrid were measured and arranged in descending order of length (Table 1). The chromosomes from O. meyeriana and O. sativa were compared based on the assumption that karyotypes were similar between corresponding chromosomes among different species of Oryza 30, 31. According to Table 1, we could reach the conclusion that the total chromosomal length of O. meyeriana was 1.69 times that of O. sativa. Each chromosome of O. meyeriana was elongated compared with the corresponding one of O. sativa, but not the same in the elongated ratio for different chromosomes. Chromosomes 4 and 10 carrying nucleolar organizing regions (NORs) were the most and the least elongated chromosomes of O. meyeriana, respectively.
The 45S rDNA has been reported to locate on chromosomes 9 and 10 of O. sativa L. subsp. indica 32. One locus of 45S rDNA was detected on chromosome 9 of O. meyeriana (unpublished data). FISH was also performed on mitotic metaphase chromosomes of the hybrid with probes of 45S rDNA and genomic DNA from O. meyeriana. The result showed that one 45S rDNA site was located on the G genome and two on the A genome (Figure 1D).
Genome spatial distribution and organization in interphase nuclei of the hybrid
In interphase nuclei of the hybrid, heterochromatin is organized as clearly distinguishable chromocenters, which are visible as bright, fluorescent domains after DAPI staining (Figure 2A). In general, 9-12 chromocenters were detected per nucleus. Bicolor GISH with the two parental genomic probes revealed that nearly all the chromocenters corresponded to the G genome-specific DNA. In addition, each of the two parental genomes tended to occupy a separate domain in the nucleus (Figure 2B–2D).
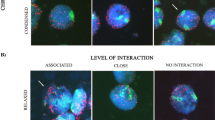
GISH on interphase nuclei and meiotic chromosomes of the interspecific hybrid. Chromosomes counterstained with DAPI. (A) Interphase nuclei stained with DAPI showing large and heavy stained chromocenters. (B) Discrimination of the two different genomes in interphase nuclei.The images of A, C and D were combined. (C) Distribution of the A genome within the nuclei. (D) Distribution of the G genome within the nuclei. (E, F, G) Multicolor GISH with the two genomic probes of A (red) and G (green) on pachytene, diplotene and meiotic metaphase I. (H) Meiotic metaphase I chromosomes probed with genomic DNA from O. meyeriana (red) and 45S rDNA (green). Arrows indicate the position of NORs.
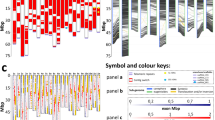
The distribution of the number of chromocenters per nucleus was compared between the hybrid and its parental species (Figure 3). In the nuclei of O. sativa, usually a few prominent chromocenters formed or the formed chromocenters were small. In O. meyeriana, the number of chromocenters varied widely in different nuclei and they were smaller and fainter than those in the hybrids (data not shown).
Chromosome pairing in the hybrid
The ability to differentiate the chromosome sets in the interspecific hybrid by GISH has provided new information on the interaction between homoeologous chromosomes during meiosis. Figure 2E showed a typical example from several hundred mid-pachytene nuclei of the diploid hybrid. At this stage, most chromosomes from O. sativa and O. meyeriana existed as univalent and were not associated with each other. Only one or two homoeologous pairings between the A genome and G genome per nucleus were detected at the terminal regions of chromosome arms. At diplotene, most chromosomes from different genomes were clearly discernible without intertwist. However, it is interesting that the hybrid showed low frequencies of bivalents among O. sativa chromosomes and among O. meyeriana chromosomes (Figure 2F). At meiotic metaphase I, three types of chromosomal associations were distinguished by GISH (Figure 2G), i.e. O. sativa-O. sativa (A-A), O. sativa-O. meyeriana (A-G) and O. meyeriana-O. meyeriana (G-G). The A-G chromosome pairing configurations observed in the hybrid included bivalents and trivalents (Table 2). Using 45S rDNA as a chromosome marker, there was no pairing to be detected between chromosomes carrying NOR (Figure 2H).
Discussion
The GISH method has been used to study differentiation and relationships among the A, B, C, D and E genomes of the genus Oryza 5, 13, 14, 15, 16, 17. The different hybridization stringencies and the rates of blocking DNA used in GISH procedures, and the fluorescence intensities observed between interspecific chromosomes within a nucleus provide phylogenetically important information for elucidating relationships among species in Oryza. GISH identifies A and C genomes in the F1 hybrid between O. sativa and O. officinalis using 16 folds blocking DNA 16. A comparison of the fluorescence signal intensity between C and D genomes within O. latifolia, and B and C genomes in O. minuta, indicated that co-hybridization existed between C and D genomes and B and C genomes 13. Under the different stringent washing conditions, the multicolor GISH results indicated that the C, D and E genomes shared a substantial amount of similar sequences 5. In the present study, GISH was clearly able to discriminate between the chromosomes of O. sativa and O. meyeriana in the F1 hybrid without blocking DNA, and co-hybridization was hardly detected. This result clearly showed considerable divergence of the middle to highly repetitive DNA sequences and confirmed the wide genetic differences between the two species. These observations are in agreement with the results that O. meyeriana is considered as the most distinct species from those with an A genome and occupies the most basal position in the phylogeny of the genus 2, 3.
The quantity of repetitive sequence is the main factor that determines the genome size of similar ploidy 33, 34. Using FISH technique, Uozu et al. 35 analyzed the distribution patterns of genome-specific repetitive DNA sequences in the genus Oryza, except for the G genome. They concluded that the variation in genome size and chromosome morphology is caused by repetitive sequences. According to our GISH results, the genomes of O. sativa and O. meyeriana were evidently different in chromosome length and heterochromatin contents. Obviously, the increase in genome size of O. meyeriana was mainly due to the amplification of repetitive sequences. This result is consistent with the conclusion reported by Aggarwal et al. 2, who demonstrated that the uniqueness of the G genome is due to the species-specific highly repetitive DNA sequences by using total genomic hybridization. Furthermore, because the whole chromosomal regions of O. meyeriana were evenly enhanced with DAPI stain (Figure 1C), our results proved that the chromosomes of O. meyeriana had been extended and evenly enhanced with repetitive sequences during the genome evolution.
The sizes of the A, B, C, E and F genomes of the diploid Oryza species were determined by flow cytometry and image analyses, and the results demonstrated that the total chromosomal length correlated well with the nuclear DNA content 35. We also estimated the DNA content of O. meyeriana according to chromosomal length in the F1 hybrid. This estimation was based on the assumption that (1) the haploid genome size of O. sativa is 430lMb and (2) the euchromatic and heterochromatic regions contain equal amounts of DNA per micrometer of chromosome length. However, the DNA content is usually underestimated for relatively heterochromatic chromosomes 36. The DNA content of O. meyeriana might be more than 727lMb/1C because the species contains a large amount of heterochromatin.
Cytogenetic methods were used to detect the relationships among the genomes of the Oryza species, based on the observation of chromosome pairing behavior at meiotic metaphase I of interspecific hybrids 37, 38, 38, 39, 40. However, chromosome pairing could not discriminate easily the genome constitution of the bivalents in hybrids. Multicolor GISH, using two different genomic probes simultaneously, not only gave high resolution in identifying the chromosome pairing between different genomes in an interspecific hybrid from pachytene to meiotic metaphase I but also provided a direct approach for characterizing genomic differentiations at both chromosomal and DNA levels. Our results showed that multicolor GISH could bring a better result than traditional cytogenetic methods.
Abbreviations
- GISH:
-
(genomic in situ hybridization)
- DAPI:
-
(4,6-diamidino-2-phenylindole)
- FISH:
-
(fluorescence in situ hybridization)
- PBS:
-
(phosphate-buffered saline)
- NOR:
-
(nucleolar organizing region)
References
Vaughan DA . The genus Oryza L. Current status of taxonomy. IRRI Res Paper Series 1989; 138:1–21.
Aggarwal RK, Brar DS, Khush GS . Two new genomes in the Oryza complex identified on the basis of molecular divergence analysis using total genomic DNA hybridization. Mol Gen Genet 1997; 254:1–12.
Ge S, Sang T, Lu BR, Hong DY . Phylogeny of rice genome with emphasis on origins of allotetraploid species. Proc Natl Acad Sci USA 1999; 96:14400–14405.
Chang TT . Conservation of rice genetic resources: luxury or necessity? Science 1984; 224:251–256.
Li CB, Zhang DM, Ge S, Lu BR, Hong DY . Differentiation and inter-genomic relationships among C, E and D genomes in the Oryza officinalis complex (Poaceae) as revealed by multicolor genomic in situ hybridization. Theor Appl Genet 2001; 103:197–203.
Snowdon RJ, Köhler W, Friedt W, Köhler A . Genomic in situ hybridization in Brassica amphidiploids and interspecific hybrids. Theor Appl Genet 1997; 95:1320–1324.
Raina SN, Mukai Y, Yamamoto M . In situ hybridization identifies the diploid progenitor species of Coffea arabica (Rubiaceae). Theor Appl Genet 1998; 97:1204–1209.
Mukai Y, Nakahara Y, Yamamoto M . Simultaneous discrimination of the three genomes in hexaploid wheat by multicolor fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome 1993; 36:489–495.
Leith AR, Schwarzacher T, Mosgöller W, Bennett MD, Heslop-Harrison JS . Parental genomes are separated throughout the cell cycle in a plant hybrid. Chromosoma 1991; 101:206–213.
Li HJ, Guo BH, Li YW, et al. Molecular cytogenetic analysis of intergeneric chromosomal translocations between wheat (Triticum aestivum L.) and Dasypyrum villosum arising from tissue culture. Genome 2000; 43:756–762.
Schwarzacher T, Anamthawat-Jónsson K, Harrison GE, et al. Genomic in situ hybridization to identify alien chromosomes and chromosome segments in wheat. Theor Appl Genet 1992; 84:778–786.
Leggett JM, Markhand GS . The genomic identification of some monosomics of Avena sativa L. cv. Sun II using genomic in situ hybridization. Genome 1995; 38:747–751.
Fukui K, Shishido R, Kinoshita T . Identification of the rice D-genome chromosomes by genomic in situ hybridization. Theor Appl Genet 1997; 95:1239–1245.
Shishido R, Apisitwanich S, Ohmido N, et al. Detection of specific chromosome reduction in rice somatic hybrids with the A, B, and C genomes by multi-color genomic in situ hybridization. Theor Appl Genet 1998; 97:1013–1018.
Yan H, Min S, Zhu L . Visualization of Oryza eichingeri chromosomes in intergenomic hybrid plants from O. sativa × O. eichingeri via fluorescent in situ hybridization. Genome 1999; 42:48–51.
Yan HH, Cheng ZK, Liu GQ, et al. Identification of Oryza sativa × Oryza officinalis F1 and backcross progenies using genomic in situ hybridization. Acta Genet Sin 1999; 26:157–162.
Li CB, Zhang DM, Ge S, Lu BR, Hong DY . Identification of genome constitution of Oryza malampuzhaensis, O. minuta, and O. punctata by multicolor genomic in situ hybridization. Theor Appl Genet 2001; 103:204–211.
Vaughan DA . The wild relatives of rice: a genetic resources handbook. Manila: International Rice Research Institution, 1994:34–60.
Qian W, Ge S, Hong DY . Assessment of genetic variation of Oryza granulata detected by RAPDs and ISSRs. Acta Bot Sin 2000; 42:741–750.
Qian W, Ge S, Hong DY . Genetic variation within and among populations of a wild Oryza granulata from China detected by RAPDs and ISSRs markers. Theor Appl Genet 2000; 102:440–449.
Ou SH . Rice disease, 2nd edition. UK: The Cambrian News Ltd, 1985:109–201.
Huang YL, Shu LH, Zhu LL, et al. Characterization of the intersectional hybrid between Oryza sativa and O. meyeriana indigenous to China. J Wuhan Univ (Nat Sci Ed) 2000; 46:739–744.
Gerlach WL, Bedbrook JR . Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 1979; 7:1869–1885.
Jin WW, Li ZY, Fang Q, et al. Fluorescence in situ hybridization analysis of alien genes in Agrobacterium-mediated Cry1A(b)-transformed rice. Ann Bot 2002; 90:31–36.
Sambrook J, Fritsch EF, Manitiatis T . Molecular cloning: a laboratory manual. New York: Cold Spring Laboratory Press, 1998:25–29.
Doyle JJ . Isolation of plant DNA from fresh tissue. Focus 1998; 12:13–15.
Xiong ZY, Gao Y, He GY, et al. Knob-associated tandem repeats on mitotic chromosomes in maize, Zea diploperennis and their hybrids. Chin Sci Bull 2004; 49:1354–1357.
Kapuscinski J . DAPI: a DNA-specific fluorescent probe. Biotechnic Histochem 1995; 70:220–233.
Cheng Z, Presting GG, Buell R, Wing RA, Jiang J . High-resolution pachytene chromosome mapping of bacterial artificial chromosomes anchored by genetic markers reveals the centromere location and the distribution of genetic recombination along chromosome 10 of rice. Genetics 2001; 157:1749–1757.
Kurata N, Omura T . Karyotype analysis in rice III. Karyological comparisons among four Oryza species. Japan J Breed 1982; 32:253–258.
Fukui K, Iijima K . Somatic chromosome map of rice by imaging methods. Theor Appl Genet 1991; 81:589–596.
Shishido R, Sano Y, Fukui K . Ribosomal DNAs: an exception to the conservation of gene order in rice genomes. Mol Gen Genet 2000; 263:586–591.
Flavell RB . The molecular characterization and organization of plant chromosomal DNA sequences. Annu Rev Plant Physiol 1980; 31:569–596.
Kubis S, Schmidt T, Heslop-Harrison JS . Repetitive DNA elements as a major component of plant genomes. Ann Bot 1998; 82:45–57.
Uozu S, Ikehashi H, Ohmido N, et al. Repetitive sequence: cause for variation in genome size and chromosome morphology in the genus Oryza. Plant Mol Biol 1997; 35:791–799.
Cheng Z, Buell R, Wing RA, Gu M, Jiang J . Toward a cytological characterization of rice genome. Genom Res 2001; 11:2133–2141.
Morinaga T . Cytogenetical studies on Oryza sativa L. VI. The cytogenetics of a hybrid of O. minuta Presl. and O. latifolia Desv. Jap J Bot 1943; 12:347–357.
Li HW, Chen CC, Weng TS, Wuu KD . Cytogenetical studies of Oryza sativa L. and its related species. 4. Interspecific crosses involving O. australiensis with O. sativa and O. minuta. Bot Bull Acad Sinica 1963; 4:65–74.
Hu CH . Cytogenetic studies of Oryza officinalis complex. III. The genomic constitution of O. punctata and O. eichingeri. Cytologia 1970; 35:304–318.
Katayama T . Cytogenetical studies on the genus Oryza IX. The F1 hybrids from synthesized amphiploid'species with the BBCC genomes. Jpn J Genet 1977; 52:301–307.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 39870423) and the Key Program of Ministry of Education of China.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Xiong, Z., Tan, G., He, G. et al. Cytogenetic comparisons between A and G genomes in Oryza using genomic in situ hybridization. Cell Res 16, 260–266 (2006). https://doi.org/10.1038/sj.cr.7310033
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310033
Keywords
This article is cited by
-
Genomic constitution, allopolyploidy, and evolutionary proposal for Cynodon Rich. based on GISH
Protoplasma (2022)
-
Identification and characterization of karyotype in Passiflora hybrids using FISH and GISH
BMC Genetics (2018)
-
Development of monosomic alien addition lines from the wild rice (Oryza brachyantha A. Chev. et Roehr.) for introgression of yellow stem borer (Scirpophaga incertulas Walker.) resistance into cultivated rice (Oryza sativa L.)
Euphytica (2016)
-
Characterization of an Alien Chromosome of Oryza officinalis Transferred the Genomic and Cytological Environment of Oryza sativa
Journal of Plant Biology (2010)
-
Development and characterization of interspecific hybrids between Oryza sativa and O. latifolia by in situ hybridization
Science Bulletin (2008)