Abstract
We previously demonstrated using noninvasive technologies that the interferon-gamma (IFN-γ) receptor complex is preassembled 1. In this report we determined how the receptor complex is preassembled and how the ligand-mediated conformational changes occur. The interaction of Stat1 with IFN-γR1 results in a conformational change localized to IFN-γR1. Jak1 but not Jak2 is required for the two chains of the IFN-γ receptor complex (IFN-γR1 and IFN-γR2) to interact; however, the presence of both Jak1 and Jak2 is required to see any ligand-dependant conformational change. Two IFN-γR2 chains interact through species-specific determinants in their extracellular domains. Finally, these determinants also participate in the interaction of IFN-γR2 with IFN-γR1. These results agree with a detailed model of the IFN-γ receptor that requires the receptor chains to be pre-associated constitutively for the receptor to be active.
Similar content being viewed by others
Introduction
The interferon-gamma (IFN-γ) receptor complex is composed of four transmembrane polypeptides: two ligand-binding chains (IFN-γR1, also known as the α chain), and two accessory chains (IFN-γR2, also known as the β chain) that do not bind IFN-g independently but are required for IFN-γ to exhibit any biological activity 2, 3, 4, 5, 6, 7, 8. The structure of the resting IFN-γ receptor complex has been in debate because of conflicting reports based on experiments that destroy the integrity of the cell and hence the receptor's microenvironment 9, 10, 11, 12. By using confocal microscopy coupled to fluorescence resonance energy transfer (FRET), we demonstrated that the receptor complex in its resting state in intact cells is a preformed tetramer that undergoes a conformational change when cells are treated with IFN-γ 1. After treatment with IFN-γ, the intracellular domains of the IFN-γR1 and IFN-γR2 chains move apart from one another.
Ligand-dependant conformational changes could come from proteins that are recruited to the activated receptor complex, such as Stat1. The interaction of Stat1 with IFN-γR1 requires the phosphorylation of Tyr-457, which occurs only after the IFN-γ receptor is activated by IFN-γ 13. This tyrosine residue is part of a motif (Tyr457Asp458Lys459Pro460His461) that is highly conserved among tetrapods and is critical for Stat1 recruitment to the active IFN-γ receptor complex 13, 14, 15, 16.
Clues to what holds the receptor together exist in the literature, but their roles in receptor preassembly are not determined. The Janus (Jak) kinases Jak1 and Jak2 play a critical role in IFN-γ signaling 17, 18. Jak1 and Jak2 bind to the intracellular domains of the chains of the IFN-γ receptor complex, providing it with tyrosine kinase activity 9, 10, 11, 19, 20. Mutation of the intracellular domains of human IFN-γR1 (Leu283Pro284Lys285Ser286) and of human IFN-gR2 (Pro284Pro285Ser286Ile287Pro288 ... Ile291Glu292Glu293Tyr294Leu295) revealed the sites at which Jak1 and Jak2 constitutively bind, respectively and established that the Jak kinases must be bound to the receptor chains for signaling by IFN-γ to be observed 19, 21. Curiously, the kinase activity of Jak1 is only necessary for some activities of the IFN-γ receptor, implying a primarily structural role for Jak1; in contrast the kinase activity of Jak2 was required to observe any activity from the IFN-γ receptor 22. Jak kinases bind to cytokine receptors via their NH2-terminal FERM domains 23, 24, 25, 26, 27, while their catalytic activity and nearly all observed tyrosine phosphorylations occur through their COOH-terminal kinase domains 28, 29.
There is strong species-specificity between IFN-γ and its receptor: IFN-γ from one species generally has insignificant biological activity on cells from another species. For example, human IFN-γ cannot bind to murine IFN-γR1 nor can it activate the murine IFN-γ receptor complex and vice versa. The species specificity is restricted to the extracellular domains of IFN-γR1 and IFN-γR2 and by the ligand IFN-γ: all three must be from the same species for signaling to be observed 2, 3, 4, 5, 30, 31, 32, 33, 34, 35, 36, 37, 38. In contrast, the intracellular domains can be interchanged between species without affecting IFN-γ signaling 30, 31, 37. Thus specific elements in the extracellular domains play a role in the associations between these chains.
In this report, we changed these elements of the IFN-γ receptor complex by mutagenesis and domain swapping to define their roles in the preassembly and ligand-induced changes in the IFN-γ receptor complex.
Material and methods
Cells, media, and reagents
COS-1 and U3A cells (mutant human 2fTGH cells lacking Stat1 39) were maintained in DMEM (Sigma Aldrich) supplemented with 10% (v:v) fetal bovine serum (FBS). No antibiotics were used during the passaging of these cell lines. Dimethylsulfoxide, chloroquine diphosphate, and diethylaminoethyl-dextran were purchased from Sigma-Aldrich. Lipofectamine was obtained from Qiagen, Inc. (Valencia, CA). Restriction endonucleases and T4 DNA ligase were obtained from New England Biolabs. Shrimp alkaline phosphatase was obtained from Boehringer-Mannheim. IFN-γ was obtained from PBL Biomedical Laboratories (Piscataway, NJ). Thermus acquaticus DNA polymerase was purified according to a published protocol 40. Cells were treated with 1500 units/mL IFN-γ unless otherwise indicated. Site-directed mutagenesis was performed with the QuikChange Site-directed Mutagenesis Kit from Strategene, Inc. (La Jolla, CA) according to the manufacturer's recommendations. In this report, mutated amino acids are numbered assuming the initial methionine at the beginning of the signal peptide is the first amino acid. Overlapping primers designed to create the Y457F mutant of IFN-γR1/EBFP or IFN-γR1 are (sense) 5′-CCA CCT CCT TTG GTT TTG ATA AAC CAC ATG TG-3′ and (antisense) 5′-CAC ATG TGG TTT ATC AAA ACC AAA GGA GGT GG-3′. Overlapping primers designed to create the P284L mutant on IFN-γR1/EGFP or IFN-γR1 are (sense) 5′-GGA AAA AAG CAT AAT AAT AC TT AAGTC CTT GAT CTC TGT GGT AAG-3′ and (antisense) 5′-CTT ACC ACA GAG ATC AAG GACTT AAGT AAT ATT ATG CTT TTT TCC-3′. The AflII site in both primers is underlined. Overlapping primers designed to create the PPSIP284-288AAAAA mutant on FL-IFN-γR2/GFP are (sense) 5′-AAA TAC TGG TTT CAC ACT GCC G C GGC C GC T GCC TTA CAG ATA GAA GAG TAT-3′ and (antisense) 5′-ATA CTC TTC TAT CTG TAA GGC A GCGGCCGC GGC AGT GTG AAA CCA GTA TTT- 3′. The NotI site in both primers is underlined. Nucleotides mutated from their wildtype sequences are in boldface in the sense strand, and anticodons mutated from their wildtype sequences are in boldface in the antisense strand.
Receptor constructs
Hu-IFN-γR2 was expressed with a FLAG epitope (ADYKDDDDK) on its NH2-terminus 41.
Hu-IFN-γR1 was modified by PCR in two steps in order to place the GFP variants onto the COOH-terminus. The following forward and reverse primers were used to amplify the transmembrane and intracellular domains of IFN-γR1: 5′-GTC GCTAGC GAT TCC AGT TGT TGC TGC T-3′ and 5′-GCC GCGGCCG CT AA TCCG AG AAA ATT CTT TGG AAT C-3′. The forward primer encodes an NheI restriction endonuclease site (underlined). The nucleotides in boldface encode amino acids that are mutated from the corresponding wild-type sequences. This mutation, encoding the first NH2-terminal three amino acids of the transmembrane domain of Hu-IFN-γR1, does not alter its biological function in any tested assay 1, 42. Each reverse primer encodes a BspEI restriction endonuclease site (single underline), followed by a TAG stop codon, in turn followed by an EagI restriction endonuclease site (double underline). This places a nonanucleotide sequence (in boldface) encoding Asp-Pro and a stop codon in place of the original stop codon of the wildtype receptor sequence. This mutation also does not appear to inhibit the biological function of the receptor (C Krause and S Pestka, unpublished observations). After amplification of the transmembrane and intracellular domains by PCR, the products were digested with NheI and EagI restriction endonucleases and ligated to pEF3-IFN-γR1 1 digested with NheI and NotI to free their transmembrane and intracellular domains. Ligation of cohesive ends yielded plasmid pEF3-IFN-γR1.
To insert the GFP variants into pEF3-IFN-γR1 and pcDNA3-FL-IFN-γR2, the GFP variants were amplified and the ends of their cDNA are altered by PCR. The following forward primer was used to amplify GFP from pEF2-S65T (derived from plasmid pS65T, a gift from Roger Y Tsien): 5′-GCG CT CCGGATCC CA TGA GTA AAG GAG AAG AAC-3′. The following forward primer was used to amplify EBFP, EGFP, ECFP and EYFP from pEF2-EBFP and pIRES-EGFP (a gift from Dr Jerome Langer), pEF2-ECFP and pEF2-EYFP, respectively: 5′-GCG CT CCGGATCC CA TGG TGA GCA AGG GCG AGG-3′. The forward primers encode a fused BspEI/BamHI site (underlined) that encodes a Pro-Asp-Pro tripeptide sequence (whose nonanucleotide codon sequence is in boldface) before the initiating methionine in the GFP sequence. The BspEI site is in identical frame to the natural BspEI site found in the COOH-terminal end of the intracellular domain of the human IFN-γR2 open reading frame and the BspEI site in pEF3-IFN-γR1. The reverse primer, the SP6 primer, hybridizes to the pEF2 backbone and allows amplification of a NotI restriction endonuclease site in the multicloning site of the host vector after the stop codon. The amplified product was digested with BspEI and NotI. Plasmids pEF3-IFN-γR1 and pcDNA3-FL-IFN-γR2 were digested with BspEI and NotI restriction endonucleases. Ligation of the digested PCR products encoding the various GFP mutants to the digested receptors above yielded receptors with the fluorescent proteins. A fusion between IFN-γR1 and EGFP (enhanced GFP) was used. EGFP was placed on IFN-γR1 instead of GFP because it was discovered that IFN-γR1/GFP was expressed at barely detectable levels, while use of EGFP instead of GFP at the COOH-terminus of IFN-γR1 overcame this limitation. A slash separates the intracellular domain from additions to the COOH-termini of the receptor chains such as GFP. These expression plasmids allowed the following receptor chains to be synthesized: IFN-γR1/EGFP (or EBFP, ECFP, EYFP replacing GFP), or IFN-γR1, or FL-IFN-γR2/GFP (or EBFP, ECFP, EYFP replacing GFP) or FL-IFN-γR2.
To create the chimeric receptor MH-IFN-γR2/GFP (where MH denotes a murine/human chimera) in which the extracellular domain is from Mu-IFN-γR2 (Mu, mouse) whereas the transmembrane and intracellular domain is from FL-Hu-IFN-γR2/GFP (Hu, human), we amplified the extracellular domain of Mu-IFN-γR2 from plasmid pcDNA3-Mu-IFN-γR2 37 by PCR with the following forward and reverse primers: 5′-GAT CGGTACCGG GCC ATG CGG CCT TTG-3′ and 5′-CAT GGCTAGCCA TTG CTG CAG CCT GGC GGA G-3′, where the KpnI and NheI restriction endonuclease sites used for the subsequent cloning step are underlined. Plasmid pcDNA3-FL-IFN-γR2/GFP was digested with KpnI and NheI, releasing the human extracellular domain. The large fragment retaining the transmembrane and intracellular domains of FL-IFN-γR2/GFP was ligated to the resulting PCR product digested with KpnI and NheI. Ligation of cohesive ends yielded pcDNA3-MH-IFN-γR2/GFP that expressed the chimeric fluorescent receptor MH-IFN-γR2/GFP.
Tandem vectors were synthesized as previously described so that two or more proteins would be co-expressed in a single cell more efficiently than could be achieved by transfection of individual plasmids 1, 41.
Transfections
Stable transfections were performed in U3A cells with Lipofectamine as previously published 11. Transient transfections of COS-1 cells with diethylaminoethyl-dextran were performed as reported 1, 41.
Confocal fluorescence spectroscopy
A custom-built instrument coupling a monochromator to a confocal microscope was described previously 43 with minor enhancements 1 to analyze interactions between GFP-tagged, EGFP-tagged and EBFP-tagged receptor chains. A femtosecond pulsed titanium:sapphire mode-locked tunable infrared laser with argon continuous-wave ion pump laser (both from Coherent, Inc.) was used as the primary excitation source. An independent 15 mW continuous-wave argon-ion laser was used as a secondary excitation source, and 488 nm light was isolated from the multiline argon laser emission with a prism followed by a 488 nm band-pass filter. Infrared light was filtered with a 670DCSP dichroic mirror and a 600SP short pass filter, while 488 nm light was filtered with a 500DCLP dichroic mirror and 488 nm notch filters.
Cells were washed twice with PBS, placed in a 250 μL quartz-bottomed (0.17 mm thickness) cylindrical sample well, covered with 200 μL PBS, and were allowed to settle. After settling of cells, cells possessing GFP emission when excited by 488 nm epifluorescent light (derived by defocusing the incident 488 nm laser light) were selected at random for spectral analysis. In some experiments, 380 nm light was used to qualitatively assay FRET in cells known to express GFP as determined above. The 380 nm epifluorescence light was generated from 760 nm light by passing the infrared laser light through a doubling crystal, filtering the remaining infrared light with a 400 nm short-pass filter, and defocusing the 380 nm light prior to exciting the sample. Individual cells were excited by two-photon excitation at 760 nm. Fluorescent light emitted from the samples was directed to either an Acton-150 monochromator (Roper Scientific) to be detected by a LN-CCD512TK back-illumination CCD camera (Roper Scientific) for spectral analysis or to an SPCM avalanche photodiode (Perkin-Elmer) capable of single fluorophore detection for imaging analysis. Spectral data were recorded with an exposure time of five seconds with WinSpec software (Roper Scientific), and were translated into a platform-independent data table format with Igor software (Wavemetrics). A Diaphot 300 (Nikon, Inc.) microscope shell was coupled to a Queensgate confocal scanner/stage micrometer. Areas of the imaging field ranging from 250 × 250 nm2 (the physical limit set by the frequency of laser light during two-photon excitation and the numerical aperture of the objective lens, which is 1.3 here) to 50 × 50 mm2 to probe were selected from a single cell for spectral analysis. Unless otherwise noted, areas of at least 0.4 mm2 were evaluated. At least seven cells expressing GFP were randomly selected for quantitative analysis in each transfection and the results from a representative cell presented. At least thirty more cells expressing GFP were illuminated by 380 nm epifluorescent light to judge FRET visually in cells throughout the population to ensure that a representative sample of the population was analyzed quantitatively.
To analyze FRET between ECFP and EYFP in intact cells, a similar instrument to the one above was designed. An Olympus FV300 confocal system on an IX71 microscope with an automatic stage controller was configured with helium-cadmium solid-state laser (442 nm), multiline argon ion laser (458/488/514 nm), HeNe Green laser (543 nm) and HeNe Red laser (633 nm) lines). Samples were illuminated by either transmitted light and visualized by light microscopy or differential image contrast microscopy (Nomarsky microscopy), by widefield epifluorescence with a mercury lamp as the excitation source, or by laser excitation from either one of the above. The confocal microscope was specially modified with an optional fiber output to couple to other detection devices. Fluorescent light emitted from the sample was directed to either a photomultiplier tube for confocal image analysis or to an Acton SP-150 Spectrophotometer for frequency-resolved photon collection with a CCD camera (Andor Technology, Model DV420) through the fiber output. Fluorescent light was analyzed with either Fluoview software from Olympus, Inc. (confocal image) or with Andor MCD software from Andor Technology (emission spectra). Emission spectra were calibrated manually with the software package by observing the simultaneous emission of the 488, 514, 543 and 633 nm lasers above.
Results
Stat1 recruitment is required for ligand-dependant alteration in the structure of IFN-γR1
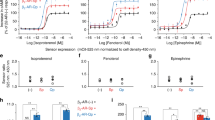
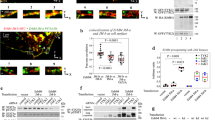
We previously demonstrated that the intracellular domains of the IFN-γR1 and IFN-γR2 move apart or rotate with respect to each other after IFN-γ is added to cells 1. Stat1 is recruited to the IFN-γ receptor complex after the IFN-γ receptor complex is activated and Tyr-457 of IFN-γR1 is phosphorylated 13. To test our hypothesis that Stat1 recruitment to Tyr-457 is required for separation of the intracellular domains of the IFN-γ receptor complex, we mutated Tyr-457 of IFN-γR1/EBFP to Phe to prevent Stat1 recruitment. We coexpressed this receptor mutant [Y457F]IFN-γR1/EBFP with FL-IFN-γR2/GFP in COS-1 cells, then measured FRET before and after IFN-γ treatment. Prior to IFN-γ treatment, there was substantial FRET between the fluorescent receptor pairs whether IFN-γR1/EBFP was mutated (Figure 1A, black line) or not (Figure 1B, black line) indicating that neither receptor preassociation nor EBFP orientation was affected by this mutation. Whereas IFN-γ produced a large reduction in FRET in the cells expressing unmutated IFN-γR1/EBFP (Figure 1B, red line), treatment of the cells containing [Y457F]IFN-γR1/EBFP with IFN-γ produced little or no change in FRET (Figure 1A, red line). This demonstrated that the major ligand-induced change in FRET requires an intact Stat1 recruitment site on IFN-γR1.
Mutation of Tyr-457 in IFN-γR1/EBFP eliminated ligand-dependant changes in FRET between IFN-γR1/EBFP and FL-IFN-γR2/GFP. (A) Fluorescence emission spectra of COS-1 cells coexpressing [Y457F]IFN-γR1/EBFP and FL-IFN-γR2/GFP. The black line represents the fluorescent emission spectrum taken from an area of a representative cell expressing this receptor pair prior to IFN-γ addition to cells. The red line represents a spectrum taken from an area of a representative cell from the same population of transfected cells after IFN-γ addition to cells (30 min). (B) Fluorescence emission spectra of COS-1 cells coexpressing IFN-γR1/EBFP and FL-IFN-γR2/GFP. The black line represents the fluorescent emission spectrum taken from an area of a representative cell expressing this receptor pair prior to IFN-γ addition to cells (25 min). The red line represents a spectrum taken from an area of a representative cell from the same population of transfected cells after IFN-γ addition to cells. (C) Fluorescence emission spectra of COS-1 cells coexpressing IFN-γR1/ECFP and FL-IFN-γR2/EYFP. The black line represents the fluorescent emission spectrum taken from an area of a representative cell expressing this receptor pair prior to IFN-γ addition to cells. The red line represents a spectrum taken from an area of a representative cell from the same population of transfected cells after IFN-γ addition to cells (23 min). (D) Fluorescence emission spectra of U3A cells coexpressing IFN-γR1/ECFP and FL-IFN-γR2/EYFP. The black line represents the fluorescent emission spectrum taken from an area of a representative cell expressing this receptor pair prior to IFN-γ addition to cells. The red line represents a spectrum taken from the same cell after IFN-γ addition (65 units/mL, 20 min). Differences in the emission spectra themselves are caused by differing exciting lasers, different GFP molecules in cells, and differing abilities for background cellular fluorescent components to be excited in each system.
To prove that Stat1 recruitment to IFN-γR1 was responsible for the major conformational change between IFN-γR1 and IFN-γR2 in response to IFN-γ, we transfected the fluorescent receptor pair IFN-gR1/ECFP and FL-IFN-γR2/EYFP into COS-1 cells and U3A cells. Cells were analyzed for FRET before and after IFN-γ treatment. The interaction between IFN-γR1 and IFN-γR2 in the absence of ligand in stably transfected cells (Figure 1D, black lines) is similar to that in transiently transfected cells (Figure 1A, 1B, 1C, black lines). Furthermore, the interaction between IFN-γR1 and IFN-γR2 is similar whether the GFP/EBFP pair or the ECFP/EYFP pair was used, even though the shape of the spectra, the emission maxima, and the wavelength axes for the figures are different. The decrease of FRET seen in COS-1 cells after treatment with IFN-γ (Figure 1C, red line) was not seen in U3A cells after treatment with IFN-γ (Figure 1D, red line).
To determine whether the ligand-dependant interaction seen between two fluorescent IFN-γR2 chains in the presence of nonfluorescent IFN-γR1 is affected when Stat1 recruitment to IFN-γR1 is inhibited, we mutated nonfluorescent IFN-γR1 in the same way as was done for IFN-γR1/EBFP above and coexpressed [Y457F]IFN-γR1 with FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP. Mutation of the Stat1 recruitment site of IFN-γR1 did not significantly affect the extent of FRET between the two IFN-γR2 chains in the absence of IFN-γ (Figure 2A, 2B, black lines). The IFN-γ-mediated decrease in FRET between the two IFN-γR2 chains observed in the presence of wild-type IFN-γR1 (Figure 2A, red lines) 1 was virtually unaltered when IFN-γR1 could not recruit Stat1 (Figure 2B, red line). Because there was no difference in FRET between the IFN-γR2 chains when Stat1 was not recruited to IFN-γR1 (cf. Figure 2A, 2B), we deduced that the decrease in FRET between IFN-γR1 and IFN-γR2 chains upon addition of IFN-γ (Figure 1B, 1C) is due to the reorientation of the IFN-γR1 chain.
The ligand-dependant interaction between the IFN-γR2: IFN-γR2 chains is not significantly affected by Stat1 recruitment. (A) COS-1 cells coexpressing IFN-γR1, FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP were probed with 760 nm light, and fluorescence emission spectra were recorded. (B) COS-1 cells coexpressing [Y457F]IFN-γR1, FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP were probed with 760 nm light, and fluorescence emission spectra were recorded. In both (A) and (B), the black emission spectra were those from cells in the absence of IFN-γ, while spectra in red were recorded from cells in the presence of IFN-γ (20 min).
The presence of Jak1 in the receptor complex is required for preassembly
IFN-γR1 and IFN-γR2 bind Jak kinases at their juxtamembrane segments 6, 7, 19, 20, 42, 44. There is tremendous variation of the cytoplasmic sequences beyond the juxtamembrane region of IFN-γR1 relative to the extracellular domains among various tetrapod species. The most strongly conserved segments of the intracellular domains of IFN-γR1 are the areas required for function such as the Stat1 binding site and the Jak1 association sites (Figure 3, left). The Jak2 association site is also strongly conserved among IFN-γR2 sequences from various tetrapod species (Figure 3, right). Mutation of these segments eliminates the ability of Jak1, Jak2 or Stat1 to bind to the receptor and results in inactive receptor complexes 15, 19, 20, 21, 42, 45. Mutation of Pro-284 in the Jak-kinase binding site of IFN-γR1 to an alanine residue virtually eliminated the ability of Jak1 to be immunoprecipitated with IFN-γR1 and resulted in a nonfunctional IFN-γ receptor complex when coexpressed with IFN-γR2 19. We mutated this proline to leucine in IFN-γR1/EGFP to create an AflII restriction site was generated so that mutated clones could be easily identified by restriction digestion. This mutant, [P284L]IFN-γR1/EGFP, was expressed on the surface of cells but could not to reconstitute an active IFN-γ receptor complex when coexpressed with IFN-γR2 (data not shown).
Sequence alignment of IFN-γR1 and IFN-γR2 from tetrapod species. (left) IFN-γR1 sequences from human, mouse, chicken, and frog were aligned and conserved regions are illustrated. Shading of conserved regions was done with the BOXSHADE algorithm with a stringency setting of 0.7. Black shading indicates identity whereas grey shading indicates conservation with a consensus sequence. Secondary structure elements, based on the crystal structure of IFN-γR1 55 are shown with a box. (right) IFN-γR2 sequences from human, pig, mouse, chicken, and frog were aligned and conserved regions are illustrated. Shading of conserved regions was done with the BOXSHADE algorithm with a stringency setting of 0.7. Black shading indicates identity with a consensus sequence, while grey shading indicates conservation with a consensus sequence. Putative secondary structure elements identified by sequence homology and based on the crystal structure of IFN-γR1 55 are shown with a box.
We coexpressed [P284L]IFN-γR1/EGFP with FL-IFN-γR2/EBFP or with IFN-γR1/EBFP in COS-1 cells. To determine if the biologically inactive receptor still maintains its constitutive interactions with IFN-γR2. We observed that FRET was greatly reduced when [P284L]IFN-γR1/EGFP was coexpressed with FL-IFN-γR2/EBFP (Figure 4A, blue vs. green lines). Addition of IFN-g to cells coexpressing [P284L]IFN-γR1/EGFP and FL-IFN-γR2/EBFP did not change the FRET pattern between [P284L]IFN-γR1/EGFP and FL-IFN-γR2/EBFP (Figure 4B, black vs. red lines). This indicates that the binding of Jak1 to IFN-γR1 is required for optimal assembly of IFN-γR1 with IFN-γR2 as well as for Stat1-dependant changes in the conformation of IFN-γR1.
Mutation of the Jak1 association site on IFN-γR1 disrupts the interaction of IFN-γR1 with IFN-γR2. (A) IFN-γR1/EGFP and FL-IFN-γR2/EBFP were coexpressed in COS-1 cells, and a representative spectrum shown by the green line was recorded upon illumination with 760 nm light. [P284L]IFN-γR1/EGFP and FL-IFN-γR2/EBFP were coexpressed, and upon 760 nm excitation, a representative spectrum shown by the blue line was observed. (B) [P284L]IFN-γR1/EGFP and FL-IFN-γR2/EBFP were coexpressed and COS-1 cells expressing this receptor pair were treated with IFN-γ. Representative spectra were taken from cells prior to and after addition of IFN-γ (12 min) (black and red lines, respectively).
The interactions between two IFN-γR2 chains in the presence or absence of Jak1 in the receptor complex were determined. We did not see a major difference in the FRET between the two IFN-γR2 chains whether or not the Jak1 recruitment site on IFN-γR1 was intact (Figure 5A, black line) or mutated (Figure 5A, purple line), possibly because there is only a small increase in FRET between two IFN-gR2 chains when IFN-γR1 is present 1. When IFN-γ was added to cells possessing FL-IFN-γR2/EBFP, FL-IFN-γR2/GFP and [P284L]IFN-γR1, there was a small decrease in FRET between the two IFN-γR2 chains (Figure 5B). This decrease in FRET was less than that seen between two IFN-γR2 chains in the presence of intact IFN-γR1 (Figure 2A). Thus, the absence of Jak1 binding to IFN-γR1 alters ligand-induced rearrangements between the IFN-γR2 chains.
Mutation of the Jak1 association site reduces ligand-dependant changes in the IFN-γR2 chains. (A) COS-1 cells coexpressing FL-IFN-γR2/GFP, FL-IFN-γR2/EBFP, and either wild-type IFN-γR1 (in black) or [P284L]IFN-γR1 (in purple) were excited with 760 nm light and fluorescence emission spectra taken. (B) A representative COS-1 cell coexpressing FL-IFN-γR2/GFP, FL-IFN-γR2/EBFP, and [P284L]IFN-γR1 was excited with 760 nm light prior to (black line) and after (red line) 25 min treatment with IFN-γ, and fluorescence emission spectra taken.
The presence of Jak2 does not contribute to the preassembly of the receptor complex
Unlike IFN-γR1, a single amino acid mutation in the intracellular domain of IFN-γR2 did not eliminate the ability of IFN-γR2 to interact with Jak2, but mutation of one of two pentapeptide regions was necessary and sufficient to inhibit the coimmunoprecipitation (and presumably the binding) of Jak2 with IFN-γR2 20. To investigate the influence of the Jak2 association site in receptor complex preformation, we mutated the first pentapeptide region of FL-IFN-γR2/GFP from Pro284Pro285Ser286Ile287Pro288 (numbering here includes the putative signal peptide but not the FLAG epitope) to alanines 20 to create [PPSIP(284-288)AAAAA]FL-IFN-γR2/GFP (designated FL-IFN-γR2DJak2/GFP). We selected a penta-alanine codon sequence such that a NotI restriction site would be created to facilitate screening of mutated FL-IFN-γR2/GFP receptor cDNAs. We coexpressed FL-IFN-γR2DJak2/GFP with either FL-IFN-γR2/EBFP or with IFN-γR1/EBFP and investigated the level of FRET relative to the FRET observed previously between FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP or between FL-IFN-γR2/GFP and IFN-γR1/EBFP 1. Mutation of the Jak2 association site from FL-IFN-γR2/GFP did not significantly affect the ability of FL-IFN-γR2DJak2/GFP to interact with FL-IFN-γR2/EBFP (Figure 6A, black vs. purple lines) or with IFN-γR1/EBFP (Figure 6B, black vs. purple lines). These data indicate that Jak2 is not essential for preassembly of the receptor chains. Furthermore, IFN-γ did not change the interaction (i.e., decreased FRET) between FL-IFN-γR2DJak2/GFP and IFN-γR1/EBFP (Figure 6C) or between FL-IFN-γR2DJak2/GFP and FL-IFN-γR2/EBFP in the presence of IFN-γR1 (Figure 6D) as observed with the native receptors. Though Jak2 is not required for receptor complex preassembly, it (like Jak1) is required for ligand-mediated restructuring that occurs in the intact fluorescent IFN-γ receptor complex in the presence of IFN-γ. The predominant role of Jak2 in activating the IFN-γ receptor complex is consistent with previous data showing the requirement for its kinase activity to observe phosphorylation of Tyr-457 of IFN-γR1 and Stat1 22.
Mutation of the Jak2 association site on FL-IFN-γR2/GFP does not affect the interaction of IFN-γR2 with IFN-γR2 or with IFN-γR1. (A) COS-1 cells coexpressing FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP were illuminated with 760 nm light and the representative spectrum shown (black line). The purple line shows the emission spectrum obtained from COS-1 cells coexpressing FL-IFN-γR2 Jak2/GFP and FL-IFN-γR2/EBFP. (B) IFN-γR1/EBFP was coexpressed either with FL-IFN-γR2/GFP (black line) or with FL-IFN-γR2 Jak2/GFP (purple line). COS-1 cells expressing each receptor pair were illuminated with 760 nm light, and the two representative fluorescence emission spectra are shown. (C) COS-1 cells coexpressing FL-IFN-γR2DJak2/GFP and IFN-γR1/EBFP were treated with IFN-γ. Representative spectra were taken from cells prior to and after addition of IFN-γ (17 min) (black and red lines, respectively). (D) COS-1 cells coexpressing FL-IFN-γR2DJak2/GFP, FL-IFN-γR2/EBFP, and nonfluorescent IFN-γR1 were treated with IFN-γ (25 min). Representative spectra were taken from cells prior to and after addition of IFN-γ (black and red lines, respectively).
Determinants in the extracellular domains differentially mediate receptor interactions
The site-directed mutagenesis of the intracellular domains described above did not disrupt the interaction seen between two IFN-γR2 chains in intact cells. Thus we suspected that mutation of the extracellular domain of IFN-γR2 may modify the interaction of IFN-γR2 with other receptor chains. To address this issue we used chimeric receptors. Substitution of the intracellular domains of either human IFN-γR1 or IFN-γR2 with their corresponding murine sequence does not alter biological activity of the human IFN-γ receptor complex 30, 31, 36, 37. However, the extracellular domains of the human and murine IFN-γR1 and IFN-γR2 chains must originate from the same species for signal transduction to occur 4, 5, 30, 31, 33, 34, 35, 36, 37. Thus we hypothesized that since the human and murine extracellular domains of each chain must match; murine-human chimeric receptor chains would interact more weakly with human receptor chains than human receptors react with each other. To test this hypothesis directly, the extracellular domain of FL-Hu-IFN-γR2/GFP was substituted with the murine homolog to create the chimeric murine:human fluorescent receptor MH-IFN-γR2/GFP. After the chimeric receptor chain was coexpressed with a human receptor chain, the FRET between the pair was compared to the FRET between the corresponding human receptor chains. Substitution of the IFN-γR2 extracellular domain with the mouse sequence affected its interaction with human receptor chains.
When MH-IFN-γR2/GFP was coexpressed with Hu-IFN-γR1/EBFP, FRET between these chains was reduced significantly (Figure 7A, blue line) relative to FRET observed between Hu-FL-IFN-γR2/GFP and Hu-IFN-γR1/EBFP (Figure 7A, green line). Therefore species-specific determinants in the IFN-γR2 extracellular domain control interactions with IFN-γR1 molecules. Addition of Hu-IFN-γ to cells expressing MH-IFN-γR2/GFP and Hu-IFN-γR1/EBFP did not change the FRET between this receptor pair (data not shown).
The species-specificity barrier decreases FRET between IFN-γR1 and IFN-γR2 chains. (A) MH-IFN-γR2 exhibits lowFRET with Hu-IFN-γR1. Fluorescence emission spectra were taken from COS-1 cells coexpressing Hu-IFN-γR1/EBFP and FL-Hu-IFN-γR2/GFP (in green) or from COS-1 cells coexpressing Hu-IFN-γR1/EBFP and MH-IFN-γR2/GFP (in blue). (B) The species-specificity barrier disrupts the interaction between IFN-γR2 molecules. Fluorescence emission spectra taken from COS-1 cells coexpressing FL-Hu-IFN-γR2/EBFP and FL-Hu-IFN-γR2/GFP (in green) or COS-1 cells coexpressing FL-Hu-IFN-γR2/EBFP and MH-IFN-γR2/GFP (in blue).
The species-specificity barrier strongly affected interactions between IFN-γR2 chains. FRET between MH-IFN-γR2/GFP and FL-IFN-γR2/EBFP almost disappeared (Figure 7B, blue line), whereas coexpression of two human IFN-γR2 chains showed considerable FRET (Figure 7B, green line). Therefore, elements in the extracellular domain controlling the interaction between IFN-γR2 chains are species-specific. Coexpression of Hu-IFN-γR1 with MH-IFN-γR2/GFP and FL-IFN-γR2/EBFP only slightly enhanced the FRET between this receptor pair (data not shown). It follows that elements in the extracellular domain of Hu-IFN-γR2 whose sequences diverge between human and murine receptors mediate the homotypic interactions.
Discussion
Previous concepts about the structure of the IFN-γ receptor complex were based on coprecipitation assays 9, 10, 11, 19, 20. Jak1 and IFN-γR1 consistently coprecipitated with antibodies against the IFN-γR1 chain. Similarly, Jak2 and IFN-γR2 consistently coprecipitated with antibodies against IFN-γR2. However, coimmunoprecipitation of IFN-γR1 and IFN-γR2 was observed only in the presence of IFN-γ; similarly, coimmunoprecipitation of IFN-γR1 and Jak2 was observed only in the presence of IFN-γ. Thus the belief developed that IFN-γ assembled the signaling complex stepwise by first binding two IFN-γR1:Jak1 complexes and then two IFN-γR2:Jak2 complexes 6, 7, 44. However, all these experiments were based on methodologies that disrupt the plasma membrane and microenvironment of the IFN-γ receptor. To determine the gross structure of the IFN-γ receptor complex in intact cells we used FRET and found that the receptor complex is preformed 1, a result that indicated that the interaction between the IFN-γR1 and IFN-γR2 chains is too labile to survive immunoprecipitation. In this report we show that Jak1 binding to IFN-γR1 is required for the interaction between IFN-γR1 and IFN-γR2 (Figure 4) that has never been observed in coimmunoprecipitations or other methods. These results underscore the importance of noninvasive technologies to better understand how the IFN-γ receptor complex and other receptor complex exist in their natural state.
In this report we investigated the domains of the IFN-γ receptor complex that mediate receptor complex formation as well as ligand-induced changes in FRET 1 by using a method that does not disturb the microenvironment of the IFN-γ receptor in intact cells. The use of FRET permits determination of the proximity of donor and acceptor chains because FRET is only observed when two proteins are very close together; this proximity usually infers a direct interaction between the proteins attached to the donor and acceptor chains. These direct interactions were disrupted by various alterations to the receptor complex described in this report. This enabled us to discover that Stat1 must bind to IFN-γR1 to induce a significant change in ligand-dependant separation of the receptor chains; and that Jak1 but not Jak2 is required for preassembly of the IFN-γR1 and IFN-γR2 chains. Furthermore, interactions between IFN-γR2 chains required species-specific residues in their extracellular domains. Species-specific residues in the extracellular domain of IFN-γR2 also influence its interaction with IFN-γR1.
We mutated IFN-γR1 at Tyr-457 to prevent Stat1 recruitment to the IFN-γR1 chain. This mutation did not inhibit FRET between [Y457F]IFN-γR1/EBFP and FL-IFN-γR2/GFP in the absence of IFN-γ, but it eliminated the ligand-mediated decrease in FRET efficiency (Figure 1A, 1B). To confirm this observation independantly, U3A cells lacking Stat1 were transfected with IFN-γR1/ECFP and FL-IFN-γR2/EYFP. These stable transformants demonstrated that the IFN-γR1 and IFN-γR2 chains are preassociated but the FRET could not be decreased in response to Hu-IFN-γ (Figure 1C, 1D). Thus, Stat1 recruitment to IFN-γR1 produces the major IFN-γ-dependant change in separation between IFN-γR1 and IFN-γR2. Because this mutation on IFN-γR1 did not affect the interaction between two IFN-γR2 chains (Figure 2A, 2B), we conclude that the interaction of Stat1 with the carboxyl terminus of IFN-γR1 primarily affects the structure of IFN-γR1, while the movement of the two IFN-γR2 chains requires the activation of the preassembled IFN-γ receptor complex by IFN-γ.
The absence of Stat1 (U3A cells; Figure 1) or the Y457F mutation on IFN-γR1 did not compromise any other aspect of the IFN-γ receptor complex as reported: IFN-γ is capable of inducing several hundred genes when Stat1 is deleted 4647, 47 and the IFN-γ receptor is capable of inducing Stat1-independant genes in the presence of Stat1 when the IFN-γR1 chain contains the Y457F mutation 47. Furthermore, when Tyr-457 is mutated, the resulting receptor is still capable of tyrosine phosphorylation of Jak1, Jak2, and Tyr-479 of IFN-γR1, which creates an evolutionarily conserved (Figure 3) recruitment site for SOCS-1 48. All except the last activity are known to require the presence and/or activity of Jak kinases 22, 46. It can be inferred from these previous studies that the IFN-g receptor complex was structurally and biologically intact. Here, we extend these studies to confirm directly that the IFN-γ receptor complex is intact structurally when Stat1 activation by IFN-γ is disrupted (Figures 1 and 2).
Jak kinases bind to the intracellular domains of cytokine receptor chains, providing the IFN-γ receptor complex with tyrosine kinase activity necessary for the activation of Stat proteins. Cells lacking either Jak1 or Jak2 are unresponsive to IFN-γ 17, 18. To determine the roles of these kinases in the preassembly of the IFN-γ receptor complex, we mutated regions of the intracellular domain of IFN-γR1 or IFN-γR2 known to interact with Jak1 or Jak2, respectively, such that they could no longer interact with a Jak kinase. Mutation of the Jak2 binding site did not change the FRET significantly between FL-IFN-γR2/GFP and FL-IFN-γR2/EBFP (Figure 6A) or IFN-γR1/EBFP (Figure 6B). This suggests that mutagenesis of the Jak2 association site in the intracellular domain of IFN-γR2 does not grossly alter the structure of the intracellular domain of IFN-γR2 or the orientation of the EBFP and GFP moieties. Consequently, we conclude that Jak2 does not contribute to the preassembly of the chains of the IFN-γ receptor complex. In contrast, mutation of the Jak1 association site greatly diminished the FRET between the IFN-γR1 and IFN-γR2 chains (Figure 4A). Interestingly, ligand-dependant interactions were seen between two IFN-γR2 chains when the Jak1 site was mutated in IFN-γR1 (Figure 5B), though they were not as great as that seen in the presence of wild-type IFN-γR1 (Figure 2B). We hypothesize that Jak1 helps to facilitate the transient interaction between the extracellular domain of IFN-γR2 and the extracellular domain of either IFN-γR1 or the IFN-g:IFN-γR1 complex by tethering IFN-γR2 to IFN-γR1 in their intracellular domains. Thus Jak1 but not Jak2 has a major role in the preassembly of the IFN-γ receptor complex by greatly enhancing the interaction between IFN-γR1 and IFN-γR2. A structural requirement for Jak1, particularly the kinase-like domain for a functional IFN-γ receptor complex was reported previously 22 but the nature of its structural role necessary for biological activity was not determined nor could a structural role for Jak2 be assessed. Our studies show that Jak2 plays no role in receptor complex preassembly but is required for ligand-mediated restructuring and is consistent with previous data showing the requirement for Jak2 kinase activity to observe any signaling by IFN-γ 22. Because Jak2 did not influence receptor preassembly, the association of IFN-γR1 with IFN-γR2 by Jak1 does not occur through Jak2, but through IFN-γR2 directly or through another IFN-γR2-associated protein.
Because exchanging the extracellular domains of the IFN-γ receptor from human to mouse renders the IFN-γ receptor complex unable to bind or respond to Hu-IFN-g, we examined if these domain changes could affect the interactions among the IFN-γ receptor chains. We found that replacing the extracellular domain of Hu-IFN-γR1 with its murine counterpart did not greatly change the FRET between it and either Hu-IFN-γR1 or Hu-IFN-γR2 (data not shown), but exchange of the extracellular domain of IFN-γR2 with the murine sequence decreased the extent of FRET significantly between both Hu-IFN-γR1 and Hu-IFN-γR2 (Figure 7). Thus it appears that the species-specificity barrier due to IFN-γR2 affects its interaction with both IFN-γR2 and IFN-γR1 chains. The interaction of IFN-γ with IFN-γR1 is species specific 30, 33, 34. Thus, we believe that the overall species-specificity of murine and human IFN-γ for their receptors operates at two levels: binding of IFN-γ to IFN-γR1, and association of IFN-γR2 with IFN-γR1 and IFN-γR2 chains through their extracellular domains.
The results with the chimeric receptors suggested that interactions between two IFN-γR2 chains are mediated predominantly by their extracellular domains. It is reasonable to speculate that regions of the extracellular domain of IFN-γR2 whose primary sequences differ between human and murine IFN-γR2 chains participate in the interaction between IFN-γR2 and the IFN-γR1 and IFN-γR2 receptor chains. Comparisons of IFN-γR2 extracellular domains of humans, pigs, mice, chickens and frogs demonstrate that the residues predicted to be within β-pleated sheets are quite strongly conserved between the sequences, but greater variation occurs between the sequences in areas between the β-pleated sheets of the extracellular domain (Figure 3, right).
Loops between β-pleated sheets in the extracellular domain mediate the receptor:receptor interaction in the erythropoietin receptor as predicted from its crystal structure. The same loops of the fibronectin III domains are used by the erythropoietin receptor to engage two different types of unrelated erythropoietin ligand agonists, an erythropoietin receptor antagonist, and the erythropoietin receptor chains 49, 50, 51, 52. These binding loops (between sheets βC and βC′ and sheets βE and βF of the NH2-terminal SD100 domain, and after sheet βG in the 310 helix between the two SD100 domains) of IFN-γR2 may in fact be responsible for the interactions of IFN-γR2 with IFN-γR1 and with IFN-γR2.
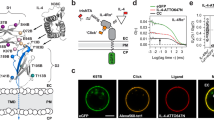
A model summarizing our data (Figure 8) is expanded from our initial model 1. The IFN-γ receptor complex is a preassembled entity (Figure 8A, B, C), as constitutive interactions are seen between two IFN-γR2 chains (Figure 6, Figure 8B), and between IFN-γR1 and IFN-γR2 (Figures 1, 4, 6, 7, 8A) in the absence of ligand. Within this receptor complex are interactions that are species specific and likely involving three loops of the extracellular domain of IFN-γR2)(Figure 8, purple shading). Within the intracellular domain (Figure 8D), IFN-γR1 and IFN-γR2 are held together by an interaction involving Jak1 (Figure 4), whereas Jak2 does not play an apparent role in receptor preassociation (Figure 6). Because the interaction between IFN-γR2 and IFN-γR1 is very weak when Jak1 is not present, we believe that there is considerable dynamic dissociation and association between the extracellular domains of IFN-γR1 and IFN-γR2 (Figure 8C); the IFN-γ receptor complex remains preassociated by the ability of Jak1 to tether IFN-γR2 to IFN-γR1 via their intracellular domains (Figure 8D). Changes in these interactions occur when IFN-γ binds to this preassembled IFN-γ receptor complex. These changes require that IFN-γR2, IFN-γR1 and IFN-γ be from closely related species. This requirement may arise because both receptor chains contact ligand directly. IFN-γR1 binds to IFN-γ directly 53, 54, 55. No evidence of a direct interaction of IFN-γ to IFN-γR2 alone exists, however IFN-γ can be crosslinked to IFN-γR2 in cells expressing IFN-γR1 11, 42, 56. Additionally, the interaction of IL-19, IL-20, and IL-24 with IL-20R1 (the orthologous ligand-binding chain of these ligands) requires the formation of a binary complex of these ligands with IL-20R2 (the orthologous accessory chain of these ligands) 57. The model proposes that first IFN-γ binds to the IFN-γR1 chains (Figure 8E, 8F), and then the IFN-γR2 chains interact with the IFN-γR1:IFN-γ:IFN-γR1 complex (Figure 8G, 8H). Jak1 keeps IFN-γR2 near IFN-γR1 so that a rapid association of IFN-γR2 to the IFN-γR1:IFN-γ:IFN-γR1 complex occurs. The Jak kinases then activate each other, and initiate the signaling cascade. Sometime after IFN-γR2 interacts with the IFN-γR1:IFN-γ:IFN-γR1 complex and probably after the Jak kinases are activated, the intracellular domains of the IFN-γR2 chains change their positions such that the FRET between them decreases (Figure 8I). The interaction of Stat1 with the IFN-γR1 chain results in a conformational change of the IFN-γR1 chain (Figure 8J) such that FRETS between IFN-γR1 and IFN-γR2 decreases.
Model of the IFN-γ receptor complex. Panels A, B, C, E, an G are views of the extracellular domains looking toward the cell membrane from above the receptor complex. Panels D, F, H, I and J are views from the plane of the plasma membrane. (A-D) The preassembled IFN-γ receptor complex in its resting state. IFN-γR1 is colored in red, IFN-γR2 in green, Jak1 in pink, and Jak2 in orange. Interactions mediated by species-specific determinants on IFN-γR2 are shown as purple shadings. The interaction of Jak1 and IFN-γR2 allows IFN-γR2 to remain associated with IFN-γR1 whether the extracellular domains of IFN-γR2 interact with those of IFN-γR1 (A), IFN-γR2(B), or are dissociated (C). (D) The constitutive interaction between the intracellular domains of IFN-γR2 and IFN-γR1 mediated by Jak1 is illustrated. (E, F) IFN-γ interacts with IFN-γR1 while the extracellular domains of the receptor complex are dissociated. IFN-γR2 remains associated with IFN-γR1 via Jak1. (G, H) IFN-γR2 associates with the IFN-γ:IFN-γR1 complex, possibly via a direct interaction with IFN-γ. This rearrangement allows the Jak kinases to become activated. (I) Initiation of signaling cascades. Phosphorylations (small black dots) occur when the Jak kinases are activated. At this stage, the IFN-γR2 intracellular domains change their conformation such that FRET between them decrease. (J) Stat1, dimerized through the NH2-terminal domains prior to its tyrosine phosphorylation 59, 60, 61, is recruited to the receptor when IFN-γR1 is phosphorylated; after its tyrosine phosphorylation and dissociation from the receptor complex, it forms SH2-mediated homodimers 62.
To summarize, the IFN-γ-dependant movement between the IFN-γR1 and IFN-γR2 chains required that Stat1 bind IFN-γR1. Mutation of the Jak1 association site inhibited the interaction between IFN-γR1 and IFN-γR2, indicating that Jak1 is necessary for the interaction of these chains. Mutation of the Jak2 association site on IFN-γR2 has no effect on receptor preassociation, suggesting there is no obligatory role for Jak2 in preassembly of the receptor complex. Exchange of the extracellular domain of human IFN-γR2 from the human to the murine sequence disrupts its interactions with human IFN-γR2 and with human IFN-γR1 chains, implying that IFN-γR2 uses species-specific determinants in these interactions. Finally, interaction of the IFN-γ receptor complex with IFN-γ results in two conformational changes dependant on both Jak1 and Jak2. One involves a spreading of the two IFN-γR2 chains from each other. The second involves a Stat1-mediated movement of the IFN-γR1 chains from the IFN-γR2 chains.
References
Krause CD, Mei E, Xie J, et al. Seeing the light: preassembly and ligand-induced changes of the interferon g receptor complex in cells. Mol Cell Proteomics 2002; 1:805–15.
Rashidbaigi A, Langer JA, Jung V, et al. The gene for human interferon receptor is located on chromosome 6. Proc Natl Acad Sci USA 1986; 83:384–8.
Jung V, Rashidbaigi A, Jones C, et al. Human chromosomes 6 and 21 are required for sensitivity to human interferon-γ. Proc Natl Acad Sci USA 1987; 84:4151–5.
Soh J, Donnelly RJ, Kotenko S, et al. Identification and sequence of an accessory factor required for activation of the human interferon-γ receptor. Cell 1994; 76:796–803.
Hemmi S, Böhni R, Stark G, et al. A novel member of the interferon receptor family complements functionality of the murine interferon γ receptor in human cells. Cell 1994; 76:803–10.
Pestka S, Kotenko SV, Muthukumaran G, et al. The interferon γ (IFN-γ) receptor: a paradigm for the multichain cytokine receptor. Cytokine Growth Factor Rev 1997; 8:189–206.
Stark GR, Kerr IM, Williams BR, et al. How cells respond to interferons. Annu Rev Biochem 1998; 67:227–64.
Pestka S, Krause CD, Sarkar D, et al. Interleukin-10 and related cytokines and receptors. Annu Rev Immunol 2004; 22:929–79.
Igarashi K, Garotta G, Ozmen L, et al. Interferon-γ induces tyrosine phosphorylation of interferon-γ receptor and regulated association of protein tyrosine kinases, Jak1 and Jak2, with its receptor. J Biol Chem 1994; 269:14333–6.
Sakatsume M, Igarashi K, Winestock KD, et al. The Jak kinases differentially associate with the α and β (accessory factor) chains of the interferon γ receptor to form a functional receptor capable of activating STAT transcription factors. J Biol Chem 1995; 270:17528–34.
Kotenko SV, Izotova LS, Pollack BP, et al. Interaction between the components of the interferon γ receptor complex. J Biol Chem 1995; 270:20915–21.
Marsters SA, Pennica D, Bach E, et al. Interferon γ signals via a high-affinity multisubunit receptor complex that contains two types of polypeptide chain. Proc Natl Acad Sci USA 1995; 92:5401–5.
Greenlund AC, Farrar MA, Viviano BL, Schreiber RD . Ligand-induced IFN γ receptor tyrosine phosphorylation couples the receptor to its signal transduction system (p91). EMBO J 1994; 13:1591–600.
Farrar MA, Fernandez-Luna J, Schreiber RD . Identification of two regions within the cytoplasmic domain of the human interferon-γ receptor required for function. J Biol Chem 1991; 266:19626–35.
Cook JR, Jung V, Schwartz B, et al. Structural analysis of the human interferon γ receptor: a small segment of the intracellular domain is specifically required for class I major histocompatibility complex antigen induction and antiviral activity. Proc Natl Acad Sci U S A 1992; 89:11317–21.
Greenlund AC, Morales MO, Viviano BL, et al. Stat recruitment by tyrosine-phosphorylated cytokine receptors: an ordered reversible affinity-driven process. Immunity 1995; 2:677–87.
Watling D, Gushin D, Muller M, et al. Complementation by the protein tyrosine kinase JAK2 of a mutant cell line defective in the interferon-γ signal transduction pathway. Nature 1993; 366:166–70.
Müller M, Briscoe J, Laxton C, et al. The protein tyrosine kinase JAK1 complements defects in interferon-α/β and -γ signal transduction. Nature 1993; 366:129–35.
Kaplan DH, Greenlund AC, Tanner JW, et al. Identification of an interferon-γ receptor a chain sequence required for JAK-1 binding. J Biol Chem 1996; 271:9–12.
Bach EA, Tanner JW, Marsters S, et al. Ligand-induced assembly and activation of the γ interferon receptor in intact cells. Mol Cell Biol 1996; 16:3214–21.
Usacheva A, Sandoval R, Domanski P, et al. Contribution of the Box 1 and Box 2 motifs of cytokine receptors to Jak1 association and activation. J Biol Chem 2002; 277:48220–6.
Briscoe J, Rogers NC, Witthuhn BA, et al. Kinase-negative mutants of JAK1 can sustain interferon-γ-inducible gene expression but not an antiviral state. EMBO J 1996; 15:799–809.
Tanner JW, Chen W, Young RL, et al. The conserved box 1 motif of cytokine receptors is required for association with JAK kinases. J Biol Chem 1995; 270:6523–30.
Zhao Y, Wagner F, Frank SJ, Kraft AS . The amino-terminal portion of the JAK2 protein kinase is necessary for binding and phosphorylation of the granulocyte-macrophage colony-stimulating factor receptor bc chain. J Biol Chem 1995; 270:13814–8.
Frank SJ, Yi W, Zhao Y, et al. Regions of the JAK2 tyrosine kinase required for coupling to the growth hormone receptor. J Biol Chem 1995; 270:14776–85.
Cacalano NA, Migone TS, Bazan F, et al. Autosomal SCID caused by a point mutation in the N-terminus of Jak3: mapping of the Jak3-receptor interaction domain. EMBO J 1999; 18:1549–58.
Haan C, Is'harc H, Hermanns HM, et al. Mapping of a region within the N-terminus of Jak1 involved in cytokine receptor interaction. J Biol Chem 2001; 276:37451–8.
Duhé RJ, Farrar WL . Characterization of active and inactive forms of the JAK2 protein-tyrosine kinase produced via the baculovirus expression vector system. J Biol Chem 1995; 270:23084–9.
Gauzzi MC, Velazquez L, McKendry R, et al. Interferon-γ-dependent activation of Tyk2 requires phosphorylation of positive regulatory tyrosines by another kinase. J Biol Chem 1996; 271:20494–500.
Hibino Y, Kumar CS, Mariano TM, et al. Chimeric interferon g receptors demonstrate that an accessory factor required for activity interacts with the extracellular domain. J Biol Chem 1992; 267:3741–9.
Böhni R, Hemmi S, Aguet M . Signaling steps involving the cytoplasmic domain of the interferon-γ receptor α-subunit are not species-specific. J Biol Chem 1994; 269:14541–5.
Jung V, Jones CA, Rashidbaigi A, et al. Chromosome mapping of biological pathways by fluorescence-activated cell sorting and cell fusion: human interferon-γ receptor as a model system. Somatic Cell Mol Genet 1988; 14:183–92.
Gray PW, Leong S, Fennie EH, et al. Cloning and expression of the cDNA for the murine interferon γ receptor. Proc Natl Acad Sci U S A 1989; 86:8497–501.
Hemmi S, Peghini P, Metzler M, et al. Cloning of murine interferon γ receptor cDNA: expression in human cells mediates high-affinity binding but is not sufficient to confer sensitivity to murine interferon γ. Proc Natl Acad Sci U S A 1989; 86:9901–5.
Hibino Y, Mariano TM, Kumar CS, et al. Expression and reconstitution of a biologically active mouse interferon γ receptor in hamster cells. Chromosomal location of an accessory factor. J Biol Chem 1991; 266:6948–51.
Gibbs VC, Williams SR, Gray PW, et al. The extracellular domain of the human interferon γ receptor interacts with a species-specific signal transducer. Mol Cell Biol 1991; 11:5860–6.
Muthukumaran G, Donnelly RJ, Ebensperger C, et al. The intracellular domain of the second chain of the interferon-γ receptor is interchangeable between species. J Interferon Cytokine Res 1996; 16:1039–45.
Izotova LS, Kotenko SV, Mirochnitchenko OV, Pestka S . Cloning and functional characterization of chinese hamster IFN-γ. Eur Cyt Network 1998; 9:418.
McKendry R, John J, Flavell D, et al. High-frequency mutagenesis of human cells and characterization of a mutant unresponsive to both a and g interferons. Proc Natl Acad Sci U S A 1991; 88:11455–9.
Pluthero FG . Rapid purification of high-activity Taq DNA polymerase. Nucleic Acids Res 1993; 21:4850–1.
Kotenko SV, Krause CD, Izotova LS, et al. Identification and characterization of a second chain of the interleukin-10 receptor complex. EMBO J 1997; 16:5894–903.
Kotenko SV, Izotova LS, Pollack BP, et al. Other kinases can substitute for Jak2 in signal transduction by interferon-γ. J Biol Chem 1996; 271:17174–82.
Bopp MA, Sytnik A, Howard TD, et al. The dynamics of structural deformations of immobilized single light-harvesting complexes. Proc Natl Acad Sci U S A 1999; 96:11271–6.
Bach EA, Aguet M, Schreiber RD . The IFNg receptor: a paradigm for cytokine receptor signaling. Annu Rev Immunol 1997; 15:563–91.
Farrar MA, Campbell JD, Schreiber RD . Identification of a functionally important sequence in the C terminus of the interferon-g receptor. Proc Natl Acad Sci U S A 1992; 89:11706–10.
Gil MP, Bohn E, O'Guin AK, et al. Biologic consequences of Stat1-independent IFN signaling. Proc Natl Acad Sci U S A 2001; 98:6680–5.
Ramana CV, Gil MP, Han Y, et al. Stat1-independent regulation of gene expression in response to IFN-γ. Proc Natl Acad Sci U S A 2001; 98:6674–9.
Qing Y, Costa-Pereira AP, Watling D, Stark GR . Role of tyrosine 441 of interferon-g receptor subunit 1 in SOCS-1-mediated attenuation of STAT1 activation. J Biol Chem 2005; 280:1849–53.
Syed RS, Reid SW, Li C, et al. Efficiency of signalling through cytokine receptors depends critically on receptor orientation. Nature 1998; 395:511–6.
Livnah O, Johnson DL, Stura EA, et al. An antagonist peptide-EPO receptor complex suggests that receptor dimerization is not sufficient for activation. Nat Struct Bio 1998; 5:993–1004.
Middleton SA, Barbone FP, Johnson DL, et al. Shared and unique determinants of the erythropoietin (EPO) receptor are important for binding EPO and EPO mimetic peptide. J Biol Chem 1999; 274:14163–9.
Livnah O, Stura E, Middleton SA, et al. Crystallographic evidence for preformed dimers of erythropoietin receptor before ligand activation. Science 1999; 283:987–90.
Fountoulakis M, Zulauf M, Lustig A, Garotta G . Stoichiometry of interaction between interferon g and its receptor. Eur J Biochem 1992; 208:781–7.
Walter MR, Windsor WT, Nagabhushan TL, et al. Crystal structure of a complex between interferon-γ and its soluble high-affinity receptor. Nature 1995; 376:230–5.
Thiel DJ, le Du MH, Walter RL, et al. Observation of an unexpected third receptor molecule in the crystal structure of human interferon-γ receptor complex. Structure Fold Des 2000; 8:927–36.
Krause CD, Lunn CA, Izotova LS, et al. Studies on signaling of covalent heterodimers of interferon-γ: evidence for one-sided signaling in the active tetrameric receptor complex. J Biol Chem 2000; 275:22995–3004.
Pletnev S, Magracheva E, Kozlov S, et al. Characterization of the recombinant extracellular domains of human interleukin-20 receptors and their complexes with interleukin-19 and interleukin-20. Biochemistry 2003; 42:12617–24.
Braunstein J, Brutsaert S, Olson R, Schindler C . STATs dimerize in the absence of phosphorylation. J Biol Chem 2003; 278:34133–40.
Ota N, Brett TJ, Murphy TL, et al. N-domain-dependent nonphosphorylated STAT4 dimers required for cytokine-driven activation. Nat Immunol 2004; 5:208–15.
Mao X, Ren Z, Parker GN, et al. Structural bases of unphosphorylated STAT1 association and receptor binding. Mol Cell 2005; 17:761–71.
Chen X, Vinkemeier U, Zhao Y, et al. Crystal structure of a tyrosine phosphorylated STAT-1 dimer bound to DNA. Cell 1998; 93:827–39.
Acknowledgements
We would like to thank Ellen Feibel for assistance in preparation of this manuscript, Roger Y Tsien for the plasmid encoding S65T, and Jerome Langer for the plasmid encoding EGFP. This work was supported by NIH grants from the National Institute of Allergy and Infectious Diseases R01-AI043369, R01-AI36450 and R01-AI059465.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Krause, C., Lavnikova, N., Xie, J. et al. Preassembly and ligand-induced restructuring of the chains of the IFN-γ receptor complex: the roles of Jak kinases, Stat1 and the receptor chains. Cell Res 16, 55–69 (2006). https://doi.org/10.1038/sj.cr.7310008
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310008
Keywords
This article is cited by
-
The interferons and their receptors—distribution and regulation
Immunology & Cell Biology (2012)
-
Effect of amino acid substitutions in the human IFN-γR2 on IFN-γ responsiveness
Genes & Immunity (2011)
-
Modulation of the activation of Stat1 by the interferon-γ receptor complex
Cell Research (2006)