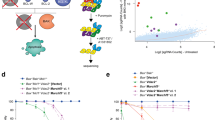
Abstract
The transcription factor E2F1 does not only induce cell proliferation but also shows the strongest proapoptotic effect of all E2F family members as part of an antitumor safeguard mechanism. We have recently identified KIAA0767 as a novel p53-independent target of E2F1. Here, we investigated the biological function of interaction. Overexpression studies of KIAA0767, termed D(eath)-I(nducing)-P(rotein), revealed its strong proapoptotic effect. DIP greatly reduced cell viability in several in vitro systems accompanied by typical apoptotic features such as caspase-3 activation and cleavage of poly(ADP-ribose)-polymerase. Endogenous DIP levels increased following E2F1 activation. Yet, inhibition of endogenous DIP function by small interfering RNA rescued p53-negative cells from E2F1-induced apoptosis, indicating that DIP is an essential mediator of the p53-independent E2F1 death pathway. Localization studies showed that DIP localizes to the mitochondria, where endogenous DIP is upregulated following E2F1 induction. These results provide new insights to the incompletely understood regulatory mechanisms of E2F1-induced apoptosis.
Similar content being viewed by others
Introduction
The transcription factor E2F1 regulates gene expression of a vast number of proteins and plays a central role in cell cycle regulation. E2F1 is the best-examined factor of the seven structurally related E2F transcription factor family members E2F1–E2F7. All E2F proteins except E2F7 function in heterodimeric complexes with DP family (DP1 or DP2) proteins transactivating or repressing gene expression. E2F activity is controlled in many ways,1 including interaction with the retinoblastoma gene product (Rb). Hypophosphorylated Rb and its pocket protein family members p107 and p130 repress E2F activity2, 3 by shielding E2Fs transactivation domain leading to restriction of cell cycle progression.4, 5 The E2F family consists of three subgroups according to their pocket protein family binding affinity and transcriptional activation. E2F1, E2F2, and E2F3 exclusively bind Rb2 and are strong transcriptional activators. The second group seems to be involved in active repression of E2F target genes6 whose members E2F4 can associate with all pocket proteins7, 8 (Rb, p107, and p130), while E2F5 solely binds p130.9 2A7E, the third group, lacks a transactivation domain and does not interact with any pocket protein10, 11 but negatively regulates E2F-responsive transcription by complexation with chromatin modifiers.12, 13 The newest family member E2F7 has recently been described as a factor capable of blocking cellular proliferation.14
Phosphorylation of the pocket proteins by cyclin–cdk (cyclin-dependent kinase)complexes15, 16 in late G1 phase leads to release from E2Fs and S-phase entry. Deregulated E2F activity due to disruption of the E2F-Rb pathway has been observed in many human tumors as a result of different oncogenic changes. Amplification of cyclin D or loss of the cdk inhibitor p16 leads to accelerated phosphorylation of Rb. Loss of Rb or expression of viral oncoproteins such as the human papilloma virus protein E7 also results in disruption of the E2F/Rb complex and leads to unrestrained E2F activity.15
Although several reports describe the inevitability of E2F1 for cell cycle progression, a large number of studies clearly implicates the role of this transcription factor for the induction of apoptosis. Under deregulated conditions many studies strongly support the notion of E2F1 as a tumor suppressor rather than an oncogene.17, 18, 19 In mice lacking E2F1, a high incidence of tumor development was observed further supporting this role of E2F1.20 Recently, the marked box domain has been identified as the essential sequence for apoptotic activity of E2F1.21 Apoptosis induction by E2F1 is initiated via p53-dependent pathways and independently of the p53 tumor suppressor. p53 accumulates following E2F1 expression22 through activation of p14ARF which in turn interacts with mouse double minute 2 (Mdm2), thereby inhibiting p53 ubiquitination and degradation.23, 24 Lately, additional ADP ribosylation factor 1 (ARF)-independent p53-mediated pathways have been described25, 26 and some reports even imply a negative feedback of ARF since it seems to target E2F1 for degradation.27 Furthermore, in the absence of ARF, p53 is phosphorylated by E2F1 presumably via ATM (ataxia telangiectasia mutated) and/or ATR protein kinases since this modification is abolished in the presence of caffeine.28
Additionally, tissue culture and mouse experiments show that E2F1-induced apoptosis occurs independently of p53.29, 30, 31 E2F1, for example, transactivates the gene encoding the apoptosis protease-activating factor 1 (Apaf1).32 This initiates the assembly of Apaf1 with cytochrome c (cyt c) followed by procaspase-9 activation and successive initiation of proapoptotic effector caspases such as caspase-3. We and others have previously reported that the p53-homologue p73 partially mediates E2F1-induced apoptosis independently of p53.33, 34 Yet, data from microarray analysis suggest the involvement of proapoptotic Bcl-2 family members on the p53-autonomous death pathway.35, 36 Altogether, E2F1-induced apoptosis utilizes various pathways, p53-dependent as well as independently of p53, reflecting its key role in initiating the cell death machinery. Nevertheless, the molecular mechanisms underlying E2F1-induced apoptosis are not fully understood.
Thus, many studies recently made use of microarray technology to identify novel targets and/or networks of E2F1-regulated downstream effectors36, 37, 38 that are possibly involved in programmed cell death. We have previously reported the identification of KIAA0767 as a novel E2F1 target gene found to be upregulated on messenger RNA (mRNA) level in Saos-2 cells that conditionally activate E2F1 by addition of 4-hydroxytamoxifen (4-OHT).36 Here, we explored the biological function of the so far undescribed E2F1 target expressed by an adenoviral (Ad) vector in an established p53-null cell system. KIAA0767 overexpression lead to reduced cell viabilty and elevated sub-G1 cell populations with classical proapoptotic features including chromatin condensation, membrane blebbing, and nucleosomal DNA fragmentation (the KIAA0767 protein was thus termed death-inducing protein (DIP)). DIP-induced apoptosis is accompanied by the activation of caspase pathway enzymes and poly(ADP-ribose)-polymerase (PARP) cleavage. Furthermore, we demonstrated the involvement of endogenous DIP in E2F1-mediated apoptotic pathways. Levels of endogenous DIP protein increased following an E2F1 stimulus and selective small interfering RNA (siRNA) interference of DIP expression elucidated a decrease of E2F1-related apoptosis, clearly demonstrating its role as a mediator of E2F1-induced apoptosis in the absence of p53. Subcellular localization studies revealed that DIP localizes to the mitochondria and is substantially upregulated in the mitochondrial fraction following E2F1 activation.
Results
DIP is transcriptionally and translationally activated by E2F1 initiation
We have previously reported the upregulation of many genes following E2F1 activation on a cDNA microarray.36 The KIAA0767 gene product whose function has been so far unknown showed a substantial increase on mRNA level following E2F1 induction. The biological relevance of KIAA0767 upregulation, here termed DIP, and its potential role in E2F1 signaling, was then analyzed in detail. A human p53-negative osteosarcoma Saos-2 cell line that stably expresses the ERE2F1 fusion protein has been previously decribed34 and was used to conditionally regulate E2F1 activation through 4-OHT administration. DIP transcript levels peaked at around 20 h after E2F1 induction (Figure 1a) compared to nondetectable mRNA levels in cells lacking activated E2F1.
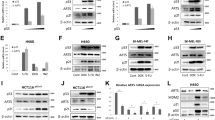
DIP induction by E2F1. (a) Serum-starved Saos-2 cells stably transfected with ERE2F1 were grown in the presence of 4-hydroxy-tamoxifen (4-OHT) at the indicated times followed by RNA isolation. Semiquantitative RT-PCR analysis of DIP was carried out under linear amplification conditions using specific primer pairs. p73 and cyclin E were used as positive controls. E2F1 mRNA levels are shown as a negative control. The data were normalized to GAPDH values. (b) Western blot of 4-OHT-treated Saos-2 ERE2F1 cells and parental Saos-2, and H1299 cells infected with Ad-ERE2F1. Equal amounts of whole-cell extracts were assayed for DIP, ERE2F1 and endogenous E2F1 protein. Actin was used as a loading control
To check whether endogenous DIP protein expression is also enhanced upon E2F1 activation, a custom made antibody against the epitope lrrldkirfrghkrdd representing amino acids 2–16 of the protein (for complete protein sequence, see supplemental data) was produced by Eurogentec (Seraing, Belgium). A double band of the protein of ∼55 kD size indicative of increased DIP levels was detected as early as 24 h after E2F1 inititiation in both the inducible Saos-2 cell line and in Saos-2 and H1299 small lung carcinoma cells infected with an Ad vector expressing ERE2F1 in the presence of 4-OHT (Figure 1b). The extent of DIP protein expression directly correlated with the amount of activated fusion protein. In both Ad-ERE2F1-infected cell lines, the signal of the ectopic ERE2F1 is too strong to allow detection of endogenous E2F1 on the same immunoblot.
DIP reduces cell viability of tumor cells and normal fibroblasts
We have previously described the use of recombinant adenoviral vectors as a means to transfer transgenes into target cells with an efficiency that allows a functional analysis of the produced protein in the entire population of cells.39, 40 Therefore, we constructed an adenovirus expressing the DIP gene product (Ad-Dip; Figure 2a). Several in vitro systems were assayed for cell viability 4 days after infection with Ad-Dip, or virus expressing the tumor suppressor p53 (Ad-p53wt), or the green fluorescent protein (Ad-GFP), respectively. As shown in Figure 2b, compared to control vector-infected cells, which were set as 100%, DIP protein expression resulted in a significant loss of cell viability by approximately 80–90% in all tested cell lines independent of the endogenous p53 status. In contrast to p53, DIP's killing potential is not restriced to tumor cells (as shown in karyotypically normal VH6 fibroblasts).
Analysis of Dip-mediated cytotoxicity in human cancer cells and primary fibroblasts. (a) Schematic representation of the Ad-Dip vector. (b) Cell viability was measured 96 h after infection with Ad-GFP (white), Ad-p53 wild type (gray) or Ad-Dip (black) by MTT assay. Ad-GFP-infected cells were set as 100%. The number of viable cells is the mean±S.D. of three different wells; bars±S.D. Triplicate experiments were performed for each cell line
DIP-mediated cell death is caused by apoptosis
To clarify whether the observed loss of viability upon DIP overexpression is due to apoptosis, cells were analyzed for their cell cycle profiles by FACS (fluorescence-activated cell sorter) (Figure 3). Compared to Ad-GFP-transduced cells, Ad-Dip-infected H1299 cells showed a significant increase of sub-G1 cells (M1) indicative of apoptosis, as early as 24 h postinfection (hpi). At 72 h after infection, the majority of DIP-expressing cells (∼70%) are shifted towards the apoptotic sub-G1 population similar to cells which overexpress p53. Ad-p53wt-infected cells illustrated this effect more pronounced with enhanced portions of apoptotic cells in sub-G1 compared to Ad-Dip-infected cells emphasizing a faster apoptosis induction.
DIP expression in these cells produced typical apoptotic features with remarkable changes in the cellular and nuclear morphology, characterized by membrane blebbing, cell shrinkage, intense staining of condensed chromatin, and nucleosomal DNA fragmentation41, 42, 43 as analyzed by Hoechst 33342 staining and laser scan microscopy (Figure 4). The described morphologic changes appeared in H1299 cells that overexpress DIP between 48 and 72 h after infection, compared to p53-expressing cells, where apoptotic bodies appeared 24 h earlier followed by giant cells which completely detached from the surface of the cell culture dish at 72 h after infection. In contrast, cells infected with the GFP vector grew normally without any morphological changes. Analysis of genomic DNA from Ad-Dip infected H1299 cells revealed typical nucleosomal fragments at 3 days after infection (data not shown).
Analysis of cellular integrity of H1299 cells following DIP expression. Cells infected with Ad-GFP, Ad-p53wt or Ad-Dip vector for different time points were stained with Hoechst 33342 and analyzed by laser scan microscopy at 20-fold magnification. Ad-Dip and Ad-p53wt-expressing cells showed characteristic proapoptotic features. Results are shown from one of two independent experiments
Caspases are essential components of the apoptotic machinery. Once activated by cleavage, they trigger a cascade that leads to morphological changes associated with apoptosis.44 To initially characterize the signaling pathway implicated in DIP-mediated apoptosis, we examined the activation of caspase-3 and PARP cleavage as key events in the apoptotic pathway. As demonstrated in Figure 5a, increased expression of DIP in H1299 cells resulted in caspase-3 activation, as revealed by the appearance of an ∼17 kDa product which corresponds to the 17 kDa subunit of activated caspase-3. The observed caspase-3 activation could be completely blocked by the wide-spectrum caspase inhibitor, zVAD-fmk (N-benzyloxycarbonyl-Val-Ala-Asp-fluoromethylketone), on the protein level (Figure 5b). Consistenly, the apoptotic activity of DIP in p53-null H1299 cells, measured by flow cytometry at 48 h after infection, was significantly antagonized by the pan-caspase inhibitor, leading to a ∼33% reduction of DIP-induced apoptosis (Figure 5c). As shown in Figure 6, exposure of cells to DIP was also accompanied by cleavage of the 116 kDa intact form of PARP to the 89 kDa fragment intimately linked to the induction of apoptosis. Both in Saos-2 and H1299 cells infected with Ad-Dip, PARP cleavage became evident at 24–72 hpi with full-length PARP greatly decreasing. Again, Ad-p53-infected cells showed faster kinetics of apoptosis induction illustrated by the appearance of cleaved PARP within 12 h in both cell lines. PARP was not affected by GFP expression.
Apoptosis induction by DIP involves caspase-3 activation in H1299 cells. (a) Equal amounts of total cell extracts from cells infected with Ad-Dip over 96 h were separated by SDS-PAGE and analyzed by Western blotting using an antibody detecting the 17 kDa subunit of cleaved caspase-3 (Casp-3). Actin was used as a loading control. (b) The inhibition of DIP-mediated caspase-3 activation by the caspase inhibitor zVAD-fmk (50 μM) was analyzed by Western blotting of whole-cell extracts and (c) quantitated by flow cytometry. Apoptosis (as determined by cells with a sub-G1 DNA content) 48 h after infection is as indicated. Each graph represents the mean±S.D. of three independent experiments
Cleavage of PARP in DIP-expressing cells. Western blot analysis of (a) H1299 cells and (b) Saos-2 cells infected with Ad-Dip, Ad-GFP, or Ad-p53wt at various time points after virus administration. Full-length PARP and the 89 kDa proteolytic cleavage product are indicated. DIP protein forms four distinct bands between 50 and 64 kDa. Actin was used as a loading control
Structural analysis and mapping of functional domains
The DIP encoding gene alias KIAA0767 (GenBank Accession number XM_027105) localizes to chromosome 22q13.31. (NC_000022; GeneID 23151). The complete cDNA of 1737 base pairs (bp) in length encodes a single open reading frame (ORF) of 578 amino acids (Figure 7). The predicted DIP protein has a calculated molecular weight of approximately 62 kDa. Mapping by BLAST database searches revealed a nuclear localization signal in the N-terminal region of the protein comprising amino acids 110–121, at least two transmembrane regions encompassing amino acids 246–268 and 345–367, and a GRAM domain at the C-terminus often found in membrane-associated proteins such as glycosyltransferases and myotubularins. Moreover, our database searches (http://www.expasy.org/tools/) predicted multiple putative phosphorylation and glycosylation sites including a tyrosine-kinase phosphorylation site at position 383–390, three myristoylation sites, and one glycosaminoglycan attachment region (aa 531–534; compare Table 1 and Figure 7), which argues for a vast number of modified forms of the protein presumably in different cellular contexts or subcellular structures. Furthermore, the TargetP server v1.01 database for subcellular localization predicted a 40% presumption for the first 130 N-terminal amino-acid residues and a probability of 76% for amino acids 375–500 of being localized in the mitochondria. Conversely, the probability for amino acids 248–375 being part of the secretory pathway was predicted as 93%. As shown in Figure 6, exposure of cells to adenovirally expressed DIP gives rise to four distinct bands between 50 and 64 kDa, whereas endogenous DIP appears as a double band of ∼55 kDa (compare Figure 1b). However, among the multiple bands of ectopically expressed DIP, the strongest band corresponds to the endogenous DIP band detectable in the immunoblot of Figure 1b, suggesting that the 55 kDa protein is the full-length form, whereas the additional bands of higher molecular weight are likely due to post-translational modifications of the full-length protein. This typical multiple band protein pattern was also confirmed when a His-tagged DIP protein version was analyzed in immunoblots with anti-His epitope antibodies (data not shown), arguing for the specificity of the bands.
Schematic structure of the DIP encoding gene on chromosome 22q13.31 and its functional domains. The coding sequence of DIP spans over 17 exons resulting in a protein of 578 amino acids. Several database predicted domains, such as a nuclear localization signal, transmembrane domains and a GRAM domain, and multiple putative protein modification sites are indicated
DIP is localized in the mitochondria and becomes markedly upregulated herein during E2F1-induced apoptosis
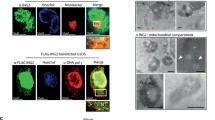
While our custom made antibody raised against DIP was not useful for protein detection of fixed cells, the DIP cDNA was cloned into the pcDNA 3.1/myc-His. vector that efficiently expresses His-tagged DIP protein. After transfection of this plasmid into H1299 cells, the subcellular localization of DIP was examined by immunofluorescence and confocal laser scanning microscopy using an antibody directed against His (Figure 8a). In pcDNA 3.1-DIP/myc-His-transfected cells, DIP was stained in a perinuclear punctate pattern indicative of intramitochondrial localization (Figure 8a, VI, VIII). Intramitochondrial localization of DIP was confirmed by colabeling with a mitochondrion-selective marker (Figure 8a, VII, VIII). No signal other than the mitochondrial green was observed in control-transfected cells (Figure 8a, II–IV). To confirm the immunofluorescence data, we analyzed the mitochondrial fraction of H1299 cells for DIP expression after Ad-Dip infection. As demonstrated in Figure 8b, ectopic DIP protein strongly appears in the mitochondria at 24 h after vector treatment. The purity of the fractions was ascertained by reprobing the immunoblot with antibodies against the mitochondrial specific marker protein Cox4. Since DIP is a downstream target of E2F1, we also investigated the effect of E2F1 activation on the pattern of endogenous DIP distribution in these cells. As shown in Figure 8c, DIP abundance is markedly upregulated in the mitochondria between 24 and 48 h peaking at 24 h in response to increased E2F1 stimulation.
Subcellular localization of DIP protein. (a) H1299 cells were transfected with pcDNA3.1-DIP/myc-His-expressing His-tagged full-length DIP and the control vector pDNA3.1/myc-His. Cells were fixed and stained for His-tagged DIP using anti-His and Alexa Fluor 546 antibodies (II and VI). Nuclei and mitochondria were stained by DAPI (I and V) and MitoTracker Green FM (III and VII), respectively. Cells were assayed for laser scan microscopy (magnification 63X). (b, c) Western blot analysis of ectopic (b) or endogenous (c) DIP in the mitochondrial fraction at various time points after transient infection of H1299 cells with Ad-Dip and Ad-ERE2F1, respectively. Equal amounts of proteins were assayed. Cox4 is shown as a mitochondrial marker
E2F1-induced apoptosis is abrogated by DIP suppression
Strikingly, endogenous DIP is substantially upregulated in response to E2F1 activity and increased levels of ectopic DIP result in the induction of apoptosis. Thus, activation of endogenous DIP might contribute to E2F1-induced apoptosis. Reasoning that inhibition of physiological DIP activity would block E2F1-induced apoptosis, we examined whether the E2F1 death promoting function is compromised in DIP-defective cells. Ad-ERE2F1-transduced H1299 cells, which showed strongly elevated DIP transcript levels at 12–20 h after 4-OHT stimulation (Figure 9a), were coinfected with a DIP siRNA expression vector (Ad-siDip). Effective inhibition of high-level DIP expression by this construct was demonstrated by reverse transcription-polymerase chain reaction (RT-PCR) analysis (Figure 9b). Increased amounts of siRNA against DIP completely abolished DIP transcription at the time point of maximum E2F1 induction, whereas DIP mRNA expression of cells infected with the unspecific control Ad-siGFP was not affected. To analyze the effect of DIP inhibition on E2F1-induced cytotoxicity, H1299 cells coexpressing activated ERE2F1 and siDip were analyzed by flow cytometry, as indicated in Figure 9c. FACS profiles revealed a substantial decrease of E2F1-related cell killing when the function of DIP was blocked. Expression of siDip resulted in a significant reduction of apoptotic cells to a level approximately 50% of that seen in H1299 cells expressing activated ERE2F1 with intact DIP function (P=0.046 at day 2). In contrast, E2F1-mediated apoptosis was not abolished by the control vector encoding siGFP. This result clearly indicates that inhibition of DIP function at least partially rescues cells from E2F1-induced apoptosis.
Reduction of E2F1-induced cell function by inhibition of DIP. (a, b) Semiquantitative RT-PCR analysis of DIP transcript levels in Ad-ERE2F1-infected H1299 cells at various time points after 4-OHT treatment in the absence (a) and presence of ectopically expressed DIP siRNA (Ad-siDip) compared to cells coinfected with Ad-siGFP (b). GAPDH is used as control. A schematic structure of siRNA expressing Ad-vectors is shown. (c) H1299 cells treated as in (b) for 1 and 2 days were analyzed by flow cytometry. The percentage of cells with a sub-G1 DNA content is as indicated (Ad-ERE2F1+Ad-siGFP, gray; Ad-ERE2F1+Ad-siDip, black). Each graph represents the mean±S.D. of two independent experiments. *AdsiGFP versus Ad-siDip, P=0.046 (day 2)
Discussion
The importance of the transcription factor E2F1 for the induction of apoptosis either in association with p53 or alone is evident by a large number of studies. Especially on the p53-independent apoptotic pathway, novel E2F1-regulated downstream target genes have been described,34, 36, 38, 45, 46, 47 implicating E2F1 in the regulation of different aspects of programmed cell death. In this study, we investigated the biological function of our newly identified E2F1 target gene DIP (KIAA0767)36 in a p53-negative cellular context. Our results demonstrate that upregulation of endogenous DIP by E2F1 stimulation as well as ectopic expression of DIP result in the induction of apoptosis in several experimental settings. Overexpressed DIP was capable of activating caspase-3 and cleavage of PARP, a known substrate for effector caspases. More importantly, inhibition of endogenous DIP expression significantly diminished E2F1-induced apoptosis, indicating that regulation of DIP protein expression by E2F1 contributes to its apoptotic function in the absence of p53. We also show that DIP localizes to the mitochondria where endogenous DIP levels are readily induced after 24 h of E2F1 activation.
Our current knowledge of the mechanism by which E2F1 promotes cell death in a p53-independent manner includes direct transcriptional activation of genes involved in apoptosis such as p73,33, 34 Apaf-1,32, 48 caspases 3, 7, 8, and 9,45 and the proapoptotic BH3-only proteins PUMA, Noxa, Bim, and Hrk/DP5.49 So far, three major mechanisms of E2F1-induced apoptosis have been described: (i) inhibition of antiapoptotic signaling;50 (ii) the stabilization of p53 via transcription of p14ARF;1 and (iii) independently of p53 via induction of several proapoptotic targets.32, 33, 34, 36, 48, 49 The increased levels of ARF or p73 result in the transcriptional activation of Bax (and other p53/p73 target genes such as PUMA;51 which in turn causes Bax mitochondrial translocation and cyt c release. In addition, the BH3-only family members PUMA, Noxa, Bim, Hrk/DP5 are also directly upregulated by E2F1,49 while the mechanism of activation of two other family members, Bad and Bid, shown to be elevated through E2F1 overexpression36 remains to be elucidated. Ultimately, the E2F1-mediated stimulus leads to sequential activation of caspases consisting of initiators and effectors forming a proteolytic cascade to induce DNA fragmentation and cell death.44 Within the mitochondrial pathway, cyt c complexes with Apaf-1 to cleave procaspase-9 into its active form. Moreover, enforced E2F1 expression has been previously reported to increase the levels of Apaf-1 and caspase-3, -7, -8 and -9 directly,32, 45, 48 thus bypassing the mitochondrial route.
Structural analysis of DIP revealed an up to 76% probability for this protein of being located in the mitochondrion. Consistent with its morphological features, endogenous DIP protein was clearly upregulated in the mitochondrial fraction in response to E2F1. Together with the data from DIP overexpression, demonstrating ectopically expressed DIP in the mitochondria, it is very likely that DIP is a mitochondrial protein. In our experiments, increased levels of DIP have been shown to induce activation of apoptosis-associated effector caspases such as caspase-3 followed by PARP cleavage. Apparently, DIP-mediated apoptosis occurs not exclusively via the caspase-dependent pathway, since inhibition of cysteine proteases by the pan-caspase inhibitor zVAD-fmk lead to an incomplete block of apoptosis, implicating that caspase-independent pathways might also be involved. Importantly, together with recently published data indicating a direct modulation of the human cytochrome c1 promoter52 and AIF upregulation in the cytosol by E2F1,53 our results support a model in which increased E2F1 activity triggers apoptosis also by stimulating proapoptotic factors in the mitochondrion. The fact that E2F1 affects death-promoting molecules on multiple levels again emphasizes its role as a key mediator of apoptosis. The detailed mechanism whereby the novel DIP converts E2F1-stimulated apoptosis and its subcellular fate remains to be elucidated.
Materials and Methods
Cell culture
Early passage human VH6 foreskin fibroblasts were obtained from M Roggendorf (University of Essen). H1299 cells were kindly provided by B Opalka (University of Essen). Saos-2 cells were purchased from DSMZ; Braunschweig, Germany. HCT 116 (p53−/−) and HCT 116 (p53+/+) were kindly provided by B Vogelstein. Retrovirally infected Saos-2 ERE2F1 cells have been previously described.34 Cells were generally maintained in Dulbecco's modified Eagle's medium (DMEM, Invitrogen; Groningen, The Netherlands) supplemented with 10% fetal calf serum (FCS, Biochrom; Berlin, Germany) and 1% penicillin G/streptomycin sulfate (Invitrogen; Groningen, The Netherlands). Media for maintenance of Saos-2 ERE2F1 cells contained puromycin (Sigma; Steinheim, Germany) at a concentration of 1 μg/ml. E2F1 activity in Saos-2 ERE2F1 cells was induced by 1 μ M 4-OHT.
Adenovirus vector construction
DIP alias KIAA0767 was full-length cloned using primers 5′-atgctaaggaggttggacaaaatc-3′ and 5′-gctgtccccgccagacg-3′ with PfuTurbo hot start polymerase (Stratagene; Heidelberg, Germany) in pCR-TOPO (Invitrogen; Groningen, The Netherlands). All adenoviral vectors were generated using the AdEasy-system54 (kindly provided by B Vogelstein). After sequence verification, DIP cDNA was subcloned in pAdTrack-CMV-plasmid. Virus was generated by homologous recombination following cotransformation with pAdEasy1 in Escherichia coli BJ5183. All viruses were propagated, purified, and titrated as described.39 Adenoviral infections were carried out at multiplicities of infection (MOIs) which allow 100% transduction of each cell line. This represents MOIs 35, 10, 50, and 35 for Saos-2 cells, H1299 cells, VH6 fibroblasts, and HCT 116 cells (regardless of their p53 status), respectively. Transduction efficiency was monitored by GFP-positive cells, since each Ad vector expresses GFP in addition to its particular transgene as a fluorophore marker. The vectors AdCMV-GFP (Ad-GFP),39 AdCMV-p53 (Ad-p53wt),55 and AdCMV-ERE2F1 (Ad-ERE2F1)40 have been previously described. Adenoviral vectors expressing siRNAs are described below.
MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium-bromide) cell viability assay
For MTT assay, cells were seeded on 96-well plates at 104 cells per well and infected 16 h later. After infection, cell viability was assayed at indicated times in triplicate wells using the CellTiter96 AQueous One Solution Cell Proliferation Assay (Promega; Mannheim, Germany).
Flow cytometry
For flow cytometry analysis, cells were harvested at indicated times after infection, fixed in 70% ethanol and stained for DNA content with propidium iodide. Analysis was carried out in a FACSVantage flow cytometer (Becton Dickinson, San Diego, USA) using CellQuest Software.
Hoechst staining
At 24, 48, 72, and 96 hpi, Hoechst 33342 was added to the cell culture medium at 1 μg/ml. Cells were incubated at 37°C/5% CO2 and subsequently subjected to fluorescence microscopy.
Western blotting
Cells were lysed in RIPA buffer (50 mM Tris-Cl, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) and total protein concentration was quantitated by Bradford assay. Equal amounts of each sample were separated by SDS-polyacrylamid gel electrophoresis (PAGE), transferred to nitrocellulose membranes (Amersham-Biosciences; Freiburg, Germany) and probed with antibodies against E2F1 (Santa Cruz Biotechnology; #sc-251, Heidelberg, Germany), p53 (Santa Cruz Biotechnology; #sc-6243, Heidelberg, Germany), cleaved caspase-3 (Cell Signaling Technology; #9661, Frankfurt, Germany), full-length PARP (Cell Signaling Technology; #9542, Frankfurt, Germany), cleaved PARP (Cell Signaling Technology; #9541, Frankfurt, Germany), or custom made anti-Dip antibody directed against epitope aa 2–16. Reprobing of the blot with actin antibody (Santa Cruz Biotechnology; #sc-1616, Heidelberg, Germany) served as gel loading control.
Semiquantitative RT-PCR
RT-PCR was performed on total RNA prepared by RNeasy Mini Kit (Qiagen; Hilden, Germany). Following DNase I treatment, 1 μg RNA was reverse transcribed using Omniscript RT (Qiagen; Hilden, Germany) and Oligo-dT. PCR amplification was performed as previously described.34 A minimum number of cycles was performed to obtain a clear signal within the linear amplification phase. Following specific primer pairs were used: Dip: 5′-gtctgtagctggggaccagt-3′ and 5′-tcaggacaagctgagggagc-3′; cyclin E: 5′-caacatacagacccacagag-3′ & 5′-cctttaggtcaagtagcacc-3′; p73: 5′-gacggaattcaccaccatcct-3′ and 5′-ccaggctctctttcagcttca-3′; E2F1: 5′-ggtgaggtccccaaagtcac-3′ and 5′-gccaccatagtgtcaccacc-3′; GAPDH: 5′-cacagtccatgccatcac-3′ and 5′-caccaccctgttgctgta-3′.
Localization studies
H1299 cells were transfected with pCDNA3.1-DIP/myc-His-expressing His-tagged full-length DIP or parental pCDNA 3.1/myc-His vector by electroporation as described34 and seeded on coverslips. At 24 h post transfection, cells were fixed with medium containing 3.7% formaldehyde and permeabilized with PBS=/0.2% Triton X-100. His-tagged DIP was visualized using anti-His antibody # R930-25 (Invitrogen; Groningen, The Netherlands) and anti-mouse Alexa Fluor 546 # A-11018 (Molecular Probes; Leiden, The Netherlands). Nuclei and mitochondria were stained using DAPI (Sigma; Steinheim, Germany) at 10 mg/ml and mitochondrion-selective MitoTracker® Green FM # M-7514 (Molecular Probes; Leiden, The Netherlands)56 at 200 nM concentration, respectively.
siRNA technology
Gene expression of endogenous Dip was suppressed using adenoviruses expressing siRNAs against this protein. Ad-siDip vectors were created, purified and titrated using DIP-specific oligonucleotides 5′-tcgaggactctatgctggtatcaatcgattgataccagcatagagtcc-3′ and 5′-ctagggactctatgctggtatcaatcgattgataccagcatagagtcc-3′ with the GeneSuppressor System (Biocarta; Hamburg, Germany) according to the supplier's protocol. Ad-siGFP vectors (part of the kit) were constructed as negative control.
Subcellular fractionation
Mitochondrial and cytosolic protein fractions were separated and purified using the ApoAlert Kit (BD Clontech; Heidelberg, Germany) according to the manufacturer's protocol. Protein concentrations were quantitated by Bradford assays. Equal amounts of proteins were analyzed by SDS-PAGE using Cox 4 mitochondrial protein antibody (ApoAlert Kit) and custom made Dip antibody.
Statistical analysis
Variations in DNA content in the sub-G1 population of cells were compared with an unpaired Student's t test. The statistical test was one-sided.
Abbreviations
- DIP:
-
death-inducing-protein
- PARP:
-
poly(ADP-ribose)-polymerase
- siRNA:
-
small interfering RNA
- Rb:
-
retinoblastoma
- cdk:
-
cyclin-dependent kinase
- Mdm2:
-
mouse double minute 2
- ARF:
-
ADP ribosylation factor 1
- ATM:
-
ataxia telangiectasia mutated
- Apaf1:
-
apoptosis protease-activating factor 1
- ER:
-
estrogen receptor
- 4-OHT:
-
4-hydroxytamoxifen
- mRNA:
-
messenger RNA
- Ad:
-
adenovirus
- GFP:
-
green fluorescent protein
- FACS:
-
fluorescence-activated cell sorter
- zVAD-fmk:
-
N-benzyloxycarbonyl-Val-Ala-Asp-fluoromethylketone
- bp:
-
base pair
- ORF:
-
open reading frame
- RT-PCR:
-
reverse transcription-polymerase chain reaction
- cyt c:
-
cytochrome c
- DMEM:
-
Dulbecco's modified Eagle's medium
- MOI:
-
multiplicity of infection
- CMV:
-
cytomegalovirus
- MTT:
-
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium-bromide
- hpi:
-
hours postinfection
- PAGE:
-
polyacrylamid gel electrophoresis
- Casp-3:
-
caspase-3
References
Müller H and Helin K (2000) The E2F transcription factors: key regulators of cell proliferration. Biochim. Biophys. Acta 1470: M1–M12
Dyson N (1998) The regulation of E2F by pRb-family proteins. Genes Dev. 12: 2245–2262
Lam EWF and LaThangue NB (1994) DP and E2F proteins: coordinating transcription with cell cycle progression. Curr. Opin. Cell. Biol. 6: 859–866
Helin K (1998) Regulation of cell proliferation by the E2F transcription factors. Curr. Opin. Genetics. Dev. 8: 28–35
Nevins JR (1998) Toward an understanding of the functional complexity of the E2F and retinoblastoma families. Cell Growth Differ. 9: 585–593
Trimarchi JM and Lees JA (2002) Sibling rivalry in the E2F family. Nat. Rev. Mol. Cell Biol. 3: 11–20
Moberg K, Starz MA and Lees JA (1996) E2F-4 switches from p130 to p107 and pRb in response to cell cycle reentry. Mol. Cell. Biol. 16: 1436–1449
Takahashi Y, Rayman JB and Dynlacht BD (2000) Analysis of promoter binding by the E2F and pRb families in vivo: distinct E2F proteins mediate activation and repression. Genes. Dev. 14: 804–816
Hijmans FM, Voorhoeve PM, Beijersbergen RL, van't Veer LJ and Bernards R (1995) E2F-5, a new E2F family member that interacts with p130 in vivo. Mol. Cell. Biol. 15: 3082–3089
Gaubatz S, Wood JG and Livingston DM (1998) Unusual proliferation arrest and transcriptional control properties of a newly discovered e2f family member, E2F-6. Proc. Natl. Sci. Acad. USA 95: 9190–9195
Trimarchi JM, Fairchild B, Verona R, Moberg K, Andon N and Lees JA (1998) E2F-6, a member of the e2f family that can behave as a transcriptional repressor. Proc. Natl. Sci. Acad. USA 95: 2850–2855
Ogawa H, Ishiguro K, Gaubatz S, Livingston DM and Nakatani Y (2002) A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296: 1132–1136
Trimarchi JM, Fairchild B, Wen J and Lees JA (2001) The 2A7E transcription factor is a component of the mammalian Bmi1-containing polycomb complex. Proc. Natl. Sci. Acad. USA 98: 1519–1524
De Bruin A, Maiti B, Jakoi L, Timmers C and Leone G (2003) Identification and characterization of E2F7, a novel mammalian E2F family member capable of blocking cellular proliferation. J. Biol. Chem. 278: 42041–42049
Sherr CJ (1996) Cancer cell cycles. Science 274: 1672–1677
Weinberg RA (1995) The retinoblastoma protein and cell cycle control. Cell 81: 323–330
DeGregori J, Leone G, Miron A, Jakoi L and Nevins JR (1997) Distinct roles for E2F proteins in cell growth control and apoptosis. Proc. Natl. Acad. Sci. USA 94: 7245–7250
Kowalik TF, DeGregori J, Schwarz JK and Nevins JR (1995) E2F1 overexpression in quiescent fibroblasts leads to induction of cellular DNA synthesis and apoptosis. J. Virol. 69: 2491–2500
Qin XQ, Livingston DM, Kaelin Jr WG and Adams PD (1994) Deregulated transcription factor E2F-1 expression leads to S-phase entry and p53-mediated apoptosis. Proc. Natl. Acad. Sci. USA 91: 10918–10922
Yamasaki L, Jacks T, Bronson R, Goillot E, Harlow E and Dyson NJ (1996) Tumor induction and tissue atrophy in mice lacking E2F-1. Cell 85: 537–548
Hallstrom TC and Nevins JR (2003) Specificity in the activation and control of transcription factor E2F-dependent apoptosis. Proc. Natl. Acad. Sci. USA 100: 10848–10853
Hiebert SW, Packham G, Strom DK, Haffner R, Oren M, Zambetti G and Cleveland JL (1995) E2F-1 : DP-1 induces p53 and overrides survival factors to trigger apoptosis. Mol. Cell. Biol. 15: 6864–6874
Bates S, Phillips AC, Clark PA, Stott F, Peters G, Ludwig RL and Vousden KH (1998) p14ARF links the tumour suppressors RB and p53. Nature 395: 124–125
Kamijo T, Weber JD, Zambetti G, Zindy F, Roussel MF and Sherr CJ (1998) Functional and physical interaction of the ARF tumor suppressor with p53 and Mdm2. Proc. Natl. Acad. Sci. USA 95: 8292–8297
Russell JL, Powers JT, Rounbehler RJ, Rogers PM, Conti CJ and Johnson DG (2002) ARF differentially modulates apoptosis induced by E2F1 and Myc. Mol. Cell. Biol. 22: 1360–1368
Tolbert D, Lu X, Yin C, Tantama M and Van DT (2002) p19(ARF) is dispensable for oncogenic stress-induced p53-mediated apoptosis and tumor suppression in vivo. Mol. Cell. Biol. 22: 370–377
Martelli F, Hamilton T, Silver DP, Sharpless NE, Bardeesy N, Rokas M, DePinho RA, Livingston DM and Grossman SR (2001) p19ARF targets certain E2F species for degradation. Proc. Natl. Acad. Sci. USA 98: 4455–4460
Rogoff HA, Pickering MT, Debatis ME, Jones S and Kowalik TF (2002) E2F1 induces phosphorylation of p53 that is coincident with p53 accumulation and apoptosis. Mol. Cell. Biol. 22: 5308–5318
Holmberg C, Helin K, Sehested M and Karlstrom O (1998) E2F-1 induced p53-independent apoptosis in transgenic mice. Oncogene 17: 143–155
Hsieh J-K, Fredersdorf S, Kouzarides T, Martin K and Lu X (1997) E2F-1 induced apoptosis requires DNA binding but not transactivation and is inhibited by the retinoblastoma protein through direct interaction. Genes Dev. 11: 1840–1852
Phillips AC, Stewart S, Ryan KM, Helin K and Vousden KH (1997) Induction of DNA synthesis and apoptosis are separable functions of E2F-1. Genes Dev. 11: 1853–1863
Moroni MC, Hickmann, ES, Denchi EL, Capara G, Colli E, Cecconi F, Müller H and Helin K (2001) Apaf-1 is a transcriptional target for E2F1 and p53. Nat. Cell Biol. 3: 552–558
Irwin M, Martin MC, Phillips AC, Seelan RS, Smith DI, Liu W, Flores ER, Tsai KY, Jacks T, Vousden KH and Kaelin Jr WG (2000) Role for the p53 homologue p73 in E2F-1-induced apoptosis. Nature 407: 645–648
Stiewe T and Pützer BM (2000) Role of the p53 homolog p73 for E2F1-induced apoptosis. Nat. Genet. 26: 464–469
Ma Y, Croxton R, Moorer RLJ and Cress WD (2002) Identification of novel E2F1-regulated genes by microarray. Arch. Biochem. Biophys. 399: 212–224
Stanelle J, Stiewe T, Theseling CC, Peter M and Pützer BM (2002) Gene expression changes in response to E2F1 activation. Nucleic Acids Res. 30: 1859–1867
Ishida S, Huang E, Zuzan H, Spang R, Leone G, West M and Nevins JR (2001) Role for E2F in control of both DNA replication and mitotic functions as revealed from DNA microarray analysis. Mol. Cell. Biol. 21: 4684–4699
Müller H, Bracken AP, Vernell R, Moroni MC, Christians F, Grassilli E, Prosperini E, Vigo E, Oliner JD and Helin K. (2001) E2Fs regulate the expression of genes involved in differentiation, development, proliferation, and apoptosis. Genes Dev. 15: 267–285
Pützer BM, Stiewe T, Crespo F and Esche H (2000) Improved safety through tamoxifen-regulated induction of cytotoxic genes delivered by Ad vectors for cancer gene therapy. Gene Ther. 7: 1317–1325
Stanelle J, Stiewe T, Rödicker F, Köhler K, Theseling CC and Pützer BM (2003) Mechanism of E2F1-induced apoptosis in primary vascular smooth muscle cells. Cardiovasc. Res. 59: 512–519
Kerr JFR, Wyllie AH and Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26: 239–257
Wyllie AH, Kerr JFR. and Currie AR (1980) Cell death: the significance of apoptosis. Int. Rev. Cytol. 68: 251–306
Wyllie AH (1980) Glucocorticoid-induced thymocyte apoptosis is associated with endogenous endonuclease activation. Nature 284: 555–556
Thornberry NA and Lazebnik Y (1998) Caspases: enemies within. Science 281: 1312–1316
Nahle Z, Polakoff J, Davuluri RV, McCurrach ME, Jacobson MD, Narita M, Zhang MQ, Lazebnik Y, Bar-Sagi D and Lowe SW (2002) Direct coupling of the cell cycle and cell death machinery by E2F. Nat. Cell Biol. 4: 859–864
Polager S, Kalma Y, Berkovich E and Ginsberg D (2002) E2Fs up-regulate expression of genes involved in DNA replication, DNA repair and mitosis. Oncogene 21: 437–446
Susin SA, Lorenzo HK, Zamzami N, Marzo I, Snow BE, Brothers GM, Mangion J, Jacotot E, Costantini P, Loeffler M, Larochette N, Goodlett DR, Aebersold R, Siderovski DP, Penninger JM and Kroemer G (1999) Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 397: 441–446
Furukawa Y, Nishimura N, Furukawa Y, Satoh M, Endo H, Iwase S, Yamada H, Matsuda M, Kano Y and Nakamura M (2002) Apaf-1 is a mediator of E2F-1-induced apoptosis. J. Biol. Chem. 277: 39760–39768
Hershenko T and Ginsberg D (2004) Up-regulation of BH3-only proteins by E2F1: role in apoptosis. J. Biol. Chem. 279: 8627–8634
Phillips AC, Ernst MK, Bates S, Rice NR and Vousden KH (1999) E2F-1 potentiates cell death by blocking antiapoptotic signaling pathways. Mol. Cell 4: 771–781
Melino G, Bernassola F, Ranalli M, Yee K, Zong WX, Corazzari M, Knight RA, Green DR, Thompson C and Vousden KH (2004) p73 induces apoptosis via PUMA transactivation and Bax mitochondrial translocation. J. Biol. Chem. 279: 8076–8083
Luciakova K, Barath P, Li R, Zaid A and Nelson BD (2000) Activity of the human cytochrome c1 promoter is modulated by E2F. Biochem. J. 351: 251–256
Vorburger SA, Pataer A, Yoshida K, Liu Y, Lu X, Swisher SG and Hunt KK (2003) The mitochondrial apoptosis-inducing factor plays a role in E2F-1-induced apoptosis in human colon cancer cells. Ann. Surg. Oncol. 10: 314–322
He TC, Zhou S, da Costa LT, Yu J, Kinzler KW and Vogelstein B (1998) A simplified system for generating recombinant adenoviruses. Proc Natl Acad Sci USA 95: 2509–2514
Stiewe T, Theseling CC and Pützer BM (2002) Transactivation-deficient deltaTA-p73 inhibits p53 by direct competition for DNA binding. J. Biol. Chem. 277: 14177–14185
Poot M, Zhang ZY, Kramer JA, Wells KS, Jones LJ, Hanzel DK, Lugade AG, Singer VL and Haugland RP (1996) Analysis of mitochondrial morphology and function with novel fixable fluorescent stains. J. Histochem. Cytochem. 44: 1363–1372
Acknowledgements
We thank Bärbel Pollmeier for technical assistance. We are grateful to Klaus Lennartz for support in flow cytometry and Peter Babioch for assistance in laser scanning microscopy. This work was supported by Grant 10-1934-Pü3 from the Deutsche Krebshilfe to BMP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd).
Supplementary information
Rights and permissions
About this article
Cite this article
Stanelle, J., Tu-Rapp, H. & Pützer, B. A novel mitochondrial protein DIP mediates E2F1-induced apoptosis independently of p53. Cell Death Differ 12, 347–357 (2005). https://doi.org/10.1038/sj.cdd.4401532
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401532
Keywords
This article is cited by
-
Role of Cell Cycle Re-Entry in Neurons: A Common Apoptotic Mechanism of Neuronal Cell Death
Neurotoxicity Research (2012)
-
GRAMD4 mimics p53 and mediates the apoptotic function of p73 at mitochondria
Cell Death & Differentiation (2011)
-
DAPK2 is a novel E2F1/KLF6 target gene involved in their proapoptotic function
Oncogene (2008)
-
Systematic mutation analysis of KIAA0767 and KIAA1646 in chromosome 22q-linked periodic catatonia
BMC Psychiatry (2005)
-
E2F and p53 make a nice couple: converging pathways in apoptosis
Cell Death & Differentiation (2005)