Abstract
Phosphorothioated DNA (PT-DNA) exhibits a mild anti-oxidant property both in vivo and in vitro. It was found that 8-OHdG and ROS levels were significantly lower in dnd+ (i.e. S+) E. coli., compared to a dnd− (i.e. S−) strain. Furthermore, different from traditional antioxidants, phosphorothioate compound presents an unexpectedly high capacity to quench hydroxyl radical. Oxidative product analysis by liquid chromatography-mass spectrometry and quantum mechanistic computation supported its unique anti-oxidant characteristic of the hydroxyl selectivity: phosphorothioate donates an electron to either hydroxyl radical or guanine radical derived from hydroxyl radical, leading to a PS• radical; a complex of PS• radical and OH− (i.e. the reductive product of hydroxyl radical) releases a highly reductive HS• radical, which scavenges more equivalents of oxidants in the way to high-covalent sulphur compounds such as sulphur, sulphite and sulphate. The PS-PO conversion (PS and PO denote phosphorus-sulphur and phosphorus-oxygen compounds, respectively) made a switch of extremely oxidative OH• to highly reductive HS• species, endowing PT-DNA with the observed high capacity in hydroxyl-radical neutralization. This plausible mechanism provides partial rationale as to why bacteria develop the resource-demanding PT modification on guanine-neighboring phosphates in genome.
Similar content being viewed by others
Introduction
DNA Phosphorothioation is a novel type of chemical modification originally discovered in Streptomyces species and then determined genetically in many kinds of bacteria1,2,3. The chemical modification in gene of Streptomyces lividans is found to be Rp stereo-specific and sequence-selective, which was partially rationalized theoretically4,5,6 and verified experimentally7,8; moreover, a mobile five-membered gene cluster dndABCDE9,10,11 commits the chemical modification on bacterial DNA. Basically, two biological functions were proposed for the physiological DNA modification12,13,14,15,16. Molecular recognition between PT-DNA motif and specific proteins strongly endorsed its restriction response in host bacteria12,13,14; for example, certain species (not all dnd+ bacteria) such as S. enterica serovar Cerro 87, possesses a downstream gene cluster dndFGHI after dndA-E to encode a restriction-enzyme system that distinguishes whether phosphorothioation occurred at the specific site in DNA. Besides, anti-oxidantivity of phosphorothioated DNA (PT-DNA) was aware in the previous experiment15, but then in the deep dispute: the original discovery of anti-oxidant benefit might be flawed by the use of wild-type bacteria possessing both PT modification genes and PT-dependent restriction genes; knocking out the PT modification genes would lead to higher sensitivity toward oxidative stress – it might be false positive from side-affect of remaining restriction enzymes in the wild-type bacteria14. Recently, a following-up paper has been published in Frontiers in Microbiology by Dai et al., confirming that the phantomlike resistance in wide type Streptomyces lividans to oxidative stress is relevant to PT-DNA directly, neither other dnd proteins nor RNA-related regulation16. However, more questions need addressing for the elusive biological effect, in particular at the level of biochemistry and molecular biology - For example, how are the rare copies of PT modifications able to conveys special in vivo resistance to oxidative stress? – it should be noted that the chemical modification occurs at guanine-neighbouring sites such as GpsA, GpsT, and CpsC (complementary of GG) at a relatively low frequency of 300~3000/106 nt1,2. How to explain the fact that the in vitro PT oxidation led to both sulphur removal and DNA backbone cleavage? Thus, the earlier works about the PT antioxidant activity were still preliminary and conflicting, mainly for lack of detailed mechanistic investigation in rationale on the promising hypothesis that the sparse PT sites meet anti-oxidant needs of a long nucleobase sequence.
Indeed, DNA oxidative damage is usually caused by highly reactive oxygen species (ROS), such as singlet oxygen, superoxide, hydrogen peroxide, and hydroxyl radical17. Many chemical modifications in DNA damage (e.g. DNA cleavage, DNA-protein cross links, purine oxidation, etc.) stem from the reactions with ROS, in particular with hydroxyl radical (HO•). Hydroxyl radical is very rare in a living cell, but extremely reactive, scrambling electrons and hydrogen atoms from protein and metabolites in a diffusive manner, and even from stable biomolecules such as DNA and lipids. Both the heterocyclic base and sugar-phosphate backbone are vulnerable, in particular via electron abstraction on guanine base and hydrogen abstraction on ribose 5′-C. If the nucleobase-repair systems are not able to immediately regenerate intact nucleobase, a mutation in genome would result from erroneous base pairing during replication or, consequently, DNA cleavage would occur consequently. It is also believed that metal ions such as iron and copper would enhance hydroxyl radical generation via Fenton chemistry18.
In short, DNA oxidative damage seems to begin with one-electron loss and formation of a radical cation “hole” at the encounter position19. The hole hops reversibly through electronic structure along the DNA helix, trapped at a guanine-rich place as a form of guanine radical cation (G•+)20. Hydrolysis of G•+ releases 8-hydroxy-2′-deoxyguanosine (8-OHdG), upon consequential DNA repairing in vivo. So 8-OHdG is an effective biomarker for DNA oxidative damage21.
Fischer et al. have also reported in vivo antioxidant capacity of 5′-PT-dNTP in drug development for neurodegenerative disorders, but as attributing to metal-ion coordination and Fenton inhibition22. In contrast, low molecular-weight free thiols such as glutathione (GSH), cysteine (CYS), and homocystine can ameliorate oxidative damage in an alternative fashion, via an oxidative dimerization towards RSSR23.
In this work we measured the both the 8-OHdG and ROS intracellular levels of dnd+ (S+) and dnd− (S−) strains of E. coli without intervention of restriction genes dndFGHI to verify the anti-oxidant property of PT-DNA. Then we revealed its anti-oxidant mechanism with in vitro oxidation of pJTU1238 plasmid and phosphorothioate dinucleotide, using gel electrophoresis, circular dichroism spectroscopy, high performance liquid chromatography - mass spectroscopy, and quantum chemistry. Here we propose a unique anti-oxidant mechanism, different from metal-ion coordination18 and sulfur-sulfur dimerization23 in the literatures.
Results
In vivo Anti-oxidant Effect
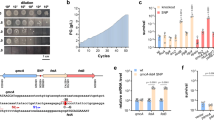
Figure 1a shows DCF fluorescence of dnd+ (S+) and dnd− (S−) E. coli strains over time under oxidative stress mimicked with Fenton’s reagent of 6 μM FeSO4 and 0.5 mM H2O2. Compared to the starting point, the intracellular ROS level represented by DCF fluorescence climbed up by 18.0 ± 8.4, 40.9 ± 10.0, 42.7 ± 4.7, 45.2 ± 7.8, 62.1 ± 11.7, and 94.8 ± 10.2% in the dnd− culture, at 1, 5, 10, 15, 20, and 25 min, respectively. By contrast, the intracellular ROS accumulation in E. coli (S+) strain fluctuates at 3.8 ± 5.5, 22.5 ± 5.7, 14.9 ± 12.9, 27.3 ± 6.6, 40.5 ± 4.9, and 35.0 ± 12.7%, respectively.
(a) The intracellular ROS levels measured by the DCF fluorescent intensity at excitation of 485 nm and emission of 528 nm, treated with 0.5 mM H2O2; (b) the time-resolved peroxide concentrations monitored with Raman vibration at 876 cm−1 and initial concentration of 30 mM H2O2; (c) the 8-OHdG level and (d) the GSH:GSSG ratio in the samples treated for 4 hours with moderate concentrations of H2O2 (the treatments included 6.0 μM FeSO4 as Fenton’s catalyst; asterisk and octothorp mark the statistical significance, p < 0.05).
Figure 1b shows the real concentration of hydrogen peroxide of dnd+ (S+) and dnd− (S−) E. coli strains over time under oxidative stress, using 30 mM H2O2 (similar to the experimental condition in ref. 16), monitored by the O-O Raman vibrational band at 876 cm−1. Accordingly, hydrogen peroxide consumed rapidly in an exponential-decay fashion for a variety of reductants in the living organisms, and the regression analysis of five replicas resulted in decay time constants of 8.5 ± 1.7 and 9.8 ± 1.9 min in the cases of the S+ and S− cultures, respectively. This observation was in agreement with the previous experiments of wide type S. lividans and the dnd− mutant treated with 20 mM H2O2, reported by Dai et al. (as shown in Fig. S2 of ref. 16).
As shown in Fig. 1c, when the strains were charged with a moderate Fenton’s reagent, the 8-OHdG level arose about 12.5 ± 14.7~58.3 ± 2.3% in the S+ strain with increasing H2O2 concentrations from 0.5 to 2.0 mM; conversely, the values in the S− strains rocketed up by 32.5 ± 3.3~180.0 ± 4.1%. In Fig. 1d, the measured GSH:GSSG ratio dropped down dramatically from 135.4 ± 20.2 to 75.4 ± 2.2 in the E. coli (S−) strain. Conversely, the values increased in the E. coli (S+) strains from 139.3 ± 12.5 to 276.8 ± 38.4. In literatures, the normal GSH:GSSG level of about 200 decreased to 10~1 under oxidative stress24, and a 2.5 fold increase of the 8-OHdG level was reported in 10 T½ cells in the presence of 0.1 mM H2O225.
In vitro Anti-oxidant Tests of Plasmid PT-DNA
Figure 2 showed the in vitro experimental results of plasmid oxidation with H2O2 concentrations in a range of 0.0 to 1.5 μM for 12 hours in the dark, with comparison to a ABTS pre-oxidized PT-DNA sample ([PT] ≈ 0.19 μM). It is almost indistinguishable in the image of gel electrophoresis (Fig. 2a) and the circular dichroism (CD) spectra (Fig. 2b) owing to the technical limitation; so digital quantitation on the two in vitro experiments was carried out.
(a) The image of plasmid DNA gel electrophoresis, where the 63 μg/mL DNA samples treated with H2O2 up to 1.5 μM for 12 hours; (b) the circular dichroism spectra of the DNA samples; (c) the quantitated nicked DNA percentage in total amount of DNA, upon the oxidation; (d) the quantitated helicity upon the oxidative damage at 1.0 μM H2O2.
Figure 2c show quantitation of the electrophoresis image by software image J26. Nicked circular DNA percentage in the total plasmid DNA (supercoil plus linear and nicked circular) was used to exhibit DNA oxidative damage27. The nicked-DNA percentage of PT plasmid slightly increased by 1.6 ± 1.6 and 2.6 ± 1.7% at 0.5 and 1.0 μM H2O2, respectively. However, the values were measured as 4.0 ± 2.9 and 8.9 ± 1.7% in the dnd− plasmid, and −0.4 ± 2.0 and 9.4 ± 1.6% in the pre-oxidized PT-DNA, respectively. All the three plasmid samples exhibited severe oxidative damage in the presence of 1.5 μM H2O2, with the nicked-DNA percentages of 12.7 ± 1.4, 17.7 ± 1.7, and 24.5 ± 1.6% for the PT-, normal plasmid, and pre-oxidized PT-, respectively.
Figure 2d shows quantitation of DNA conformational damage by oxidative stress, represented with a ratio of integrated areas under the positive band at 275 nm over the negative band at 245 nm in CD spectra. It has been reported that declining the positive ellipticity and inclining the negative ellipticity characterize oxidative damage on plasmid DNA27. For purpose of controlling errors from peak-wavelength shift, the integrated value in half-band width was computed and their ratios were used to denote DNA helicity. The ratios drop down by slopes of −1.2 ± 0.6, −3.1 ± 1.1, and −7.1 ± 0.8 (×10−3 min−1) for the PT-, normal, and pre-oxidized PT-DNA plasmid, respectively. On the other hand, the absolute values of ratio in the order of dnd− > dnd+ > pre-oxidized PT-DNA plasmid indicated that DNA oxidation damaged the secondary structure significantly while DNA phorosphorothioation disturbed the B-helical conformation slightly5.
Comparison of PS, GSH, CYS, and vitC
Owing to the μM technical limitation of plasmid PT-DNA, diethyl phosphorothioate (PS) was used as a model compound of PT-DNA in the following in vitro comparative experiments with non-physiological oxidants ABTS•+ and DPPH• radicals. GSH, CYS, and vitC appeared to possess a similar scavenging capacity with reference to 46.7 μM ABTS•+ in 30 min, with the measured IC50 values of 5.1 ± 0.2, 5.9 ± 0.1, and 7.1 ± 0.1 μM, respectively. This value was 42.3 ± 1.2 μM for PS, six times higher than those of traditional antioxidants. Similarly, the IC50 values towards 200 μM DPPH in 30 min were determined as 36.5 ± 1.5, 31.2 ± 0.2, and 25.3 ± 0.4 μM for GSH, CYS, and vitC, respectively; while the IC50 value for PS was measured to be 230.7 ± 8.1 μM, also about 6 to 8 times higher than the others. Thus, in terms of IC50 value with reference to organic oxidants ABTS•+ and DPPH•, phosphorothioate exhibited a mild anti-oxidant activity, in agreement with its weak reductive potential.
As shown in Fig. 3a, the ROS level characterized by the TBARS amount decreased by 0.0 ± 2.4, 7.3 ± 1.3, 29.3 ± 3.5, 34.2 ± 2.2, and 41.5 ± 12.5% at the PS concentrations of 0.25, 2.5, 25.0, 250.0, and 2500.0 μM, respectively. The values were 0.0 ± 4.9, 19.5 ± 1.8, 22.0 ± 6.3, and 14.6 ± 2.3% for GSH, and −2.4 ± 4.8, 7.3 ± 2.6, 19.5 ± 6.1, and 9.8 ± 5.4% for CYS, respectively. The corresponding ROS generation increased by 78.1 ± 5.5 and 14.6 ± 4.3% at 2500.0 μM in the case of GSH and CYS, respectively. Similar phenomena were observed with anti-oxidant vitC, in which the ROS generation increased by 4.9 ± 4.7, 14.6 ± 4.3, and 117.1 ± 3.4% at the concentrations of 25.0, 250.0, and 2500.0 μM, respectively.
(a) The ROS levels characterized by the amounts of TBARS with the absorbance at wavelength of 532 nm in 30 min, generated by Fenton’s reagent of 50 μM FeSO4 and 1.0 mM H2O2 at 37 °C and quenched with gradient antioxidant concentrations from 0.25 μM to 2.5 mM; (b) the cyclic voltammograms of the phosphorothioate model compound (lines 1–3: 10, 15, and 20 μM, respectively) and 2′-deoxyguanosine (insert, lines 4–6: 10, 15, and 20 μM, respectively), with potential voltages referred to a Ag/AgCl electrode; (c) the image of gel electrophoresis, in which 50 μg/mL plasmid DNA were pre-treated with the Fe(III)-modified Fenton’s reagent and gradient concentrations of antioxidants from 2.5 mM to 2.5 μM.
According to electrochemical measurement (Fig. 3b), the reduction potential of PS (0.74 V vs Ag/AgCl) is lower than that of 2′-deoxyguanosine by about 0.28 V. The standard reduction potentials of guanine, adenine, thymidine, and cytosine were reported to be about 1.29, 1.42, 1.70, and 1.60 V28,29. So the standards reduction potential of phosphorothioate was estimated as 0.94 V after calibration, higher than that of Fe(III)/Fe(II) by 0.20 V, but lower than the G•+/G potential. Therefore, the pro-oxidant effect for these antioxidants was determined to conform to the order of CYS > vitC > GSH ≫ PS (Figs S3 and S4). Similarly, oxidative damage of plasmid DNA in the DNA lesion experiment can be used to test the pro-oxidant effect in the presence of Fe(III)-modified Fenton’s reagent30. PS gave no detectable pro-oxidativity up to 2.5 mM, while the other three antioxidants exhibited the DNA degradation at the relatively high concentrations (Figs 3c and S5).
LCMS Characterization of Oxidative Products
As shown in Fig. 4a, two stereochemical phosphorothioated dinucleotides GsA (retention time of Rp- configuration at 8.5 min and Sp- at 8.1 min) both led to GA dinucleotide (retention time at 7.5 min). As a comparison, Fig. 4b shows the H2O2-dependent PH formation (PH: hydrogen-phosphonate compound) in oxidation of diethyl phosphorothioate (PS) by gradient H2O2 concentration. The HPLC experiments showed two new peaks for the PS oxidation by H2O2 (Fig. 4b, peak 1 and 3). As shown in Fig. 4c, high-resolution mass spectrum analysis indicated that the substance of peak 1 was O,O-diethyl phosphate (PO, [M + H]+ m/z: exp. 155.0469, ca. 155.0473), peak 2 O,O-diethyl thiophosphate (PS, original reactant, [M + H]+ m/z: exp. 171.0248, ca. 171.0245), and peak 3 O,O-diethyl hydrogen-phosphonate (PH, [M + H]+ m/z: exp. 139.0528, ca. 139.0524), with retention times at 1.2, 1.7, and 2.2 min, respectively. The two oxidative products (PO and PH) were proportional to H2O2 concentration in the oxidation.
The HPLC and LCMS analysis of oxidative products of (a) the mixture of 20 μM GsA-Rp and GsA-Sp with 100 μM H2O2 plus 1 μM FeSO4, and (b) the mixture of 25 μM PS with the gradient H2O2 concentrations; (c) the high-resolution mass spectra of components 1 (PO), 2 (PS), and 3 (PH), respectively; (d) GsA and GA.
In silico Justification
We carried out intensive quantum chemical calculations for the reaction pathway from PS toward PH and PO (Table S1 and Fig. S8). Figure 5 shows the most plausible mechanism in which PH was generated from two oxygen-donations of H2O2. In the first step, one molecule of H2O2 reacts with PS to form PSO anion (INT1) and release one molecule of water, with a calculated barrier of 23.2 kcal/mol in aqueous solution. Then, another molecule of H2O2 reacts with PSO anion, releasing one molecule of sulphur dioxide (SO2) and one water molecule and then leading to PH generation. The reaction barrier for the second step was computed to be 14.9 kcal/mol.
By comparison to the PS-PH conversion, the most plausible mechanism for PS-PO conversion has a shortcut path in the presence of hydroxyl radical. Electron transfer from phosphorothioate anion to HO• or G•+ seemed barrier-free for the extremely high reactivity of hydroxyl radical - a pre-reactive complex of [PS---OH]−• was formed instantly. The radical anion complex transformed to a five-coordinate adduct (INT2) via phosphor hydroxylation, with a calculated barrier of 6.9 kcal/mol. Such an addition-elimination mechanism is similar to hydrolysis of organophosphorus pesticides31. The INT2 was higher in energy by 5.9 kcal/mol than the pre-reactive complex. INT2 underwent an elimination of hydrogen-sulphur radical (HS•), forming the normal phosphate with a calculated barrier of 14.4 kcal/mol. The reaction of PS plus HO• towards PO plus HS• was exothermic by 27.8 kcal/mol. Therefore, the overall reaction barrier for the PS-PO conversion was estimated to be 14.4 kcal/mol (ΔG≠) in an aqueous solution. As Fig. 6 shows, the frontier orbitals of GA and GsA were observed at the M06 (CPCM, solvent = water)/6-311 + G(d, p)//M06/6-31 G(d) level of theory.
Discussion
The in vivo experiments indicated that the model organisms (E. coli, as shown in Fig. 1) of this study led to the consistent results as wild type S. lividans (dnd+, i.e. S+) and the knocking-out mutant (dnd−, i.e. S−) to oxidative stress reported by Dai et al.16. It should be acknowledged that the additional anti-oxidant capacity in the S+ strain isn’t major given that the decay time changed slightly for extracellular peroxide (12~13%) owing to coexistence of various anti-oxidant systems in the living organisms. Beside a slightly fast consumption of peroxide, it was first time to observe the intracellular ROS accumulation was suppressed by about 14~60% in the S+ strain, albeit with a sink at 10–15 min upon anti-ROS response time-delay as reported in the previous literature32. Dai et al. reported no significant up-regulation of anti-oxidant genes based on RNA-seq experiment of wide type S. lividans upon oxidative stress16, in addition to the consistent observation on dndFGHI-negative E. coli in this work, clarifying the previous suspicion whether the dnd− knocking-out would disturb dndABCDE/FGHI restriction-response system (“sick cell”). However, the intercellular ROS response exhibited dramatical difference between the S+ and S− cultures: the intracellular levels of 8-OHdG biomarker for DNA oxidative damage dropped significantly and those of GSSG biomarker for overall oxidation lowered as well; on the other hand, the GSH levels were similar in the two strains - consistent with no thiol peroxidase up-regulation based on RNA-seq experiment under oxidative stress16. All these results testified an effective anti-oxidant bypass in the S+ strain to maintain a reductive atmosphere in cell, which may stem from PT-DNA itself.
pJTU1238 in the E. coli strains were used to solve the anti-oxidant biochemistry of PT-DNA. Plasmid DNA was extracted from the two E. coli strains and tested for anti-oxidant activity in vitro directly. A typical PT population was estimated as ~1/1000 nt in the S+ strain based on the previous deep sequencing1, so the PT concentration in a stock solution of extracted dnd+ plasmid with optical density of 4.0 at 260 nm is quantified as ~200 μg/mL in ds-DNA weight or ~0.6 μM in PT equivalent. Owing to no further endogenous PT regeneration after the E. coli digestion, the anti-oxidant capacity of plasmid PT was limited at the micromolar (μM) level, significantly lower than those in vivo oxidant levels in Fig. 115,33. The plasmid experiment in Fig. 2d indicated that the oxidative damage occurred 2.6 times slower in the PT-DNA, and upon pre-oxidation, the pre-oxidized PT-DNA appeared loss of the additional anti-oxidant property in the in vitro anti-oxidant experiments, consistent with the results from the gel-image quantitation (Fig. 2c).
Although the anti-oxidant property of the extracted plasmid touched limit of determination at 1 μM H2O2, it is worthwhile to note that the low-frequency PT in genome is in agreement with economical efficiency in biology, due to the low-frequency of DNA oxidative damage in reality, e.g. usually ~1 lesion per 10000 nt upon gamma radiolysis34.
Intriguingly, the antioxidant effect of phosphorothioate compound showed unexpected selectivity to hydroxyl radical in the anti-oxidant chemistry, compared to traditional antioxidants. It has been postulated that PT “chelator” might coordinate ferrous iron and shut down Fenton reaction35. But this seems unlikely in our scenario: first, the in vitro anti-oxidant activity of plasmid DNA was observed even without an iron catalyst (due to difficult manipulation of low concentration H2O2); second, the PT consumption was observed at the same level of molar equivalent of H2O2 in the extracted plasmid experiments, indicating that PT is consumable in the anti-oxidant process. There is no evidence that PT would be inert toward hydroxyl radical, irrespective of whether PT coordinates the metal ion or not. Moreover, the coordination of PS to Fe(II) was relatively weak – compared to Fe(II)-EDTA (IC50 = 1.27 ± 0.05 μg/ml at 12.5 μM [Fe2+]), the absorbance of Fe(II)-ferrozine complex did not change in the presence of PS from 0.01 to 5.00 mM (Fig. S6). It is likely that even a strong chelator like EDTA (ethylene di-amine tetra-acetic acid) cannot inhibit oxidative stress mimicking with Fenton chemistry. Probably this is due the high sensitivity of the biological system to trace amount of oxidative damage on the DNA molecule, and so PT-metal coordination is likely to be unimportant in its anti-oxidant activity. It should also be mentioned that both GSH and CYS exhibited severe side-effect at concentrations above 250.0 μM, that is, more ROS was generated (pro-oxidant effect). Indeed, hydroxyl radical - the dominant ROS product of Fenton reaction - is the most lethal to DNA molecule, responsible for the formation of G•+ radical cation and then resulting 8-OHdG36,37.
Regarding the issue of PT-DNA backbone cleavage during oxidation, we repeated LCMS analysis of GsA oxidation of our previous work15, using Fenton’s reagent, instead. Oxidant ABTS•+ resulted in the same product as the Fenton’s reagent, but no GHA was detected in the presence of excessive amount of oxidants. The GHA formation observed in the previous work was due to peroxyacetic acid (PAA) usage or high concentration of H2O215,16. Importantly, the GsA oxidation led to only GA, no PSSP disulphide was observed as in the PS oxidation. In the absence of Fenton catalyst, no GA product was detected in the mixture of 200 μM H2O2 and 20 μM GsA after 30 min of incubation in the Tris-HCl buffer (pH = 7.4) (Fig. S7). Thus, the PT-DNA oxidation by hydroxyl radical likely resulted in a clean conversion to PO-DNA, i.e. the normal form of phosphate.
Therefore, the oxidation of phosphorothioate by ROS generated by Fenton’s reagent is different from free thiols. For example, CYS oxidation by H2O2 involves in a two-step reaction, forming sulfenic acid (CSOH) intermediate and then disulfide (CSSC), sulphenic, sulphinic or even sulphonic acid with increasing ROS concentrations38,39. The oxidation of phosphorothioate dinucleotide could desulfurize to normal phosphate40, in particular for the dilute PT encountering hydroxyl radical in a weak alkaline circumstance.
Owing to the experimental limitation, we distinguished the question whether the hydrogen-phosphonate species (PH) is an intermediate in the PS-PO conversion or a branched by-product of H2O2-dependent oxidation by quantum-chemistry calculations. As shown in Fig. 5, the PH generation needs two molecules of H2O2 with a higher reaction barrier, consistent with the HPLC-MS experimental observation in which the PH formation was highly dependent on the concentration of H2O2. The neural form of PH has two tautomeric structures, either as protonation on a phorsphorous center or as oxygen atom41. Hydrogen phosphonate would hydrolyze P-O bond into two parties at a rate of 3.2 × 105 M/s at 25 °C42. Thus, unlike phosphate, hydrogen phosphonate hydrolyzes quickly and causes DNA backbone cleavage, as the dnd+ phenotype in PT-DNA gel electrophoresis.
Based on the quantum calculations, hydrogen-sulfur radical HS• was generated in the PS-PO conversion (Fig. 5). Opposite to extremely oxidative hydroxyl radical, HS• is extremely reductive! HS• will degrade rapidly to sulphur (S8), sulfite (SO32−) and sulfate (SO42−), creating a specific reductive atmosphere in the surrounding compartment43,44.
At this point we tackle the question of how ROS invasion can precisely hit the PT site in vivo and in vitro when a PT site has an average scatter of 100~10000 nt. We believe that radical G-hopping in DNA oxidative damage helps to explain why the guanine-neighbouring PT site captures DNA damage efficiently. As shown in Fig. 6, both the HOMO and LUMO of GA anion and radical coincided each other in population, indicating that electron donation occurred on the guanine moiety and phosphate group in the oxidation of normal DNA, which is in agreement with the previous studies15. By contrast, the HOMO of GsA anion was consistent with the HOMO of GA and the SOMO of GsA radical in population, indicating that the guanine moiety of GsA is able to donate electron as well; however, the LUMO of GsA radical was computed to be dominant for the sulfur atom in population, indicating the likely formation of a phosphorothioate radical, which, as a result, keeps the guanine moiety intact in the PT-DNA oxidation. Such an electron transfer was further confirmed by the distributions of Mulliken charge and spin density of the two forms of GsA and GA, where the GA radical presented a significant charge separation between the nucleobases (+0.41 and +0.29) and phosphate (−0.97) groups with an unpaired electron on the guanine moiety (0.922), while the GsA radial only loses the negative charge (−1.296 → −0.253) and casts most of spin density (1.021) on the phosphorothioate group. We conclude that the phosphorothioate radical is hydrolyzed as via the same TS3 and TS4 transition states as shown in Fig. 5, leading to the formation of normal phosphate linkage. This also explains why the GsA was more reactive in scavenging ROS than PS, since the guanine moiety played a role in bridging electron transfer, and the relatively higher pH environment benefitted the formation of the [PS-OH]•− pre-reaction state, which can also be accelerated in the presence of hydroxyl radical as an oxidant.
In summary, due to the exceptional chemical nature of phosphorothioate, E. coli (S+) possesses DNA-intramolecular PT antioxidant activity, which acted as a guard against oxidative stress via effectively scavenging hydroxyl radical (the most lethal species to genetic materials), decreased the intracellular ROS level, and therefore limited the 8-OHdG production and promoted the GSH:GSSG ratio, and thereby provided additional protection from DNA oxidative damage. The reductivity of phosphorothioate is powerful in protecting the most vulnerable moiety (guanine) of the DNA molecules, but not too intensive to induce a pro-oxidant side effect in cell.
In vitro hydrogen phosphonate species was a branched by-product at the high H2O2 concentration. The PS-PO in vivo recycling does not require a hydrogen phosphonate intermediate, averting the dilemma of the PH-related DNA cleavage and genomic integrity discussed previously in the literatures. Figure 7 shows that the single-molecular PS-PO shortcut path led to the formation of highly reductive species, HS•, which as available for scavenging more surrounding ROS including O2−• and H2O2. The oxidation of HS• towards sulphur (S8), sulfite (SO32−) and sulfate (SO42−) provided additional antioxidant capacity (see Fig. S9). Moreover, the computations clarified the reasons of why phosphorothioation frequently occurs at a guanine-rich location. Thus, at this stage, the biologically functional loop of anti-oxidative PT-DNA was closed with the dnd-gene-induced phosphate S-incorporation and the oxidative PS-PO recycling. It is also reasonable that eukaryote develops complicated karyotheca substructure tightly packing genetic materials away from mitochondria, while a prokaryote needs additional protector from oxidative metabolites. However, whether the PS-PO conversion cascades an in vivo ROS-signal to regulate PT-DNA expression in the ROS response needs further investigation. The novel mechanism of anti-ROS property of PT-DNA may be applicable in anti-tumor and anti-aging pharmaceutics.
Methods
General
E. coli DH10B harboring pJTU1238 was prepared as the E. coli (S+) strain, and the one with dndC-disrupted pJTU1238 was used as the control in this work (namely the E. coli (S−) strain). The E. coli (S+) strain displayed the dnd phenotype in electrophoresis but the E. coli (S−) strain did not. All chemicals were of analytical grade and used without further purification. 8-OHdG ELISA kit was obtained from the Shanghai Enzyme-linked Biotechnology company. Plasmid Mini preparation kit was purchased from DingGuo Biotech company.
DCFH-DA determination
In the in vivo ROS-level test, the 0.2 mg/ml DCFH-DA stock solution was added into the E. coli cultures to make the final concentration of 10 μM. After 30 min incubation at 37 °C, the suspension was washed with Tris-HCl (pH 7.4) and diluted to 3 × 108 cfu. Then 150 μl samples in 96-well plate were treated with Fenton’s reagent. The intracellular ROS level was measured with the DCF fluorescence and the reaction condition was selected according to Fig. S1.
8-OHdG levels
The sample for the measurement of 8-OHdG levels was prepared in a four-hour incubation with Fenton’s reagent, sonication in ice-cold Tris-HCl, and centrifugation at 8000 G to remove cellular debris. In the 8-OHdG test, the supernatant was mixed with the stop reagent of the ELISA kit, and the absorbance at 450 nm was measured to compare to a calibration curve constructed with the standard 8-OHdG solutions at 2.5 to 40 pg/ml.
Pre-oxidation of plasmid PT-DNA
This procedure was developed based on the ABTS+ oxidation of GsA, in which GA was determined as the only oxidative product by HPLC and LCMS. Accordingly, the plasmid PT-DNA extracted from the dnd+ (S+) E. coli was treated with 46.7 mM ABTS•+ and then purified as the pre-oxidized Ox(S+) plasmid DNA sample (Figs S2 and S7).
Raman spectra
Raman spectra were recorded on a Bayspec Raman spectroscopy. For each kinetic measurement, H2O2 was added into the S+ or S− E. coli strains washed by 10 mM PBS (E. coli OD600 ~0.75, peroxide ~30 mM, in 10 mM PBS). Then the time-resolved spectra were collected in a rate of one spectrum per 7 seconds. The Raman intensity at the wavenumber of 876 cm−1 (O-O vibration) was used to monitor the peroxide concentration45.
CD spectra
CD spectra were measured on a Jasco J-815 spectropolarimeter. For each kinetic measurement, after running a blank spectrum with buffer (10 mM HEPES, pH 7.4), H2O2 (final concentration was 1 μM) was added into the plasmid DNA (final concentration was 63 μM), and then 30 spectra were recorded in a rate of 1 spectrum per minute.
Gel electrophoresis
A typical gel electrophoresis was set on 1.5% agarose gels with 0.5 × TBE running buffer (45 mM Tris, 4.5 mM boric acid, 1 mM EDTA) at 100 V. Thereafter, the gels were stained with Geneview, visualized and digitally photographed by Alphalimager HP Gel Imaging system.
Antioxidant assessment
Oxidant ABTS•+ was prepared by reacting ABTS with potassium persulfate according to the literature46. Using 46.7 μM ABTS•+ solution, the scavenging rate (SR) was calculated in percentage of its feature absorbance at 734 nm after 30-min reacting with different amounts of the antioxidant. IC50 value was estimated as the antioxidant concentration resulting in SR = 50%.
A similar method was used for the 0.1 mM DPPH oxidant. Instead, the feature absorbance at 520 nm was used to calculate the DPPH concentration47. IC50 value was estimated as the antioxidant concentration resulting in SR = 50%.
The in vitro ROS level was measured by the modified TBARS assay48. The mixture of antioxidant and Fenton’s reagent were vortexed and then settled for 30 min at 37 °C. TBA was dissolved in 10% trichloroacetic acid, with the final concentration of 0.67%. The solution was then heated in boiled water for 20 min. Finally the amount of TBARS formed in each sample was assessed with the absorbance at 532 nm.
The ferric-ferrous conversion was monitored with a modified colorimetric method, based on the absorbance of the Fe(II)-Ferrozine complex at wavelength of 562 nm.
Electrochemistry
Electrochemical measurement was performed with a CHI 660D electrochemical analyser, with a three-electrode system, i.e. a saturated AgCl electrode, a platinum wire, and a platinum disc electrode used as the reference, auxiliary, and working electrodes, respectively.
HPLC-MS analysis
The phosphorothioate oxidation was carried out in a mixed solution of 25 μM PS or 20 μM GsA-Rp and GsA-Sp with different concentrations of oxidants (H2O2 or Fenton reagents), respectively. The mixtures were settled for 2 hours in the dark at room temperature before the HPLC-MS analysis. In the analysis, 10 μl of reaction sample was loaded on Agilent’s C18 reversed phase column (150 × 4.6 mm, 1.5 μm), water and acetonitrile with 0.1% acetic acid were used as eluent A and eluent B in HPLC, respectively, and the flow rate was 0.3 ml/min with a gradient elution B from 5 to 95% in 10 min. Drying gas flow: 8 L/min; nebulizer pressure: 35 psi; drying gas temperature: 320 °C; capillary voltage: 3500 V (using the positive detection mode and scanning between 50 and 900 m/z for mass spectrometer analysis).
Quantum chemical calculation
All DFT (density functional theory) calculations were carried out with Gaussian 09 package49. Geometries were first optimized at the B3LYP/6-31 + G* level. In a typical calculation, the M06 and B3LYP levels of theory with the larger basis sets of 6-311 + G(d, p) were used to evaluate the geometric and energetic results in the gas phase. Solvent effect was considered with single point calculations of a continuum model (SMD) on reactants, transition states, intermediates, and products.
Additional Information
How to cite this article: Wu, T. et al. Mechanistic Investigation on ROS Resistance of Phosphorothioated DNA. Sci. Rep. 7, 42823; doi: 10.1038/srep42823 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Cao, B. et al. Genomic mapping of phosphorothioates reveals partial modification of short consensus sequences. Nat. Commun. 5, 3951–3979 (2014).
Wang, L. et al. DNA phosphorothioation is widespread and quantized in bacterial genomes. Proc. Natl. Acad. Sci. 108, 2963–2968 (2011).
Wang, L. et al. Phosphorothioation of DNA in bacteria by dnd genes. Nat. Chem. Biol. 3, 709–710 (2007).
Zhang, Y. C. et al. Theoretical study on steric effects of DNA phosphorothioation: B-helical destabilization in Rp-phosphorothioated DNA. J. Phys. Chem. B 116, 10639–10648 (2012).
Chen, L. et al. Theoretical study on the relationship between Rp-phosphorothioation and base-step in S-DNA: based on energetic and structural analysis. J. Phys. Chem. B 119, 474–481 (2015).
Karwowski, B. T. The influence of phosphorothioate on charge migration in single and double stranded DNA, a theoretical approach. Phys. Chem. Chem. Phys. 17, 21507–21516 (2015).
Lan, W. et al. Structural investigation into physiological DNA phosphorothioate modification. Sci. Rep. 6, 25737 (2016).
Cao, B. et al. In vivo analysis of phosphorothioate modification of DNA reveals substrate recognition by a multiprotein complex. Sci. Rep. 5, 12513 (2015).
Xiong, W., Zhao, G., Yu, H. & He, X. Interactions of Dnd proteins involved in bacterial DNA phosphorothioate modification. Front Microbiol 6, 1–7 (2015).
Zheng, T. et al. DndEi exhibits helicase activity essential for DNA phosphorothioate modification and ATPase activity strongly stimulated by DNA substrate with a GAAC/GTTC motif. J. Bio. Chem. 291, 1492–1500 (2016).
Zhou, X. et al. A novel DNA modification by sulphur. Mol. Microbio. 57, 1428–1438 (2005).
Gan, R. et al. DNA phosphorothioate modifications influence the global transcriptional response and protect DNA from double-stranded breaks. Sci. Rep. 4, 6642 (2014).
He, W. et al. Regulation of DNA phosphorothioate modification in Salmonella enterica by DndB. Sci. Rep. 5, 12368 (2015).
Cao, B. et al. Pathological phenotypes and in vivo DNA cleavage by unrestrained activity of a phosphorothioate-based restriction system in Salmonella. Mol. Microbiol. 93, 776–785 (2014).
Xie, X. et al. Phosphorothioate DNA as an antioxidant in bacteria. Nucleic Acids Res. 40, 9115–9124 (2012)
Dai, D. et al. DNA phosphorothioate modification plays a role in peroxides resistance in Streptomyces lividans . Front. Microbiol. 7, 1380 (1016).
Dizdaroglu, M. & Jaruga, P. Mechanisms of free radical-induced damage to DNA. Free. Radic. Res. 46, 382–419 (2012).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Kobayashi, K. & Tagawa, S. Direct observation of guanine radical cation deprotonation in duplex DNA using pulse radiolysis. J. Am. Chem. Soc. 125, 10213–10218 (2003).
Giese, B., Amaudrut, J., Köhler, A. K., Spormann, M. & Wessely, S. Direct observation of hole transfer through DNA by hopping between adenine bases and by tunneling. Nature 412, 318–320 (2001).
Valavanidis, A., Vlachogianni, T. & Fioktakis, C. 8-Hydroxy-2′-deoxyguanosine (8-OHdG): A critical biomarker of oxidative stress and carcinogenesis. J. Env. Sci. Health. C 27(2), 120–139 (2009).
Amir, A. et al. Nucleoside-5′-phosphorothioate analogues are biocompatible antioxidants dissolving efficiently amyloid beta-metal ion aggregates. Dalton. Trans. 41, 8539–8549 (2012).
Nordberg, J. & Arner, E. S. Reactive oxygen species, antioxidants, and the mammalian thioredoxin system. Free Radic. Biol. Med. 31, 1287–1312 (2001).
Masip, L., Veeravalli, K. & Georgiou, G. The many faces of glutathione in bacteria. Antioxid. Redox Signal. 8, 753–762 (2006).
Yarborough, A., Zhang, Y. J., Hsu, T. M. & Santella, R. M. Immunoperoxidase detection of 8-hydroxydeoxyguanosine in aflatoxin B1-treated rat liver and human oral mucosal cells. Cancer Res. 56, 683–688 (1996).
Abramoff, M. D., Magelhaes, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Butenko, N. et al. DNA cleavage activity of VIVO(acac)2 and derivatives. J. Inorg. Biochem. 103, 622–632 (2009).
Kupan, A. et al. Guanine oxidation by electron transfer: one- versus two-electron oxidation mechanism. Chembiochem. 7, 125–133 (2006).
Burrows, C. J. & Muller, J. G. Oxidative nucleobase modifications leading to strand scission. Chem. Rev. 98, 1109–1152 (1998).
Steenken, S. & Jovanovic, S. V. How easily oxidizable is DNA? One-electron reduction potentials of adenosine and guanosine radicals in aqueous solution. J. Am. Chem. Soc. 119, 617–618 (1997).
Dyguda-Kazimierowicz, E., Roszak, S. & Sokalski, W. A. Alkaline hydrolysis of organophosphorus pesticides: the dependence of the reaction mechanism on the incoming group conformation. J. Phys. Chem. B 118, 7277–7289 (2014).
Hagensee, M. E. & Moses, R. E. Repair response of Escherichia coli to hydrogen peroxide DNA damage. J. Bacteriol. 168, 1059–1065 (1986).
Zhou, X., Deng, Z., Firmin, J. L., Hopwood, D. A. & Kieser, T. Site-specific degradation of Streptomyces lividans DNA during electrophoresis in buffers contaminated with ferrous iron. Nuc. Acids Res. 10, 4341–4352 (1988).
Bergeron, F., Auvre, F., Radicella, J. P. & Ravanat, J. L. HO• radicals induce an unexpected high proportion of tandem base lesions refractory to repair by DNA glycosylases. Proc. Natl. Acad. Sci. 107, 5528–5533 (2010).
Mata, J. E. et al. Evidence of enhanced iron excretion during systemic phosphorothioate oligodeoxynucleotide treatment. J. Toxicol. Clin. Toxicol. 38, 383–387 (2000).
Giese, B. Long distance charge transport in DNA: The hopping mechanism. Acc. Chem. Res. 33(9), 631–636 (2000).
Kawanishi, S., Hiraku, Y. & Oikawa, S. Mechanism of guanine-specific DNA damage by oxidative stress and its role in carcinogenesis and aging. Mutation Research-Reviews in Mutation Research 488, 65–76 (2001).
Forman, H. J. & Torres, M. Reactive oxygen species and cell signaling: respiratory burst in macrophage signaling. Am. J. Respir. Crit. Care Med. 166, S4–8 (2002).
Luo, D., Smith, S. W. & Anderson, B. D. Kinetics and mechanism of the reaction of cysteine and hydrogen peroxide in aqueous solution. J. Pharm. Sci. Pharmacol. 94, 304–316 (2005).
Wu, L. et al. Desulfurization of phosphorothioate oligonucleotides via the sulfur-by-oxygen replacement induced by the hydroxyl radical during negative electrospray ionization mass spectrometry. J. Mass Spectrom. 47, 836–844 (2012).
Stawinski, J. & Kraszewski, A. How to get the most out of two phosphorus chemistries? Studies on H-phosphonates. Acc. Chem. Res. 35, 952–960 (2002).
Westheimer, F., Huang, S. & Covitz, F. Rates and mechanisms of hydrolysis of esters of phosphorous acid. J. Am. Chem. Soc. 110, 181–185 (1988).
Kotronarou, A., Mills, G. & Hoffmann, M. R. Oxidation of hydrogen sulfide in aqueous solution by ultrasonic irradiation. Environ. Sci. Technol. 26, 2420–2428 (1992).
Schippers, A. & Sand, W. Bacterial leaching of metal sulfides proceeds by two indirect mechanisms via thiosulfate or via polysulfides and sulfur. Appl. Environ. Microbiol. 65, 319–321 (1999).
Stewart, S. P. et al. Determination of hydrogen peroxide concentration using a handheld Raman spectrometer: detection of an explosives precursor. Forensic Sci. Int. 216, e5–8 (2012).
Re, R. et al. Antioxidant activity applying an improved ABTS radical cation decolorization assay. Free. Radic. Biol. Med. 26, 1231–1237 (1999).
Villano, D., Fernandez-Pachon, M. S., Moya, M. L., Troncoso, A. M. & Garcia-Parrilla, M. C. Radical scavenging ability of polyphenolic compounds towards DPPH free radical. Talanta 71, 230–235 (2007).
Bekdeser, B., Ozyurek, M., Guclu, K. & Apak, R. Novel spectroscopic sensor for the hydroxyl radical scavenging activity measurement of biological samples. Talanta 99, 689–696 (2012).
Frisch, M. et al. Gaussian 09. Revision A. 02. Gaussian. Inc., Wallingford, CT, 2009 (2009).
Acknowledgements
This work was supported by the National Basic Research Program of China “973” (No. 2013CB966802), National Science Foundation of China (Nos 21377085), MOE New Century Excellent Talents in University (No. NCET-12-0354). T.W. thank Dr. Timo Törmäkangas and Ms. Limeng Chen for the helpful discussion.
Author information
Authors and Affiliations
Contributions
Y.L.Z. and Z.D. conceived and designed the investigation. T.W. and H.Q. conducted the in vivo and in vitro experiments. X.L.W. and T.S. performed the quantum chemistry calculations. Y.L.Z., T.W., Z.W., J.L., and L.B. wrote up the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wu, T., Huang, Q., Wang, XL. et al. Mechanistic Investigation on ROS Resistance of Phosphorothioated DNA. Sci Rep 7, 42823 (2017). https://doi.org/10.1038/srep42823
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep42823
This article is cited by
-
Bio-responsive polymers for dual 31P/19F-magnetic resonance to detect reactive oxygen species in vivo
Communications Materials (2024)
-
Phosphorothioate-DNA bacterial diet reduces the ROS levels in C. elegans while improving locomotion and longevity
Communications Biology (2021)
-
Structural basis for the recognition of sulfur in phosphorothioated DNA
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.