Abstract
N6-Methyladenosine (m6A) is currently one of the most intensively studied post-transcriptional modifications in RNA. Due to its critical role in epigenetics and physiological links to several human diseases, it is also of tremendous biological and medical interest. The m6A mark is dynamically reversed by human demethylases FTO and ALKBH5, however the mechanism by which these enzymes selectively recognise their target transcripts remains unclear. Here, we report combined biophysical and biochemical studies on the specificity determinants of m6A demethylases, which led to the identification of an m6A-mediated substrate discrimination mechanism. Our results reveal that m6A itself serves as a ‘conformational marker’, which induces different conformational outcomes in RNAs depending on sequence context. This critically impacts its interactions with several m6A-recognising proteins, including FTO and ALKBH5. Remarkably, through the RNA-remodelling effects of m6A, the demethylases were able to discriminate substrates with very similar nucleotide sequences. Our findings provide novel insights into the biological functions of m6A modifications. The mechanism identified in this work is likely of significance to other m6A-recognising proteins.
Similar content being viewed by others
Introduction
All cellular RNAs undergo a range of post-transcriptional modifications, which are important mechanisms employed by nature to increase their structural and functional diversity1,2. To date, at least 100 chemically distinct modifications have been identified3,4, among which N6-methyladenosine (m6A) is currently one of the most important and most intensively studied epigenetic modifications5,6,7. It is universally conserved across all three domains of life, including archaea, bacteria and eukarya. In eukaryotes, m6A occurs predominantly as internal modification in messenger RNA (mRNA), where it is highly enriched near the stop codon and in the 3′-untranslated regions (3′UTRs)8,9,10. Recent methylome profiling studies in mouse and human revealed that m6A modification is pervasive throughout the transcriptome and is present in the transcripts of more than 7,600 coding genes and 300 non-coding genes. However, majority of the m6A modifications are uniquely distributed within the DR(m6A)CH consensus motif, where D denotes A, G or U, R denotes G or A and H denotes A, C or U11,12,13. The exact physiological relevance of m6A remains to be determined, although its widespread occurrence in mRNAs implies important roles associated with the regulation of gene expression and mRNA functions, such as alternative splicing, translation efficiency and mRNA stability14,15,16,17. It is also increasingly clear that dysregulation of pathways controlled by m6A modifications may underlie the pathogenesis of a range of human diseases, such as obesity18,19,20,21,22,23, neurological disorders24 and, potentially, male infertility25.
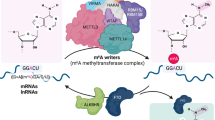
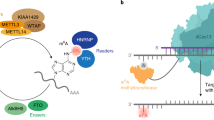
The m6A landscape is dynamically regulated by a complex interplay between various families of m6A-specific proteins, termed ‘writers’, ‘readers’ and ‘erasers’, which add, interpret and remove the m6A mark, respectively5. For instance, the addition of N6-methyl group on adenosine is catalysed by m6A methyltransferase complexes, such as METTL3-METTL14-WTAP26,27 and this can be directly reversed by human m6A demethylases FTO (fat mass and obesity-associated protein)28 and ALKBH5 (AlkB homologue 5)25, both of which are medically-important enzymes belonging to the iron- and 2-oxoglutarate (2OG)-dependent family of AlkB oxygenases (Fig. 1)18,29. The FTO and ALKBH5 are highly specific for m6A, with little or no activity for other methylated-nucleotides, such as N1-methyladenosine (m1A, a cytotoxic lesion in DNA) and 5-methylcytosine (m5C, another ubiquitous epigenetic modification found in DNA and RNA)30,31. Although FTO is also able to demethylate N3-methylthymidine (m3T; kcat/Km ~0.007 min−1μM−1) and N3-methyluracil (m3U; kcat/Km ~0.014 min−1μM−1), it does so with significantly lower efficiency compared with m6A substrates (kcat/Km ~0.3–0.8 min−1μM−1) (Figs 1 and 2)28,30,32. Hence, to date, m6A is the only known physiologically-relevant substrate for FTO and ALKBH5. The factors determining their ‘methylated-nucleotide specificity’ remain to be elucidated, although we30 and others29,32,33,34, have recently shown that at least part of their specificity could be due to distinct structural features within their nucleotide-binding sites and catalytic domains, which enables multiple specific interactions with m6A (Fig. 1). It is also beginning to emerge that adjunct structural elements, such as the nucleotide-recognition lid domain and the L1 loop, which are unique to FTO and ALKBH5, are likely important for their ‘methylated-nucleotide specificity’34.
Oxidative demethylation reactions catalysed by FTO and ALKBH5.
(a) In the removal of m6A mark by FTO, the N6-methyl group is first oxidised to a hydroxymethyl group, which fragments to give formaldehyde and the demethylated base. ALKBH5 likely catalyses the direct removal of the N6-methyl group from m6A. m3T and m3U are also demethylated by FTO, but with significantly lower efficiency28,30,31. Superimposition of views from the crystal structure of AlkB-m6A complex (green stick, PDB ID 4NID)28 with a structure of (b) FTO (salmon residues) (PDB ID 4CXW)30 and (c) ALKBH5 (orange residues) (PDB ID 4NJ4)29. The multiple specific interactions between m6A and the active sites likely enable ‘methylated-nucleotide specificity’.
Substrate selectivity and sequence preference of human m6A demethylases FTO and ALKBH5.
(a) The extent of demethylation of substrates was determined by HPLC after a 1 hour (FTO) or 30-minute (ALKBH5) incubation at 37 °C, pH 7.4. Full length human FTO and catalytically-active human ALKBH566–292 were used. Steady-state kinetics analyses of the demethylation of consensus (6, 10 and 11) and non-consensus m6A substrates (13) by (b) FTO and (c) ALKBH5. The Km and kcat values were determined by keeping a constant FTO or ALKBH5 concentration of 0.5 μM. Errors represent S.D. of three replicates.
The specificity of FTO and ALKBH5 is not only observed at the nucleotide level, it is also apparent at the transcript level. This is evident from the observation that despite ubiquitous expression of both FTO and ALKBH5 in mammalian cells, only a fraction of m6A sites is found to be demethylated in any given mRNA24. Thus the activity of FTO and ALKBH5 is likely transcript-specific, where only certain selected m6A-containing transcripts are being actively demethylated by the enzymes. Consistent with this proposal, FTO and ALKBH5 have highly distinct physiological functions. In particular, FTO has been shown in several studies to be strongly linked with obesity35,36, while ALKBH5 appears to be essential for spermatogenesis25. In order to achieve such distinct biological functions, FTO and ALKBH5 likely target certain mRNA specifically for demethylation.
Conceivably, the substrate specificity of FTO and ALKBH5 may be facilitated through subcellular localisation of the proteins and/or their differential expression in different tissues (FTO is most abundantly expressed in the hypothalamus18 whereas ALKBH5 is highly expressed in the testis25). It is also possible that the biological system may tolerate some off-target m6A demethylation, particularly in view that m6A is a reversible modification. However, additional mechanisms are likely in place to enable FTO and ALKBH5 to recognise specific m6A-sites.
A number of studies37,38,39,40,41 have demonstrated that m6A methylation can directly impact the thermodynamic stability and conformations of DNA/RNA. In particular, recent studies by Micura et al.38 on various self-complementary duplexes showed that m6A has a general destabilising effects on duplex base-pairing and may promote secondary structure change in certain sequence contexts. The structural influence of m6A is also evident in cellular RNA and appears to be physiologically-relevant. For instance, recent study investigating the structural profiles of RNA in living cells revealed that m6A-modified sites exhibit specific structural signatures; a loss of m6A modifications (through mettl3 knockout) was accompanied by a significant loss of these structural signatures42. This observation concurs with transcriptome-wide RNA structural mapping work by Kool et al.39 where sites adjacent to m6A were generally found to have a strong tendency towards unpaired structure. In a separate study, it was found that m6A modification on MALAT1 (Metastasis Associated Lung Adenocarcinoma Transcript 1; a lncRNA) caused significant alteration to its local structure, which likely serve to facilitate the binding of its regulatory protein HNRNPC43,44. Inspired by these interesting observations, we envisaged that m6A-induced conformational change could provide a basis for substrate discrimination by m6A demethylases.
Here, we report combined thermodynamic, spectroscopic, gel-shift, thermophoretic and biochemical studies on the determinants of substrate specificity for human m6A demethylases. Our results reveal that FTO and ALKBH5 do not exhibit strict sequence requirements for substrate specificity and the highly-conserved GG(m6A)CU consensus motif is, unexpectedly, not a crucial determinant for selectivity. Our results further reveal that m6A serves as a ‘conformational marker’ which dynamically regulates the overall conformation of the modified RNA, and, hence, the substrate selectivity of m6A demethylases. Remarkably, the introduction of m6A modification induces different conformational outcomes in different RNAs sequences and this profoundly impact their interactions with m6A-recognising proteins, including FTO and ALKBH5. Through the structural message encoded by the ‘m6A mark’, FTO and ALKBH5 are not only able to discriminate substrates with very similar primary nucleotide sequences, but also those that contain the same consensus motif. Our findings, therefore, provide new insights into the biological functions of m6A methylation. The unique recognition strategy identified in this work is likely of significance to other m6A-recognising proteins and, more widely, other RNA-binding proteins.
Results and Discussion
Substrate selectivity of FTO and ALKBH5 is not strictly dependent on specific recognition of m6A consensus motif
To date, the substrate preferences and sequence requirements of the m6A demethylases have not been systematically studied. It is not clear if the G(m6A)C and A(m6A)C consensus motifs (where m6A predominantly resides) are essential for substrate recognition. Moreover, besides mRNA, recent studies have also identified m6A modifications in non-coding RNAs, such as transfer RNA (tRNA), ribosomal RNA (rRNA), small nuclear RNA (snRNA) and long non-coding RNA (lncRNA), where they are not found within the same consensus motifs45,46,47. These findings raise the question of whether FTO and ALKBH5 are able to accept m6A on non-consensus sites. Conceivably, other unidentified m6A demethylases may exist which specifically regulate m6A marks on non-coding RNAs.
To explore these interesting questions and to clarify the substrate specificity of FTO and ALKBH5, we analysed the activities of FTO and ALKBH5 against a series of m6A-containing oligonucleotides using a HPLC-based assay (Fig. 2). The m6A substrates investigated consist of short DNAs and RNAs of varying lengths and sequences (Fig. 2a). They are either based on the m6A consensus motifs G(m6A)C (i.e. 2, 4–9, 11, 15) and A(m6A)C (i.e. 3, 10, 12), or are based on random sequences (i.e. 13, 14). We initially determine the minimum substrate length that is required for enzyme recognition. Our results revealed that the m6A nucleotide 1 itself is a very poor substrate for FTO and ALKBH5 (Fig. 2a). The 3-mer core consensus motifs, G(m6A)C 2 and A(m6A)C 3, also gave negligible demethylation yields (~2–5%), even after prolonged incubation (Fig. 2a). This implies that the residues surrounding the m6A site are likely involved in crucial interactions with the active sites of FTO and ALKBH5. Indeed, the addition of either a guanosine residue at position −2 (relative to m6A in 2 i.e. 4) or a uridine residue at position +2 (i.e. 5) resulted in marked improvements in demethylation yields with both enzymes (~19–31%; Fig. 2a). Interestingly, short 5-mer substrates, such as 6, 8–10 (~64–70%) were found to have similar FTO demethylation yields as that of longer 14-mer substrate 11 (~78%, Fig. 2a), thus the minimum sequence that can be recognised by FTO appears to be only five nucleotides in length. Consistent with these results, kinetic analyses with FTO revealed substantial activity for 5-mer 6 (kcat/Km = 0.68 min−1μM−1), which is comparable to that of 14-mer 11 (kcat/Km = 0.77 min−1μM−1) (Fig. 2a,b and Supplementary Fig. S1). ALKBH5 also exhibited a similar activity profile, although it showed a slight preference for 11 (kcat/Km = 0.098 min−1μM−1) over 6 (kcat/Km = 0.060 min−1μM−1, Fig. 2a,c and Supplementary Fig. S1). Notably, a guanosine residue at +2 position to m6A is clearly disfavoured, as indicated by the significant reduction in activity for 7 by FTO (~11%) and ALKBH5 (~9%). This concurs with the DR(m6A)CH consensus motif, where ‘H’ is never found to be a guanosine residue. Overall, our results suggest that the short, 5-mer GG(m6A)CU sequence is likely sufficient to define a demethylation site for both FTO and ALKBH5.
We next examine if the m6A demethylases demonstrate any preference for either of the two consensus motifs G(m6A)C and A(m6A)C. Our assay data indicate that FTO and ALKBH5 are able to demethylate GG(m6A)CU 6 and GA(m6A)CA 10 with comparable efficiencies (Fig. 2b,c). In agreement with this result, the demethylation yields for 14-mer G(m6A)C-based substrate 11 were also found to be similar to that for 14-mer A(m6A)C-based substrate 12 (Fig. 2a), implying that m6A demethylases likely do not discriminate between the two consensus motifs. The m6A methyltransferases, on the contrary, strictly favour G(m6A)C over A(m6A)C consensus motifs48,49, consequently G(m6A)C-based sequences are, depending on species, two- to twelve-fold more abundant than A(m6A)C-based sequences8,9,10.
Our results further revealed that FTO and ALKBH5 do not discriminate between RNA and DNA substrates, as shown by their substantial activities towards both DNA substrates 6 and its RNA equivalent 9. Importantly, both FTO and ALKBH5 are able to recognise m6A modifications on non-consensus sites. This is clearly demonstrated by their significant catalytic activities towards the arbitrary RNA sequence 13 (kcat/Km (FTO) = 0.39 min−1μM−1 and kcat/Km (ALKBH5) = 0.053 min−1μM−1), which, notably, is only ~2-fold lower than consensus substrate 11 (Fig. 2 and Supplementary Fig. S2). In addition, there are also considerable demethylase activities towards the non-consensus DNA substrate 14 (~63% (FTO), ~32% (ALKBH5), Fig. 2a and Supplementary Fig. S2). These observations are consistent with a recent report which showed that ALKBH5 is able to demethylate the non-consensus sequence rAUUGUCU(m6A)UUGCAGC, although with reduced demethylation yields (~20%)25.
Taken together, our results suggest that the GG(m6A)CU consensus motif, while preferred, is not absolutely essential for substrate recognition by FTO and ALKBH5. It is also apparent that FTO and ALKBH5 have the potential for a high degree of promiscuity, although the efficiency of m6A demethylation varies. Hence, sequence information alone is likely insufficient in regulating the substrate specificity of FTO and ALKBH5. We, therefore, propose that additional mechanisms that do not depend exclusively on the specific recognition of the primary nucleotide sequence are likely involved.
m6A modification destabilises RNA duplexes
We considered the possibility that m6A methylation might modulate substrate specificity of m6A demethylases by fine-tuning the conformation of the modified RNA. Precedence for this possibility comes from observations that conceptually similar epigenetic modifications, in particular N-methylation on histone lysine and arginine frequently result in remodelling of chromatin structures, which alters their interactions with DNA50,51. This is further supported by the number of studies (highlighted in the introduction) which showed that m6A modification can directly impact the secondary structure of RNAs both in vitro and in vivo37,38,39,40,41,42,43,44. To the best of our knowledge, to date, m6A has not been explored as a potential determinant of substrate specificity for FTO and ALKBH5.
We begin by studying the effect of m6A modification on RNA conformations. To this end, we employed several 12-mer palindromic RNAs 16–23 as model sequences (Table 1 and Supplementary Table S1). Due to their self-complementary nature, they can inherently adopt two main secondary structures in solution, namely (1) duplex conformation, by engaging in intermolecular base pairing and (2) hairpin conformation, by folding back on themselves. Such structural versatility enables them to mimic the dynamic RNA structures observed under physiological conditions, hence they are particularly suited for our study.
We first examine the secondary structure of the unmethylated RNA 16 (rCCGGAAUUCCGG) by performing UV-melting analysis under physiologically relevant conditions (i.e. in 10 mM sodium phosphate buffer containing 150 mM NaCl, pH 7.4). At a total strand concentration of 5 μM, 16 showed a monophasic, sigmoidal melting profile, indicating the presence of a single structural species (Fig. 3a). In addition, van’t Hoff plot over a concentration range of 1–100 μM 16 revealed melting temperatures (Tm) that are linearly dependent on strand concentration, suggesting that 16 likely exists as a bimolecular duplex structure (Fig. 3d and Supplementary Fig. S3). This is supported by CD analysis of 16, which shows a dominant positive UV absorption band at 264 nm and a negative absorption band at 212 nm, which are characteristics of an A-form double helix structure (Fig. 3g). The strong hyperchromicity observed in the CD spectrum and UV-melting profiles of 16 further suggests that the duplex structure of 16 is likely to be extensively, if not fully, based-paired.
Effects of m6A modification on the overall conformation of model RNA sequences.
(a–c) UV-melting profiles of the methylated RNA sequences and their corresponding reference sequences (colour-coded) at 5 μM strand concentration. The presence of m6A in 19 and 21 triggered a dramatic duplex to hairpin transformation, as indicated by the biphasic melt curves. (d–f) Van’t Hoff analyses showed dependence of melting temperatures on strand concentrations. At concentrations <5 μM, 19 and 21 gave invariable Tm of 47.5 °C and 45 °C, respectively due to hairpin formation. (g–i) Overlay of the CD spectra of methylated and unmethylated pair (both at 5 μM strand concentration) indicates significant conformational change on m6A modification of 18 and 20.
To determine the structural influence of m6A, we replaced the adenine base at strand position 5 of 16 with an m6A residue, which generated 17 (rCCGG(m6A)AUUCCGG) (Table 1). As with its unmethylated form, 17 was also found to assume an A-form duplex structure, as demonstrated by a monophasic, concentration-dependent melting profile (Fig. 3a,d and Supplementary Fig. S3). In addition, the CD spectrum of 17 superimposes well with that of 16, implying that m6A modification did not cause any significant conformational change in 17 (Fig. 3g). The presence of m6A, however, did cause an overall destabilisation of the duplex structure of 17 (Tm = 58.0 °C;  = −13.9 kcal/mol), as indicated by its less favourable free Gibbs enthalpy compared with that of unmethylated duplex 16 (Tm = 63.0 °C;
= −13.9 kcal/mol), as indicated by its less favourable free Gibbs enthalpy compared with that of unmethylated duplex 16 (Tm = 63.0 °C;  = −15.4 kcal/mol, Table 1). The magnitude of destabilisation caused by a single m6A base is ~1.5 kcal/mol, which is consistent with measurements by others37,38,39,40,41. In particular, recent work by Kool et al.38 on multiple sequence contexts indicates that the amount of destabilisation per m6A base ranges from 0.5–1.7 kcal/mol, depending on the level of m6A substitution and sequence contexts. Early study by von Hippel et al.37 on m6A-containing RNA polymers also indicates a similar level of destabilisation (0.35–0.95 kcal/mol).
= −15.4 kcal/mol, Table 1). The magnitude of destabilisation caused by a single m6A base is ~1.5 kcal/mol, which is consistent with measurements by others37,38,39,40,41. In particular, recent work by Kool et al.38 on multiple sequence contexts indicates that the amount of destabilisation per m6A base ranges from 0.5–1.7 kcal/mol, depending on the level of m6A substitution and sequence contexts. Early study by von Hippel et al.37 on m6A-containing RNA polymers also indicates a similar level of destabilisation (0.35–0.95 kcal/mol).
Thermodynamic analysis indicates that the observed duplex destabilisation is primarily due to a less exothermic enthalpy (ΔΔH° = 2.5 kcal/mol) which counteracts the favourable change in entropy (ΔΔS° = 2.9 cal/mol/K) (Table 1). In view of the lack of major conformational change between 16 and 17 (Fig. 3g), the introduction of m6A in 17 likely did not cause significant disruption of its duplex base-pairing. Notably, recent NMR studies suggests that m6A likely base pair with U in a Watson-Crick manner in double-stranded RNA38. Apparently, this is achieved via rotation of the N6-methylamino group from its energetically preferred syn-conformation (with respect to N1 in the purine ring) to the higher-energy anti-conformation. Although this orientation is partially compensated for by hydrogen bonding and Watson-Crick geometry matching with U, the resulting m6A·U base pair is expected to be intrinsically unstable and this introduces an element of instability in the duplex structure.
m6A modification discriminates RNA sequences by triggering different conformational outcomes
The above results are interesting because they imply that m6A can, in principle, trigger an overall conformational change in RNA if it occurs on a relatively unstable duplex, where the energy cost to form alternative structures will be relatively low. As a proof of principle, we designed a thermodynamically less stable analogue of 17 by replacing both its ‘CCGG’ segments (rCCGGAAUUCCGG; underlined) with ‘CGCG’, to generate rCGCGAAUUCGCG 19 (Tm = 55.0 °C;  = −12.6 kcal/mol, Table 1). 19 is expected to be thermodynamically less stable compared to 17 due to less favourable nearest neighbour effects52.
= −12.6 kcal/mol, Table 1). 19 is expected to be thermodynamically less stable compared to 17 due to less favourable nearest neighbour effects52.
As anticipated, the UV-melting analysis of 19 revealed an interesting biphasic melting profile, indicating the presence of two structural species under the experimental conditions (Fig. 3b). The first melt transition at Tm ~47.5 °C has significantly lower hyperchromicity (~14%) than generally observed for a duplex transition, implying a reduced number of base-pairs in this structure. Moreover, van’t Hoff analysis revealed a relatively constant Tm over 1–5 μM i.e. the concentration range at which the first melt transition was detected (Fig. 3e and Supplementary Fig. S4). These observations are indicative of a monomolecular hairpin structure. In contrast, the second melt transition (Tm ~ 55.0 °C), which dominates the melting profile of 19 at concentrations above 10 μM, showed strong hyperchromicity (~21%) and a concentration-dependent Tm, which are suggestive of a duplex transition (Fig. 3b,e and Supplementary Fig. S4). Hence, 19 likely exists as both hairpin and duplex structures under our experimental conditions. These data concur with the CD spectrum of 19 which showed characteristics of both A-form duplex and B-form hairpin (Fig. 3h).
We were unable to confirm the presence of hairpin structure with 1H NMR study as this species was only formed at ≤5 μM. However, poly-acrylamide gel electrophoresis (PAGE) of 19 under non-denaturing conditions clearly revealed the existence of both duplex (lower mobility band) and hairpin structures (higher mobility band), particularly at low strand concentrations (Fig. 4a). Notably, the ‘hairpin bands’ are inherently faint compared with the ‘duplex bands’, due to the lower number of base-paired sites that can associate with the staining dye. We estimated that at least 20% and 35% of 19 were present as hairpin conformation at 5 μM and 2.5 μM strand concentrations, respectively. Importantly, there was no evidence of hairpin structure in the native PAGE and UV-melting analyses of unmethylated 18 (Figs 3b and 4a). Hence, hairpin formation in 19 was clearly triggered by m6A modification and likely via duplex-to-hairpin conversion.
m6A-induced conformational change regulates the substrate selectivity of FTO and ALKBH5.
(a) Non-denaturing PAGE analysis at 4 °C showed significant hairpin formation in 19 (~20–35% hairpin conversion) and 21 (~30% hairpin conversion) at 5 μM and 2.5 μM strand concentrations, but not in 17 (b) This was accompanied by a dramatic increase in the demethylation yields of 19 and 21, in sharp contrast to 17. Notably, hairpin formation was not observed for their unmethylated equivalents 16, 18 and 20, implying that hairpin formation was triggered by the presence of m6A. RNAs 24, 11 and 25 are sequences with duplex, random coil and hairpin structure, respectively. The GG(m6A)CU consensus motifs are highlighted in yellow. Errors represent S.D. of three replicates.
To obtain additional insights into factors determining m6A-induced conformational change, we further analysed 21 (rCGCGU(m6A)UACGCG), an analogue of 19 where the bases within the centre of the palindrome (underlined) were switched from ‘(m6A)AUU’ to ‘U(m6A)UA’ (Table 1). Interestingly, m6A-induced duplex-hairpin conversion was again observed in the native PAGE analysis of 21, where similar levels of hairpin formation (~30%) were detected at ≤5 μM (Fig. 4a). This observation was verified by UV-melting and CD analyses (Fig. 3 and Supplementary Fig. S5). The thermodynamic parameters derived from UV-melting experiments suggest that duplex-hairpin transformation in 21 is primarily an entropy-driven process, as apparent from the highly favourable entropy change (ΔΔS° = 86.1 cal/mol/K) (Table 1). However, due to the large enthalpy cost (ΔΔH° = 38.3 kcal/mol), the hairpin form of 21 is only marginally more stable than its duplex form at 37 °C ( = −1.3 kcal/mol). These data concur with those obtained for the hairpin and duplex structures of 19 (ΔΔH° = 37.5 kcal/mol, ΔΔS° = 86.0 cal/mol/K,
= −1.3 kcal/mol). These data concur with those obtained for the hairpin and duplex structures of 19 (ΔΔH° = 37.5 kcal/mol, ΔΔS° = 86.0 cal/mol/K,  = −1.8 kcal/mol).
= −1.8 kcal/mol).
Our combined results, therefore, demonstrate that the introduction of a single N6-methyl group on adenine is sufficient to induce a major overall conformational change in RNA. In addition, the conformational outcomes of m6A modification is highly dependent on sequence context. In RNAs 17, 19 and 21, m6A-induced duplex-hairpin transformation was only observed for 19 and 21 and not for 17, despite all three RNAs having the same base composition and highly similar primary nucleotide sequences. Hence, a relatively limited sequence change can have a profound effect on the overall conformation of the RNA and this may provide a molecular basis for substrate selectivity of m6A demethylases.
m6A is an important molecular determinant of substrate specificity for FTO and ALKBH5
To understand the role of m6A in regulating the substrate specificity of m6A demethylases, we profiled the activities of FTO and ALKBH5 against 17, 19 and 21 using a HPLC-based assay. The assay results are summarised in Fig. 4. Overall, the activity profile is highly consistent with the structural influence of m6A on these sequences. In particular, both FTO and ALKBH5 were able to recognise and accept 19 and 21, but not 17 as substrates, suggesting that their substrate selectivity is indeed, at least partially, regulated by m6A-induced structural change (Fig. 4 and Supplementary Figs S3–S8). In line with this observation, the level of demethylation of 19 and 21 was also found to approximately correlate with the extent of m6A-induced hairpin formation under our assay conditions. At 10 μM strand concentration, where both 19 and 21 were observed to exist predominantly as duplexes, there was little or no demethylation of both substrates by FTO and ALKBH5 (Fig. 4). However, when the concentration was reduced to 5 μM and 2.5 μM, where hairpin formation was appreciable, the demethylation yields of 19 (~45% (FTO), ~25% (ALKBH5)) and 21 (~37% (FTO), ~24% (ALKBH5)) increased significantly (Fig. 4, Supplementary Figs S6 and S7). Moreover, in a negative control experiment, we observed negligible demethylase activity towards the unmethylated analogues 16, 18 and 20 at all concentrations tested (Fig. 4b). It is thus clear that m6A can critically impact the selectivity of FTO and ALKBH5 by modulating the conformation of m6A substrates. Conceptually, this mechanism could enable FTO and ALKBH5 to distinguish their bona fide targets from other potential m6A substrates, including those with very similar primary nucleotide sequences. We further postulate the structural influence of m6A may also facilitate the discrimination of substrates with the same consensus motif.
To investigate this possibility, we evaluated the activity of FTO and ALKBH5 against 15 (rGCGG(m6A)CUAGUCCGC), a palindromic substrate containing the GG(m6A)CU consensus motif (underlined). Remarkably, 15 is an extremely poor substrate for both enzymes (demethylation yields ~3% (FTO), ~4% (ALKBH5)) even though it contains the m6A consensus motif. To rationalise this result, we analysed the conformation of 15 and its binding interactions with FTO and ALKBH5. Contrary to other palindromic sequences investigated in this study, such as 19 and 21, m6A methylation of 15 did not result in any detectable duplex-hairpin conversion. Both 15 and its unmethylated analogue 22 were found to exist almost exclusively as A-form duplex structures, as determined by native PAGE, CD and UV-melting analyses (Supplementary Figs S9–S11). Apparently in its duplex form, 15 showed very poor affinity for FTO and ALKBH5, as demonstrated by biotin-labelled electrophoretic mobility shift assay (EMSA), where there was no detectable binding of biotin-15 to FTO and ALKBH5, even at 1250-fold excess of proteins (Supplementary Fig. S12). Hence, 15 was not recognised and accepted as substrate by both m6A demethylases. This enables the discrimination of 15 from other substrates containing the same consensus motif, as exemplified by 11 (rGCGG(m6A)CUCCAGAUG) and 25 (rGCGG(m6A)CUCCACCGC) (Fig. 4). In the sequence contexts of 11 and 25, m6A modification promotes a random coil and hairpin conformations, respectively, both of which are able to bind significantly stronger with FTO and ALKBH5 than duplex 15 (Supplementary Figs S11–S13). Consequently, 11 and 25 are selectively targeted by m6A demethylases (Fig. 4). Results from microscale thermophoresis (MST)-based experiments53,54 indicate that FTO and ALKBH5 have similar binding affinities for 11 (KD (FTO) ~ 97.1 μM; KD (ALKBH5) ~ 52.9 μM) and 25 (KD (FTO) ~ 91.3 μM; KD (ALKBH5) ~ 75.3 μM) (Figs 5 and 6), although their demethylation yields are significantly higher for 11 than 25 (Fig. 4). This implies that the catalysis of hairpin substrate is likely slower compared with single-stranded substrates.
Intriguingly, selectivity for 11 over 15 was also observed for other m6A-binding proteins, in particular the YTH-domain proteins YTHDF255, which showed significant binding with biotin-11, but very little or no binding to biotin-15 (Supplementary Fig. S14). Our combined results revealed that m6A achieves substrate selectivity by regulating the affinity of m6A-recognising proteins for their targets.
Conclusions
Overall, we use a combination of thermal denaturation studies, CD analyses, gel-shift techniques, microscale thermophoresis measurements and biochemical assays to investigate the factors that modulate the substrate specificity of human m6A demethylases. Consistent with reports by others37,38,39,40,41,42,43,44, our results reveal that m6A modification has a general destabilising effect on RNA duplexes. We showed that although the magnitude of destabilisation by m6A is relatively small, it could induce a major overall conformational change in certain sequence context. This is clearly demonstrated by oligos 19 and 21, where the presence of a single m6A modification is sufficient to trigger a remarkable transformation from duplex to hairpin structure.
Importantly, we revealed, through direct biophysical evidence, that such m6A-induced conformational change on RNA (or lack of) could critically influence its interactions with several m6A-recognising proteins, including FTO, ALKBH5 and YTHDF2. This provides, at least partially, a strategy by which these proteins achieve substrate specificity. In particular, through the remodelling influence of m6A, both FTO and ALKBH5 are able to selectively recognise different m6A substrates, including those with highly similar primary nucleotide sequences, as demonstrated by their distinct selectivity for 19 and 21, over 17. Intriguingly, this mechanism also likely enables the discrimination of substrates with the same consensus motif, as shown by the lack of activity of FTO and ALKBH5 for 15 compared with 11 and 25, even though all three sequences contain the GG(m6A)CU consensus motif. Thus m6A likely serves as a ‘conformational marker’ which dynamically regulates the substrate selectivity of m6A demethylases.
Unexpectedly, the highly-conserved GG(m6A)CU consensus motif, which is widely assumed to be essential, is not crucial for substrate selectivity. This is demonstrated by the significant activities of FTO and ALKBH5 towards random sequences 13 and 14 which do not contain the consensus motif. Such apparent lack of a strict sequence requirement by the demethylases is in sharp contrast to other m6A-recognising proteins, such as m6A methyltransferase METTL337,38 and m6A-binding proteins YTHDF254, where the GG(m6A)CU consensus motif is strongly preferred.
To our knowledge, this is the first report demonstrating that m6A itself serves as an important selectivity determinant for m6A demethylases. This result is likely of significance to other m6A-recognising proteins and, more widely, other RNA-binding proteins. It is, however, important to appreciate that the identified mechanism alone is unlikely to discriminate m6A modifications on unstructured RNAs. It is also unable to account for differences in specificity between FTO and ALKBH5. Several mechanisms are probably involved which collectively shape the overall selectivity profile of the enzymes. In light of the structure-determining property of the m6A mark37,38,39,40,41,42,43,44, an interesting question will be whether the physiological functions of m6A demethylases are mediated through dynamic remodelling of their respective RNA transcripts.
Material and Methods
Synthesis and purification of DNA/RNA sequences
The DNA/RNA oligonucleotides used in this study were synthesised using standard β-cyanoethyl phosphoramidite chemistry. All synthesiser reagents and phosphoramidites were purchased from Glen Research. In brief, the oligonucleotides were synthesised on a solid support by the automated DNA/RNA synthesiser (Applied Biosystems 394) using a standard 1.0 μmole phosphoramidite cycle of acid-catalysed detritylation, coupling, capping and iodine oxidation. Cleavage of the oligonucleotides from the solid support and deprotection was achieved by exposure to a 1:1 mixture of 28% aq. ammonium hydroxide and 40% aq. methylamine for 10 min at 65 °C. Deprotection of the 2’-O-TBDMS group and initial purification were carried out with Glen-Pak RNA purification cartridges according to the manufacturer’s procedure. The crude products were purified by reverse-phase HPLC using the Waters XBridge OST C18 column (2.5 micron, 10 mm × 50 mm). HPLC solvents used were: solvent A (100 mM triethylammonium acetate buffer, pH 6.5 with 5% acetonitrile) and solvent B (100 mM triethylammonium acetate buffer, pH 6.5 with 15% acetonitrile) with a flow rate of 5 mL/min. All purified oligonucleotides were characterized by MALDI-MS and capillary gel electrophoresis and were found to be at least 95% pure (Supplementary Table S1).
Expression and purification of human FTO, ALKBH5 and YTHDF2
Full length human FTO, human ALKBH566–292 and human YTHDF2385–576 were expressed and purified as previously reported, with modifications30. In brief, all constructs were transformed into E. coli BL21 (DE3) Rosetta cells. The transformed cells were grown at 37 °C until an OD600 of 0.6 was reached. Protein expression was then induced with isopropyl β-D-1-thiogalactopyranoside (IPTG, 0.5 mM, Gold Biotechnology). Cell growth was continued at 16 °C for 16 h, after which the cells were harvested by centrifugation and the resulting cell pellet was stored at −80 °C. The frozen cell pellets were then thawed, resuspended in lysis buffer and disrupted by French Press. Further purification of the protein was achieved using Ni affinity chromatography and gel filtration, as described below. Full length human FTO was sub-cloned into pNIC28-Bsa4 to generate a His6-tagged FTO1–505 construct. FTO in lysis buffer (25 mM Tris, pH 7.5, 500 mM NaCl, 40 mM imidazole and 5 mM β-mercaptoethanol (β-ME)) was purified using Ni affinity chromatography (GE healthcare), followed by gel filtration using HiLoad superdex 200 26/60 (GE healthcare) into the final buffer (25 mM Tris buffer, pH 7.5, 100 mM NaCl, 5% (v/v) glycerol and 5 mM β-ME). For human ALKBH5, a His6-tagged ALKBH566–292 construct in pNIC28-Bsa4 was used. ALKBH5 in lysis buffer (25 mM Tris, pH 8.0, 500 mM NaCl, 40 mM imidazole and 5 mM β-ME) was first purified using Ni affinity chromatography (GE healthcare), followed by anion chromatography using a 5 mL HiTrap Q HP column (GE healthcare) and gel filtration using HiLoad superdex 75 16/60 (GE healthcare) into the final buffer (20 mM Tris buffer, pH 8.0, 100 mM NaCl and 5 mM β-ME). Recombinant human YTHDF2385–576 was cloned into pGEX-6P-1 from cDNA clones and transformed into E. coli BL21 (DE3) Rosetta cells. Purification of YTHDF2 in lysis buffer (50 mM Tris, pH 8.0, 500 mM NaCl, 40 mM imidazole and 5 mM β-ME) was achieved by loading the supernatant to a 5 mL GSTrapTM HP column (GE healthcare). The protein was eluted with gluthione containing buffer (50 mM Tris, pH 8.0, 300 mM NaCl, 10 mM glutathione and 5 mM β-ME). Further purification was achieved by gel filtration using HiLoad superdex 200 26/60 (GE healthcare) into the final buffer (20 mM Tris, pH 8.0, 150 mM NaCl and 5 mM β-ME).
HPLC-based demethylase assay
The assay was modified from previously reported methods30,56. The assay was performed in triplicate for each m6A substrate, in a final reaction volume of 25 μL. Reaction consisted of FTO (2 μM) or catalytically active ALKBH566–292 (4 μM), 2-oxoglutarate (300 μM), (NH4)2Fe(SO4)2·6H2O (150 μM), L-ascorbate (2 mM), m6A-containing DNA/RNA (substrate, 10 μM) in 50 mM HEPES buffer, pH 7.4. The reaction was incubated at 37 °C for 30 min, after which the m6A-containing DNA/RNA was digested by treatment with 1 Unit of nuclease P1 in buffer containing 7 mM of sodium acetate and 0.4 mM of ZnCl2 at 37 °C for 2 h. This was followed by the addition of 1M NH4HCO3 (1 μL) and alkaline phosphatase (1 Unit). After further incubation at 37 °C for 1.5 h, an internal standard (10 μM, uridine or thymidine for DNA and RNA substrates, respectively) was added to the reaction mixture and the solution was analysed on a HPLC system. The nucleosides were separated using a Zorbax C18 column (4.6 mm × 250 mm) with a gradient of 98% solvent A (MilliQ water + 0.1% TFA) to 100% solvent B (methanol) over 25 min, at a flow rate of 1.0 mL/min at room temperature. The UV detection wavelength was set at 266 nm. Controls without enzyme were also set up. The percentage of demethylation was calculated based on the peak areas of N6-methyladenine (m6A) in the samples and in their respective controls.
Steady-state kinetics of m6A demethylation by FTO and ALKBH5
The Km and kcat values of FTO and ALKBH5 were determined by keeping a constant enzyme concentration of 0.5 μM and varying the substrate concentrations (1, 2, 3, 5 and 10 μM), according to reported methods30,56. The percentage demethylation at different substrate concentrations was plotted as a function of time (Fig. 2 and Supplementary Fig. S1). The initial velocity (V0) for each substrate concentration was determined from the slope of the curve at the beginning of a reaction. The Michaelis–Menten curve was fitted using non-linear regression and the kinetic constants (Vmax, Km) of the substrate was estimated using GraphPad Prism. All reactions were performed at 37 °C in triplicate and were adjusted to ensure that less than 20% of the substrate was consumed.
UV-based thermal denaturation studies
The melting of each oligonucleotides was performed on a Cary 3000 UV-Visible Spectrophotometer (Varian) at a total strand concentration of 5 μM (unless stated otherwise) in 10 mM sodium phosphate buffer, pH 7.4 and 150 mM NaCl. Absorbance versus temperature profiles were recorded at 260 nm. The samples were first denatured by heating to 85 °C at 10 °C/min, followed by slow cooling to 20 °C at 0.4 °C/min to ensure a complete annealing of the strands. The melting transitions were then monitored by heating to 85 °C at 0.4 °C/min. To increase the accuracy of measurements, the sixth position was used to record the temperature data points by placing a temperature probe directly in the cuvette. Up to six melting transitions were measured for each oligonucleotide and the average Tm values were calculated using Varian Cary Software. For sample preparation, lyophilised oligos were reconstituted in the buffer and their concentrations were determined by UV absorbance at 260 nm (A260) using a NanoDrop ND-1000 UV-Visible Spectrophotometer. Extinction coefficients were calculated using the nearest neighbour approximation. The extinction coefficient of oligos containing m6A was assumed to be the same as those containing adenosine.
Analysis of thermodynamic data for bimolecular duplex structures
Each oligonucleotide was measured at six different strand concentrations from 1–100 μM in buffer containing 10 mM sodium phosphate buffer, pH 7.4 and 150 mM NaCl. They were subjected to multiple melting-annealing cycles while monitoring UV absorbance at 260 nm as described above. The melting transitions for duplex structures were assumed to proceed in a two-state manner and to obey the van’t Hoff’s equation (1) below.

A plot of 1/Tm versus ln(total strand concentration) gives a straight line, where the slope is R/ΔH° and the y-intercept is ΔS°/ΔH°. Data were fitted using linear least-squares minimisation using GraphPad Prism. The free Gibbs energy (ΔG°) were calculated at 37 °C (310.15 K) using the following equation (2).

Analysis of thermodynamic data for monomolecular hairpin structures
The experimental absorbance versus temperature curves were first converted into a fraction of strands remaining hybridized (α) versus temperature curves, which were then fitted to a two‐state transition model using Varian Cary Software.
Circular dichroism (CD) spectroscopy
CD spectra were obtained with a JASCO J810 spectro-polarimeter. The measurements were carried out with 5 μM oligonucleotides (unless stated otherwise) in a 10 mM sodium phosphate buffer, pH 7.4 and 150 mM NaCl. The oligos solutions were first heated to 90 °C for 5 min and re-annealed by slow cooling to 4 °C at a rate of 1 °C/min. CD spectra were then recorded in quartz cuvettes (path length 10 mm, 400 μL) from 200 nm to 350 nm using a 10 nm/min scan speed, a spectral band width of 1 nm and a time constant of 4 s. All the spectra were subtracted with the buffer blank and smoothed using the Savitsky-Golay algorithm (polynomial order 10).
Non-denaturing polyacrylamide gel electrophoresis (PAGE) analysis
Annealed oligonucleotides were loaded to 20% native polyacrylamide gel and electrophoresis was performed at 4 °C in Tris/Borate/EDTA (TBE) running buffer (90 mM Tris, pH 8.3, 90 mM boric acid and 5 mM EDTA). The gel were stained with SYBR® Gold Nucleic Acid Gel Stain and visualized by Gel Dock XR + (Bio-Rad) and Image Lab 4.0 software (Bio-Rad). The fraction of monomolecular hairpin structures was evaluated based on the assumption that the efficiency of the staining of the base pairs in a hairpin was similar to that in a bimolecular duplex.
Microscale Thermophoresis (MST) Measurement
MST experiments53,57 were performed on a Monolith NT Label Free system (NanoTemper Technologies) at 25 °C using 40% MST power and 60% LED power for FTO and 40% LED power for ALKBH5. Laser on and off times were set at 30 s and 5 s, respectively. Zero background standard treated capillaries (NanoTemper Technologies) were used. All experiments were performed in triplicates. Oligos for MST measurement were purchased from DharmaconTM. For determination of binding affinity with FTO, twelve different concentrations of RNA ranging from 1 mM to 490 nM were used. The annealed RNAs were incubated with a mixture of FTO/NiSO4 complex (100 nM/1 mM) and NOG (500 μM) in 50 mM Tris buffer (pH 7.5; containing 150 mM NaCl and 0.05% Tween 20) at 25 °C for 20 minutes. Data derived from thermophoresis measurement and temperature-dependent change in fluorescence (T-Jump) were used to determine the binding affinities (KD) of the RNAs. Curve fitting was performed using the NT Analysis software provided. For experiments with ALKBH5, ALKBH5/MnCl2 complex (500 nM/1 mM) was used.
Electrophoretic mobility shift assay (EMSA)
RNA probes that were labelled at their 3′-ends with biotin were purchased from Keck Foundation Biotechnology Resource Laboratory. Prior to electrophoresis, 2 μL of annealed RNA probes (4 nM final concentration) were incubated with 2 μL of FTO or ALKBH5 or YTHDF2 (0.02, 0.5, 1, 2, 5 μM and other concentrations as indicated) at 4 °C for 30 min in a binding buffer (10 mM HEPES, pH 7.3, 5% glycerol, 20 mM KCl, 1 mM MgCl2, 1 mM DTT and 8 U RNasin® Ribonuclease Inhibitor (Promega)). 20 uL of the RNA-protein mixture was loaded to 7.5% native polyacrylamide gel and electrophoresis was performed at 4 °C for 90 min at 90 V using TBE running buffer. Electrophoresis for ALKBH5 and FTO were performed in buffer without EDTA. Visualisation was carried out using CL-XposureTM Film (Thermo Scientific), Gel Dock XR + (Bio-Rad) and Image Lab 4.0 software (Bio-Rad).
Additional Information
How to cite this article: Zou, S. et al. N6-Methyladenosine: a conformational marker that regulates the substrate specificity of human demethylases FTO and ALKBH5. Sci. Rep. 6, 25677; doi: 10.1038/srep25677 (2016).
References
Grosjean, H. Fine-tuning of RNA functions by modification and editing, Ch. xxiv, 442 (Springer, Berlin, New York, 2005).
Grosjean, H. & Benne, R. Modification and editing of RNA, Ch. xiii, 596 (ASM Press, Washington, DC, 1998).
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways–2013 update. Nucleic Acids Res 41, D262–7 (2013).
Motorin, Y. & Helm, M. RNA nucleotide methylation. Wiley Interdiscip Rev RNA 2, 611–31 (2011).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m(6)A RNA methylation. Nat Rev Genet 15, 293–306 (2014).
Meyer, K. D. & Jaffrey, S. R. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol 15, 313–26 (2014).
He, C. Grand challenge commentary: RNA epigenetics? Nat Chem Biol 6, 863–5 (2010).
Rottman, F. M., Desrosiers, R. C. & Friderici, K. Nucleotide methylation patterns in eukaryotic mRNA. Prog Nucleic Acid Res Mol Biol 19, 21–38 (1976).
Wei, C. M. & Moss, B. Nucleotide sequences at the N6-methyladenosine sites of HeLa cell messenger ribonucleic acid. Biochemistry 16, 1672–6 (1977).
Canaani, D., Kahana, C., Lavi, S. & Groner, Y. Identification and mapping of N6-methyladenosine containing sequences in simian virus 40 RNA. Nucleic Acids Res 6, 2879–99 (1979).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–6 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–46 (2012).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 12, 767–72 (2015).
Lee, M., Kim, B. & Kim, V. N. Emerging roles of RNA modification: m(6)A and U-tail. Cell 158, 980–7 (2014).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–20 (2014).
Lin, S. & Gregory, R. I. Methyltransferases modulate RNA stability in embryonic stem cells. Nat Cell Biol 16, 129–31 (2014).
Wang, X. et al. N(6)-methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 161, 1388–99 (2015).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–72 (2007).
Fischer, J. et al. Inactivation of the Fto gene protects from obesity. Nature 458, 894–8 (2009).
Loos, R. J. & Yeo, G. S. The bigger picture of FTO: the first GWAS-identified obesity gene. Nat Rev Endocrinol 10, 51–61 (2014).
Church, C. et al. Overexpression of Fto leads to increased food intake and results in obesity. Nat Genet 42, 1086–92 (2010).
Claussnitzer, M. et al. FTO Obesity Variant Circuitry and Adipocyte Browning in Humans. N Engl J Med 373, 895–907 (2015).
Zhao, X. et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res 24, 1403–19 (2014).
Hess, M. E. et al. The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat Neurosci 16, 1042–8 (2013).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49, 18–29 (2013).
Liu, J. et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol 10, 93–5 (2014).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep 8, 284–96 (2014).
Jia, G. et al. Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Lett 582, 3313–9 (2008).
Aik, W. et al. Structure of human RNA N(6)-methyladenine demethylase ALKBH5 provides insights into its mechanisms of nucleic acid recognition and demethylation. Nucleic Acids Res 42, 4741–54 (2014).
Toh, J. D. W. et al. A strategy based on nucleotide specificity leads to a subfamily-selective and cell-active inhibitor of N6-methyladenosine demethylase FTO. Chem Sci. 6, 112–122 (2015).
Yang, T., Cheong, A., Mai, X., Zou, S. & Woon, E. C. Y. A methylation-switchable conformational probe for sensitive and selective detection of RNA demethylase activity. Chem Commun 52, 6181–4 (2016).
Han, Z. et al. Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464, 1205–9 (2010).
Xu, C. et al. Structures of human ALKBH5 demethylase reveal a unique binding mode for specific single-stranded N6-methyladenosine RNA demethylation. J Biol Chem 289, 17299–311 (2014).
Zhu, C. & Yi, C. Switching demethylation activities between AlkB family RNA/DNA demethylases through exchange of active-site residues. Angew Chem Int Ed Engl 53, 3659–62 (2014).
Frayling, T. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–94 (2007).
Scuteri, A. et al. Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet 3, e115 (2007).
Engel, J. D. & von Hippel, P. H. Effects of methylation on the stability of nucleic acid conformations: studies at the monomer level. Biochemistry 13, 4143–58 (1974).
Roost, C. et al. Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J Am Chem Soc 137, 2107–15 (2015).
Micura, R. et al. Methylation of the nucleobases in RNA oligonucleotides mediates duplex-hairpin conversion. Nucleic Acids Res 29, 3997–4005 (2001).
Engel, J. D. & von Hippel, P. H. Effects of methylation on the stability of nucleic acid conformations. Studies at the polymer level. J Biol Chem 253, 927–34 (1978).
Kierzek, E. & Kierzek, R. The thermodynamic stability of RNA duplexes and hairpins containing N6-alkyladenosines and 2-methylthio-N6-alkyladenosines. Nucleic Acids Res 31, 4472–80 (2003).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–90 (2015).
Liu, N. et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–4 (2015).
Liu, N. et al. Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA 19, 1848–56 (2013).
Bringmann, P. & Luhrmann, R. Antibodies specific for N6-methyladenosine react with intact snRNPs U2 and U4/U6. FEBS Lett 213, 309–15 (1987).
Iwanami, Y. & Brown, G. M. Methylated bases of transfer ribonucleic acid from HeLa and L cells. Arch Biochem Biophys 124, 472–82 (1968).
Saneyoshi, M., Harada, F. & Nishimura, S. Isolation and characterization of N6-methyladenosine from Escherichia coli valine transfer RNA. Biochim Biophys Acta 190, 264–73 (1969).
Harper, J. E., Miceli, S. M., Roberts, R. J. & Manley, J. L. Sequence specificity of the human mRNA N6-adenosine methylase in vitro. Nucleic Acids Res 18, 5735–41 (1990).
Csepany, T., Lin, A., Baldick, C. J. Jr. & Beemon, K. Sequence specificity of mRNA N6-adenosine methyltransferase. J Biol Chem 265, 20117–22 (1990).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–5 (2000).
Bird, A. Molecular biology. Methylation talk between histones and DNA. Science 294, 2113–5 (2001).
Xia, T. et al. Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs. Biochemistry 37, 14719–35 (1998).
Wienken, C. J., Baaske, P., Rothbauer, U., Braun, D. & Duhr, S. Protein-binding assays in biological liquids using microscale thermophoresis. Nat Commun 1, 100 (2010).
Wienken, C. J., Baaske, P., Duhr, S. & Braun, D. Thermophoretic melting curves quantify the conformation and stability of RNA and DNA. Nucleic Acids Res 39, e52 (2011).
Xu, C. et al. Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat Chem Biol 10, 927–9 (2014).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7, 885–7 (2011).
Jerabek-Willemsen, M., Wienken, C. J., Braun, D., Baaske, P. & Duhr, S. Molecular interaction studies using microscale thermophoresis. Assay Drug Dev Technol 9, 342–53 (2011).
Acknowledgements
We like to thank Protein Production Platform (Nanyang Technological University, Singapore) for providing the cDNA of human YTHDF2 and ALKBH5 and also the cloning and small scale expression of human FTO and ALKBH566–292. This work was funded by the Singapore Ministry of Health’s National Medical Research Council (NMRC/BNIG/2008/2013) and Singapore Ministry of Education (Start-Up Grant R148-000-168-133). It was also supported by the Singapore National Research Foundation NRF-RF2009-RF001-267 and Singapore Ministry of Education grant MOE2014-T2-1-083.
Author information
Authors and Affiliations
Contributions
E.C.Y.W. designed the project. S.Z. and J.D.W.T. contributed to the work equally. S.Z. and K.H.Q.W. performed the HPLC-based demethylation assay, enzyme kinetic studies, UV-based thermal denaturation studies and thermodynamic analysis. J.D.W.T. performed the protein expression and purification, CD analyses, native PAGE analyses, MST and EMSA experiments. E.C.Y.W wrote the manuscript. All authors contributed to research discussion and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zou, S., Toh, J., Wong, K. et al. N6-Methyladenosine: a conformational marker that regulates the substrate specificity of human demethylases FTO and ALKBH5. Sci Rep 6, 25677 (2016). https://doi.org/10.1038/srep25677
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep25677
This article is cited by
-
The role of the methyltransferase METTL3 in prostate cancer: a potential therapeutic target
BMC Cancer (2024)
-
Altered m6A RNA methylation governs denervation-induced muscle atrophy by regulating ubiquitin proteasome pathway
Journal of Translational Medicine (2023)
-
Deciphering glioma epitranscriptome: focus on RNA modifications
Oncogene (2023)
-
METTL14 promotes glomerular endothelial cell injury and diabetic nephropathy via m6A modification of α-klotho
Molecular Medicine (2021)
-
Evolutionary conservation of the DRACH signatures of potential N6-methyladenosine (m6A) sites among influenza A viruses
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.