Abstract
In Nature, protons (H+) can mediate metabolic process through enzymatic reactions. Examples include glucose oxidation with glucose dehydrogenase to regulate blood glucose level, alcohol dissolution into carboxylic acid through alcohol dehydrogenase and voltage-regulated H+ channels activating bioluminescence in firefly and jellyfish. Artificial devices that control H+ currents and H+ concentration (pH) are able to actively influence biochemical processes. Here, we demonstrate a biotransducer that monitors and actively regulates pH-responsive enzymatic reactions by monitoring and controlling the flow of H+ between PdHx contacts and solution. The present transducer records bistable pH modulation from an “enzymatic flip-flop” circuit that comprises glucose dehydrogenase and alcohol dehydrogenase. The transducer also controls bioluminescence from firefly luciferase by affecting solution pH.
Similar content being viewed by others
Introduction
In nature, H+ plays an important role in modulating enzymatic reactions during metabolic process to fulfill physiological functions1,2. Examples include glucose oxidation with glucose dehydrogenase to maintain the blood glucose level, alcohol digestion into carboxylic acid through alcohol dehydrogenase and voltage-regulated H+ channels activating bioluminescence in firefly and jellyfish3. There are several bioelectronics devices that use enzymatic reactions to regulate device output such as biofuel cells4, organic electrochemical transistors biosensors5 and enzyme logics mimicking electronic circuits6. In turn, bioelectronic devices may regulate electron mediated enzymatic reactions7,8. However, not all enzymatic reactions are electron mediated and control of enzymatic reactions with devices9,10,11,12 that modulate ionic currents is desirable.
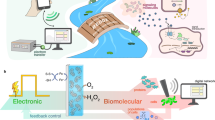
Examples of these devices include ion pumps based on conducting polymers that can deliver Ca2+ and the positive charged neurotransmitter gamma-aminobutyric to brain tissues for potential treatment of epilepsy13, metallic nanostraws14 and carbon nanotube porins that15,16 deliver selectively target molecules such as cations15, DNA14 and nicotine16. H+ also plays important roles in modulating physiological function17. We have developed bioprotonic devices that selectively control the flow of H+ including complementary field effect transistors18,19, synaptic memories20 and enzyme logic6. Recently, H+ transistors with the protein reflectin have been demonstrated21. These bioprotonic devices are based on PdHx as a transducer between H+ currents in the device or solution and e− current in the electronic circuit. Here, we develop an electrochemically-controlled PdHx biotransducer that not only records but also modulates the pH of a solution. We integrate this biotransducer with enzymatic reactions to implement H+ mediated monitoring and control of biochemical processes (Fig. 1). The biotransducer converts the input of an enzymatic flip-flop circuit – a bistable circuit with a high pH and a low pH state controlled by the enzymes glucose dehydrogenase and alcohol dehydrogenase- to a readable H+ current as the output (Fig. 1a) and it provides pH modulation as the input for the pH sensitive enzymatic reaction of luciferin and luciferase with bioluminescence as the output (Fig. 1b).
H+ mediated enzymatic reactions.
(a) Schematic of the enzymatic flip flop circuit as a bistable pH modulating system controlled by logically processed biochemical signals. The ‘set’ operation is the glucose input. The reaction between glucose and the enzyme glucose dehydrogenase (GDH) increases H+ concentration in solution and lowers the pH. Acetaldehyde (Act) reacts with the alcohol dehydrogenase (ALD) enzyme. The reaction consumes H+ in the reaction and increases the pH. The GDH and ALD AND enzyme logics regulate the solution pH, thus affect H+ transfer between solution and the contact. H+ combines with an electron at the Pd contact and forms PdH. The PdH biotransducer converts the biochemical signals into readable protonic current. (b) Schematic of firefly bioluminescence reaction integrated with pH modulating biotransducer. Luciferin is oxidized into oxy-luciferin with the presence of the luciferase enzyme. The reaction emits light. The solution pH affects the color of the light emitted. When the PdH pH modulator transfers H+ between the solution and the contact, it changes the solution pH. The bioluminescence light is the solution pH change readout.
Results
PdHx protodes measure the pH of a solution because the transfer of H+ between PdHx and the solution is affected by the difference in chemical potential of H+, or protochemical potential (μ), between the PdH (μPdH) and the solution (μpH) (Fig. S1)6,22. This difference is defined as22

where aH+ = activity of H+ in solution with pH = −log aH+, pH2 = hydrogen partial pressure in the Pd V = potential difference between Pd and solution
According to PdH ↔ Pd + H+ + e−, the H+ transfer results in a measurable electronic current. In our previous work, we have demonstrated recording of the pH readout of an enzymatic AND gate comprising glucose (Glc) and NAD+ with glucose dehydrogenase (GDH)15. In the presence of both NAD+ and Glc, GDH lowers the solution pH, which is recorded with the PdHx protodes. In the absence of either substrate, GDH is not active and the solution pH remains the same23. GDH modifies the pH only once because the enzymatic AND gate does not include a feedback loop to make the reaction reversible and return the solution to its original pH. Here, we integrate the enzyme alcohol dehydrogenase (ALD) with GDH to create an enzymatic flip-flop with a bistable pH modulation as the read out (Fig. 1a). A flip- flop is a circuit made by two AND gates. The output of the first AND gate is fed back as one of the inputs of the second AND gate and vice versa. In this fashion, a flip-flop has two stable states and can be used to store information. A set and a reset inputs switch the flip-flop between the two states. The enzymatic flip-flop consists of two types of enzymatic AND logic that use GDH and ALD as gates. GDH requires the presence of both NAD+ and Glc as inputs to function. GDH produces NADH and gluconic acid (GlcA), which lowers the solution pH from 6.0 to 4.3 (Fig. S2a). NADH, the output of the GDH, is one of the inputs for ALD with acetaldehyde (Act) as the other input. In the presence of both NADH and Act, ALD produces ethanol and NAD+. NAD+, in turn, is the input of the GDH logic gate. ALD consumes H+ during this enzymatic reaction and induces an increase in solution pH from 7 to 9.7 (Fig. S2b). The products (NADH or NAD+) of each AND gate (ALD or GDH) are connected to one of the inputs of the other gate. This connection results in a positive feedback that creates an enzymatic flip-flop24. The life-time of the flip-flop circuit depends on the life-time of the enzyme activity. While extended lifetime is not the goal of the present work, we measure at least three cycles of the bistable pH modulation and the devices are functional for at least one day when stored at low temperature. In our system, the ‘set’ operation is Glc injection, which causes a decrease in pH and an associated increase in the solution μpH6. The reset operation is Act injection, which causes an increase in pH and an associated decrease in the solution μpH. Solution pH is used as the readout.
To read out the state of the enzymatic flip-flop, we integrate it with the PdHx electrochemical biotransducer and monitor the transfer of H+ between the solution and the PdHx protode at a given applied voltage as function of the pH state (Fig. 2a) (Fig. S1). To this end, we use a standard three-electrode configuration with Pd (PdHx) as the working electrode (WE), Ag/AgCl as the reference electrode (RE) and platinum (Pt) as the counter electrode (CE). In this setup, a large enough cathodic voltage, Vc, applied to the WE (negative vs. Ag/AgCl) transfers an H+ from the solution to the Pd where H+ combines with an e− to form H. H adsorbs onto Pd to form PdHx. In general, H adsorption into the Pd nanofilm can expand its volume up to 10% after forming the PdH25, so the film thickness (50 nm Pd) will increase up to 55 nm of PdH. We monitor this H+ transfer by recording the cathodic current, Ic, resulting from the e− that flow into the Pd WE and participate in the reduction of H+ to H. A large enough reverse anodic voltage, Va, applied to the WE (positive vs. Ag/AgCl) causes an H+ to transfer from the PdHx to the solution leaving an e− behind, which leads to an anodic current Ia (electron flowing out of the Pd WE). When all of the PdHx is transformed into Pd, Ia goes to zero. The magnitude of Ia is proportional to the extent of PdHx formation in the cathodic phase because it is directly correlated to how many H+ have previously transferred from the solution into the PdHx26. We use the mangnitude of Ia as an indirect means of monitoring H+ transfer from solution into the PdHx because Ic also contains components related to charging of the solution.
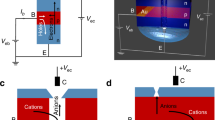
The biotransducer transduces the set-reset enzymatic flip-flop signals into current.
(a) Enzymatic flip-flop integrated with the PdH biotransducer. The electrochemical setup has a Pd working electrode, Ag/AgCl reference electrode and Pt counter electrode. Va and Vc is applied to Pd vs. Ag/AgCl. Ia and Ic is measured between Pd and Pt. The reaction between glucose and NAD+, catalyzed by the enzyme GDH, produces gluconic acid (GlcA) and NADH. Acetaldehyde (Act) reacts with the ALD enzyme and produces ethanol and NAD+. (b) Upper panel shows the pH change when adding Glc and Act into the solution. Lower panel shows Ic and Ia in response to pH change. Vc = 0 V and Va = −0.95 V is applied during the ‘set’ and ‘reset’ process respectively. Ic is recorded during Vc = 0 V and Ia is recorded during Va = −0.95 V. Ia is almost 0 when solution is above pH 6. At Va = −0.95 V, few H+ can be transferred into Pd contact. pH 5.5 induced by the addition of Glc into the solution causes Ia = 0.16 mA. The return of pH to above 6 with the addition of Act results in no Ia. (c) Truth table of the set-reset enzymatic flip-flop circuit.
In the initial state of the flip-flop, the solution pH = 7.5 and Vc = −0.95 V. According to the process map (Fig. S1) H+ do not transfer from the solution to the Pd contact, Ic = 0 A and setting Va = 0 V results in Ia =0 A because PdHx did not form when Vc was applied (Fig. 2b). We keep Vc = −0.95 V during the “set” operation when we add Glc (Fig. 2b). Adding 3 mM Glc to the solution causes the pH to drop from 7.5 to 5.5. Now, with Vc = −0.95 V, H+ transfer from the solution to the Pd contact and form PdHx. For PdHx, μPdH is higher than μpH when the pH = 5.5. When we switch to Va = 0 V, H+ transfer back to the solution. The Ia resulting from this transfer is as high as 0.16 mA. The ‘reset’ operation of adding Act in the presence of NADH engages the ALD enzymatic reaction, which consumes H+ and brings the pH back close to neutral with a value of pH = 6.5. At pH = 6.5, H+ do not transfer into the Pd when Vc = −0.95 V and PdHx does not form. As a result, when Va is set to 0 V, Ia = 0 mA.
Using these inputs and readouts, we produce a truth table for the enzymatic flip-flop (Fig. 2c). The output, or memory readout, switches between two states defined digitally as 0 (neutral pH) and 1 (low pH). These two states correspond to either Ia = 0 mA or Ia >>0 mA. The enzymatic flip-flop is activated by ‘set’ and ‘reset’ input signals applied at two levels digitally defined as 0 and 1. The presence of the ‘set’ Glc or the ‘reset’ Act is digital 1 and their absence is digital 0. Application of ‘set’ = 0 and ‘reset’ = 0 preserve the readout state regardless of its original value (0 or 1). Application of ‘set’ = 1 results in the readout state 1, while application of ‘reset’ = 1 yields the readout state 0, regardless of the original readout state (0 or 1). As in a digital flip-flop, the simultaneous application of the signals ‘set’ = 1 and ‘reset’ = 1 sends the system in opposite directions, results in an unstable readout and it is not permitted. The enzymatic flip-flop demonstrates how the Pd/PdHx biotransducer transduces H+ signals in the form of pH change from the enzymatic readouts into measurable electronic currents.
In addition to the protonic readout from an enzyme logic signal, we use this PdHx biotransducer to modulate pH in solution (Fig. 3a). When Vc is applied to the WE and H+ transfer from solution into the Pd contact, the net concentration of H+ in solution decreases and the pH increases (Fig. 3a). At the same time, when Va is applied to the WE and H+ transfer from the PdHx contact into the solution, the net concentration of H+ in the solution increases and the pH decreases (Fig. 3a). Since the voltage applied to the WE, Vc or Va, controls whether H+ flow from the solution into the PdHx contacts and vice versa, the net result is electronic pH control. In order to achieve this electronic pH control, we keep the conventional three-electrode system used in enzymatic flip-flop setup. This setup, however, does not produce the desired results because of the high catalytic property of the Pt CE. It is likely that redox reactions (4H+ + O2 + 4e− → 2H2O; 2H+ + 2e− → H2; 2H2O + 4e− → H2 + 2OH−) on the CE consume the H+ injected from the WE with no change in solution pH. To overcome this issue, we use a counter electrode that is inert and with high capacitance, similar to the electrodes used in electrochemical double layer capacitors27. We use a high capacitive carbon fabric decorated with carbon nanotube (CF/CNT), with a capacitance per unit area Ci = 6.5 mF/cm2 28. We first attempt to increase the pH of a Na2SO4 solution with initial pH = 6.8 (Fig. 3b). With Vc = −0.9 V on the WE, H+ transfer from the solution into the WE and form PdHx. The H+ concentration in the solution decreases and the pH increases as indicated by the pH paper changing color from yellow (neutral pH) to dark blue (basic pH) (Fig. 3b). The Pd WE also changes color from metallic silver into a darker gray as expected from the transformation of Pd into PdHx, as we have previously observed20. As a control, we confirm that with an Au WE the pH change is much smaller than when using the PdHx WE (Fig. S3). This is because the Au WE is not able to directly inject or sink H+ at the solution interface. The small pH change that we still observe with the Au WE may be due to H+ or OH− accumulating at the Au charged surface or other surface catalytic reactions29. However, the pH change with PdH protode is much higher than that with Au WE because the Au does not promote H+ transfer between the WE and the solution. When Va = 0.9 V is applied to the PdHx WE, PdHx injects H+ back into the solution, which returns to its original pH (Fig. S4). This pH modulation is reversible and solution pH cycles bewtween pH = 6.8 to pH = 9.2 several times (Fig. S5). While the inert CF/CNT enables pH modulation of solution by minimizing side reactions at the CE, likely some of these reactions remain and optimization of the system is required to increase the range of pH modulation.
pH regulation in solution.
(a) Schematics of pH modulator with CNT modified CF counter electrode. In (a), with Pd contact there is no Ia at Va = 0.9 V. At Vc = −0.9 V, H+ transfers from solution into the Pd and recombines with e− to form H. H adsorbs onto Pd and forms PdH. Decrease of H+ concentration in solution causes pH increase in solution. (b) Pictures of pH modulator with pH paper in solution. Yellow color of the paper indicates pH = 7.0 in solution, which is the initial pH of solution. Dark blue color indicates pH = 10.0 in solution. pH increase is due to H+ transfer from solution to Pd contact.
We demonstrate pH control in solution with an optical output by modulating the output bioluminescent enzymatic reactions (Figs 1b and 4)30. We choose to modulate the yellow-green glow that in nature is emitted by the firefly. This glow is a result of the oxidation reaction of firefly luciferin, catalyzed by the enzyme luciferase in the presence of ATP as the energy source (Fig. 1b). This enzymatic reaction is pH sensitive30. To this end, we utilize a Japanese toy (Photolight), which contains one packet of firefly luciferase and the other packet of its substrate luciferin, ATP powder and Mg2+, which is consumed during the reaction. We test the reaction at different pH and obtain the highest output at pH = 8 (Fig. 4a). The color also changes as a function of pH from yellow to bright green and ultimately to red when the solution pH changes from basic to acidic (Fig. 4a). This color change is consistent with prior reports31. We place the solution at pH = 6.7 containing the Photolight components in a PDMS chamber that includes the electrode setup used previously (Fig. 4b). When Vc = −0.9 V at the WE, H+ transfer from the solution to the Pd/PdHx WE and result in an increase of solution pH. When pH = 8 is reached in the solution, the bioluminescent output increases with a bright yellow color (Fig. 4c). When Va = 0 V at the WE, H+ transfer back into the solution, which returns to its original pH turning the bioluminescence off. This pH controlled bioluminescence can be cycled between the low pH off state and the high pH on state. We have not thoroughly mapped performance and lifetime for this process. In preliminary measurements we were able to turn the bioluminescence ON and OFF for at least four cycles. (Fig. S6).
Bioluminescence incorporated with pH modulating biotransducer.
(a) Schematic of the pH modulator integrated with bioluminescence enzymatic reaction. Change of pH lights up the solution due to the optimal pH of the enzyme reaction is reached. (b) The firefly bioluminescence at different solution pH. At acidic pH 6, solution shows red color. At neutral to basic pH, solution is yellow. The optimal pH of firefly luciferin luciferase reaction is pH 8, thus strongest light emission. (c) Bioluminescence turned on and off by the pH modulator. The initial pH of solution is 5.8. When applied Vc = −0.9 V, pH increases to reach the optimal pH for the enzymatic reaction. Bioluminescence turns on. When reversed to Va = 0.9 V, solution pH returns to initial pH and bioluminescence is off. Bioluminescence can be turned on and off for several cycles.
To decrease solution pH in addition to increase solution pH, we develop a pH modulator circuit with two Pd/PdHx WE and two separate chambers (Fig. 5a). The two chambers and Pd/PdHx WE act as H+ reservoirs for each other. The two WE are connected by a strip of proton conducting polymer (Nafion), which transfers H+ between the two Pd/PdHx contacts when a transfer voltage (VT) is applied20. Again, to monitor pH change in the two chambers we use firefly bioluminescence (Fig. 5b). We start with the chamber of the right at pH 6.5 and the chamber on the left at pH 7.5 as indicated by a reddish (right) and bright yellow (left) bioluminescent output. We start with the chamber on the right slightly acidic to ensure that there are enough H+ to transfer into the Pd WE. First, we apply Vc = −0.9 V to the Pd WE on the right and load it with H+ from the solution. Solution pH in the right chamber increases as indicated by the bioluminescent output color change from red to yellow. Second, we apply a transfer voltage Vt = 1.2 V between the two Pd WE to transfer the H+ from the right WE to the left WE along the Nafion H+ conducting bridge. At this stage, Vc = 0 V and it is likely that some of the H+ from the PdH move back to the solution with higher pH and lower protochemical potential6. This transfer results in the pH in the right chamber returning to slightly acidic. Third, we apply Va = 0.6 V to the WE on the right and transfer the H+ from the WE to the solution in the left chamber. The pH in the left chamber decreases as indicated by a change in color from bright yellow to red. If we apply Va = 0.6 V to the left chamber without previously transferring H+ from the right chamber and right WE, we do not observe any noticeable pH change.
Discussion and Conclusions
We demonstrate a Pd/PdHx based protonic biotransducer that records and modulates pH dependent enzymatic reactions. This recording and modulation is based on the pH dependence of the transfer of H+ between solution and Pd/PdHx. By coupling this biotransducer with two enzymatic AND gates, we demonstrate an enzymatic flip-flop that sets a high and a low pH state. Such integrated logic system may allow for future implantable biomedical devices controlled by physiological conditions. Further, we provide an optical output for pH modulation by using firefly bioluminescence with pH dependent color and intensity. While we modulate the pH of the entire solution, it is likely that the pH at the surface of the PdHx contacts is different due to expected electrostatic charging effects. We are able to modulate the solution pH between 6.8 and 9.6 with an increase in efficiency required to access a broader pH spectrum. This broader pH spectrum is required for controlling biological function in solutions with high buffering capacity such as physiological conditions26.
Materials and Methods
Devices are fabricated on microscope glass slide (2.5 cm × 4.5 cm VWR). 50 nm Pd with a 15 nm Cr adhesion layer is deposited via e-beam evaporation (Balzers PLS 500). PDMS wells (made from 5 mL PDMS solution) are used as solution containers and attached to the Pd substrates with PDMS. For the pH modulator circuit, 3.5 wt% agarose gel solution is used to seal between the PDMS wells and the substrates. Fully hydrated agarose gel is placed above the Nafion channel to hydrate the material. After 15 min we assume that the Nafion is fully hydrated and we begin the measurement. 200 μL Nafion 117 solution (5% concentration) from Sigma Aldrich is drop-cast on top of the patterned Pd substrate and the solution is dried in a fume hood. Glucose dehydrogenase (EC 1.1.47) is donated from TOYOBO enzymes. Alcohol dehydrogenase (EC 1.1.1.1), glucose, acetaldehyde, nicotinamide adenine dinucleotide, Na2SO4, MgSO4, universal pH indicator are purchased from Sigma Aldrich. Luciferin and luciferase are from the Photolight (Japan). Carbon fiber (TCC-3250) is donated from Toho Tenax Co. Carboxylic groups modified carbon nanotubes (20–30 nm) are purchased from Cheap Tubes.com. The CNT modified CF electrode follows a previously reported method28. Electrochemical measurements are performed with a potentiostat (BAS, model2325).
Additional Information
How to cite this article: Deng, Y. et al. Proton mediated control of biochemical reactions with bioelectronic pH modulation. Sci. Rep. 6, 24080; doi: 10.1038/srep24080 (2016).
References
Du, J. Y. et al. Protons are a neurotransmitter that regulates synaptic plasticity in the lateral amygdala. Proc. Natl. Acad. Sci. USA 111, 8961–8966 (2014).
Kawasaki, H. et al. Proton Acts as a Neurotransmitter for Nicotine-Induced Adrenergic and Calcitonin Gene-Related Peptide-Containing Nerve-Mediated Vasodilation in the Rat Mesenteric Artery. J Pharmacol Exp Ther 330, 745–755 (2009).
DeCoursey, T. E. Voltage-gated proton channels: what’s next? J. Physiol. 586, 5305–5324 (2008).
Miyake, T. et al. Enzymatic biofuel cells designed for direct power generation from biofluids in living organisms. Energy. Environ. Sci. 4, 5008–5012 (2011).
Rivnay, J. et al. High-performance transistors for bioelectronics through tuning of channel thickness. Sci. Adv. 1, e1400251 (2015).
Miyake, T., Josberger, E. E., Keene, S., Deng, Y. & Rolandi, M. An enzyme logic bioprotonic transducer. APL Mat. 3, 014906 (2015).
Heller, A. & Feldman, B. Electrochemistry in Diabetes Management. Acc. Chem. Res. 43, 963–973 (2010).
Yoshino, S., Miyake, T., Yamada, T., Hata, K. & Nishizawa, M. Molecularly Ordered Bioelectrocatalytic Composite Inside a Film of Aligned Carbon Nanotubes. Adv. Energy. Mater 3, 60–64 (2013).
Rivnay, J., Owens, R. M. & Malliaras, G. G. The Rise of Organic Bioelectronics. Chem. Mat. 26, 679–685 (2014).
Larsson, K. C., Kjall, P. & Richter-Dahlfors, A. Organic bioelectronics for electronic-to-chemical translation in modulation of neuronal signaling and machine-to-brain interfacing. Biochim. Biophys. Acta. 1830, 4334–4344 (2013).
Noy, A. Bionanoelectronics. Adv. Mater. 23, 807–820 (2011).
Tarabella, G. et al. New opportunities for organic electronics and bioelectronics: ions in action. Chem. Sci. 4, 1395–1409 (2013).
Berggren, M. et al. Electronic control of Ca(2+) signalling in neuronal cells using an organic electronic ion pump. Nat. Mater. 6, 673–679 (2007).
VanDersarl, J. J., Xu, A. M. & Melosh, N. A. Nanostraws for direct fluidic intracellular access. Nano. Lett. 12, 3881–3886 (2012).
Geng, J. et al. Stochastic transport through carbon nanotubes in lipid bilayers and live cell membranes. Nature 514, 612–615 (2014).
Wu, J. et al. Programmable transdermal drug delivery of nicotine using carbon nanotube membranes. Proc. Natl. Acad. Sci. USA 107, 11698–11702 (2010).
DeCoursey, T. E. Voltage-Gated Proton Channels: Molecular Biology, Physiology and Pathophysiology of the H-V Family. Physiol. Rev. 93, 599–652 (2013).
Zhong, C. et al. A polysaccharide bioprotonic field-effect transistor. Nat Commun 2, 476 (2011).
Deng, Y. et al. H+-type and OH− -type biological protonic semiconductors and complementary devices. Sci. Rep. 3, 2481 (2013).
Josberger, E. E., Deng, Y. X., Sun, W., Kautz, R. & Rolandi, M. Two-Terminal Protonic Devices with Synaptic-Like Short-Term Depression and Device Memory. Adv. Mater. 26, 4986–4990 (2014).
Ordinario, D. D. et al. Bulk protonic conductivity in a cephalopod structural protein. Nat. chem. 6, 597–603 (2014).
Flanagan, T. B. & Lewis, F. A. Electrode Potentials of the Palladium + Hydrogen System. Trans. Faraday. Soc. 55, 1409–1420 (1959).
Miyake, T. et al. Biofuel cell anode: NAD(+)/glucose dehydrogenase-coimmobilized ketjenblack electrode. Chem. Phys. Lett. 480, 123–126 (2009).
Zhou, M., Wang, F. A. & Dong, S. J. Boolean logic gates based on oxygen-controlled biofuel cell in “one pot”. Electrochim. Acta. 56, 4112–4118 (2011).
Mizumoto, M., Ohgai, T. & Kagawa, A. Bending behavior of Cu-plated Pd-Ni alloys ribbon driven by hydrogenation. J. Alloys. Compd. 482, 416–419 (2009).
Imokawa, T., Williams, K. J. & Denuault, G. Fabrication and characterization of nanostructured Pd hydride pH microelectrodes. Anal. Chem. 78, 265–271 (2006).
Simon, P. & Gogotsi, Y. Materials for electrochemical capacitors. Nat. Mater. 7, 845–854, doi: 10.1038/nmat2297 (2008).
Miyake, T., Haneda, K., Yoshino, S. & Nishizawa, M. Flexible, layered biofuel cells. Biosens. bioelectron. 40, 45–49 (2013).
Herrera Gallego, J., C. E. C., Calandra, A. J. & Arvia, A. J. The electrochemistry of gold in acid aqueous solutions containing chloride ions. J. Electroanal. Chem. Interfacial Electrochem. 66, 23 (1975).
Shapiro, E., Lu, C. & Baneyx, F. A set of multicolored Photinus pyralis luciferase mutants for in vivo bioluminescence applications. Protein. Eng. Des. Sel. 18, 581–587 (2005).
Ando, Y. et al. Firefly bioluminescence quantum yield and colour change by pH-sensitive green emission. Nat. photon. 2, 44–47 (2008).
Acknowledgements
This work was supported by the National Science Foundation under CAREER Award DMR-1150630 (enzymatic flip-flop and bioluminescence) and U.S. Department of Energy (DOE), Office of Science, Basic Energy Sciences (BES), under Award No. <DE-SC0010441> (H+ transfer between Pd/PdHx and solution). T.M. acknowledges a fellowship from the Japan Society for the Promotion of Science. Part of this work was conducted at the Washington Nanofabrication Facility/Molecular Analysis Facility, a member of the NSF National Nanotechnology Infrastructure Network.
Author information
Authors and Affiliations
Contributions
Y.D. and T.M. designed the experiment. Y.D., S.K. and T.M. performed the experiment. E.J. fabricated the device. Y.D., S.K. and T.M. analyzed the data. Y.D., T.M. and M.R. wrote the manuscript. All the authors revised the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Deng, Y., Miyake, T., Keene, S. et al. Proton mediated control of biochemical reactions with bioelectronic pH modulation. Sci Rep 6, 24080 (2016). https://doi.org/10.1038/srep24080
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24080
This article is cited by
-
A non-enzymatic glucose sensor enabled by bioelectronic pH control
Scientific Reports (2019)
-
A protonic biotransducer controlling mitochondrial ATP synthesis
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.