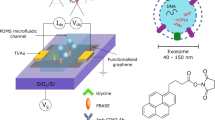
Abstract
A significant advantage of a graphene biosensor is that it inherently represents a continuum of independent and aligned sensor-units. We demonstrate a nanoscale version of a micro-physiometer – a device that measures cellular metabolic activity from the local acidification rate. Graphene functions as a matrix of independent pH sensors enabling subcellular detection of proton excretion. Raman spectroscopy shows that aqueous protons p-dope graphene – in agreement with established doping trajectories and that graphene displays two distinct pKa values (2.9 and 14.2), corresponding to dopants physi- and chemisorbing to graphene respectively. The graphene physiometer allows micron spatial resolution and can differentiate immunoglobulin (IgG)-producing human embryonic kidney (HEK) cells from non-IgG-producing control cells. Population-based analyses allow mapping of phenotypic diversity, variances in metabolic activity and cellular adhesion. Finally we show this platform can be extended to the detection of other analytes, e.g. dopamine. This work motivates the application of graphene as a unique biosensor for (sub)cellular interrogation.
Similar content being viewed by others
Introduction
Single layer graphene (SLG) is a planar sheet of sp2-bonded carbon atoms, organized into a hexagonal crystal lattice with exceptional optical, electronic, mechanical and thermal properties1,2,3. With its large contact area and high surface-to-volume ratio, graphene also has significant potential as a sensor, particularly for biological applications. Graphene oxide and chemically modified graphene (CMG) have shown the ability to detect the presence of single-strand DNA, aptamers, proteins, bacteria and viruses by quenching fluorescence of dyes attached to a part of the analyte by means of fluorescent resonant energy transfer (FRET)4. Fabrication of graphene and graphene-based field effect transistor (FET) devices and electrochemical sensors has allowed the detection of DNA, proteins, bacteria, mammalian cells, enzymes, small molecules (e.g. hydrogen peroxide, dopamine, glucose), biomacromolecules (e.g. hemoglobin) and different acids and bases4,5,6,7,8,9,10. For graphene-based FET devices, the detection mechanism is based on a change in graphene charge carrier mobility, minimum conductivity and charge neutrality point, caused by charge donation or extraction by the analyte (i.e. doping)5,11. These types of devices typically have active graphene areas of ~100 µm2 and are implemented such that the sensor response is averaged over the entire graphene surface12. Emerging topics of interest in biological analysis include single cell interrogation13,14,15,16, subcellular mapping of biochemical signaling17,18 and understanding phenotypic diversity within a cell population19,20,21. For these applications the FET graphene biosensors are less practical. The response of a single cell would not significantly alter the conductivity of the entire device. Even if one were able to make a graphene transistor small enough to show sensitivity to the excretion products of a single cell, the difficulty lies in placing only one single cell onto this transistor. While this is not technically impossible, it requires plenty of micromanipulations, on top of the clean-room work to make the transistor itself. Therefore suitable and practical cell-based applications of graphene FETs involve the measuring properties of a population of cells rather than of single cells6.
Raman spectroscopy has shown to be a reliable, fast and non-destructive technique to measure the degree of doping in graphene22,23,24,25,26, but also of other nanomaterials such as metallic nanoparticles27,28. Graphene is very sensitive to chemical dopants and even small shifts in its Fermi level result in distinctive changes in its Raman spectrum23,29. We assert that a graphene lattice can be conceptualized as an array of independently addressable optical sensors, practically limited in size only by extrinsic factors (i.e. diffraction limit, near field resolution). This offers the potential for spatial and temporal monitoring of the doping state of graphene locally and the possibility of single-molecule detection, in contrast to bulk conductivity measurements in graphene FETs. Furthermore, because graphene can be synthesized by chemical vapor deposition (CVD) methods into large macroscopic areas30,31, the high-sensitivity graphene-based sensor can extend over large detection areas.
In this work, we exploit the chemical sensitivity of single layer graphene together with the ability to make high-resolution spatial measurements via Raman spectroscopy to construct a novel graphene sensor platform. We report the pH-response of graphene and identify its two pKa values, as well as detect the metabolic footprint of isolated, living cells adhered to the graphene surface, classifying a population based on metabolic activity. Finally we show the graphene Raman biosensor can detect various other analytes such as immunoglobulin (IgG) and dopamine.
Results and Discussion
pH response of graphene measured by Raman spectroscopy reveals two pKa values
Large area (~1 cm2) monolayer graphene was synthesized via chemical vapor deposition (CVD) and transferred onto Si substrates with a 300 nm SiO2 capping layer following protocols similar to those reported earlier30,32 (full description in Methods Section). Six different graphene samples were exposed to unbuffered, aqueous solutions of varying initial pH (with pH values 0, 2, 4.5, 7.4, 12 and 14) with simultaneous micro-Raman spectroscopic mapping, as shown in Figure 1(a). Figure 1(b) shows characteristic Raman spectra of pristine graphene in air (blue curve) and of graphene exposed to extremely alkaline (red curve) and acidic (blue curve) conditions. Pristine CVD graphene is characterized by a small D peak near 1300–1350 cm−1, a G peak near 1580 cm−1 and a 2D peak between 2600 and 2700 cm−1, depending on the laser excitation source25,33. These three primary peaks are associated with various phonon modes in graphene and are sensitive to the electronic and structural properties of graphene22,23,24,25,26.
(a) Schematic of experimental setup (not to scale). A 150 μl drop of unbuffered solution with a specific pH is deposited onto a monolayer of CVD graphene supported by a SiO2/Si wafer. A cover slip (not shown) is put on top to slow down evaporation of the solution. 121 spatially distinct Raman spectra of graphene were collected both before and after it was exposed to the solution. (b) Characteristic Raman spectra of bare graphene in air (blue), graphene exposed to 1 M NaOH (red) and graphene exposed to 1 M HCl (green), showing the 3 main graphene peaks (D, G, 2D). All spectra are normalized to G peak height; inset zooms in on the G peak region. Exposure of graphene to alkaline solution decreases its peak positions, whereas contact with an acidic solution increases its peak positions. (c) Transient response of the G peak position to NaOH. A 3 M NaOH solution is added to graphene previously exposed to water, thereby changing the pH from 4.5 to 14 (red). The G peak position is observed to shift rapidly. Addition of PBS to graphene already exposed to PBS (pH is unchanged) does not shift the G peak position (blue). An exposure time of 100 ms was used.
The spectra in Figure 1(b) all show a very small D peak (indicating largely defect-free graphene) and clear differences in the G peak position, 2D peak position and 2D/G intensity ratio, corresponding to differences in the degree of doping (i.e. excess charge carriers in graphene)22,23,25,33,34. To determine how quickly graphene responds to changes in pH, a single position in a graphene sample, exposed to water, was monitored as a 3 M NaOH solution was added, changing the pH from 4.5 to 14. The trace in Figure 1(c) shows the G peak position (GPOS) shifts almost instantaneously after NaOH addition. To verify the NaOH response was indeed a pH effect and not an artifact of adding the solution, more PBS was added to a graphene sample already exposed to PBS; as expected no shift in the GPOS was observed (Fig. 1c).
For each sample, 121 (11 × 11) spatially distinct Raman spectra at 2 μm pitch were collected. Figure 2 shows scatter plots of the 2D peak position (2DPOS) (Fig. 2(a)), the 2D/G intensity ratio (I2D/IG) (Fig. 2(b)) and the G peak full width at half maximum (GFWHM) (Fig. 2(c)) versus GPOS; where the peak parameters were determined by Lorentzian fitting. Dashed lines represent doping trajectories based on data in ref. 23, in which Das et al. monitored the graphene Raman signature of electrostatically gated graphene as a function of the gate voltage VG. In doing so, they were able to establish a quantitative relationship between the number of injected carriers in graphene (dopants/cm2) and its Raman G peak position. Applying a gate voltage higher (lower) than the Dirac voltage VD (at which the number of excess charge carriers is zero, i.e. no doping) causes the injection of additional electrons (holes) into the graphene lattice. GPOS increases with either kind of excess charge carrier, whereas 2DPOS changes differently depending on the type of charge carrier: it increases for p-doping (additional holes) and decreases for n-doping (additional electrons)22,23,29. In previous work our group has shown that carriers injected or withdrawn by different substrates produce Raman peak dispersions that adhere to these trajectories24,26. Other signs of increased doping are a decreased I2D/IG and a decreased GFWHM22,23,25,33,34.
Scatter plots of Raman 2D peak position vs. G peak position (a), 2D to G intensity ratio vs. G peak position (b) and G peak width vs. G peak position (c) for graphene exposed to air (black dots) and solutions of different pH.Dashed lines are a guide to the eye adapted from ref. 23. The trend lines in (a) are shifted downwards to account for the dependence of the 2D peak position on the excitation wavelength (a 532 nm laser was used in ref. 23, as opposed to a 633 nm laser in this work). (d) Average hole concentration in graphene as a function of the pH of the solution in contact with graphene (red crosses). The hole concentration was deduced based on the relationship between GPOS and graphene dopant concentration, first shown in ref. 23 (see Fig. 3(b) in ref. 23). Dashed line represents independent binding events, whereas solid line represents negative cooperativity (with Hill coefficient n = 0.28). In the acidic regime the pre-hole-doped graphene becomes increasingly p-doped with decreasing pH and it has a pKa value of 2.9. In the alkaline regime the OH− ions compensate for (part) of the pre-existing hole-doping in graphene, which reveals a second pKa value of graphene of 14.2. The two pKa values indicate two types of binding sites for H+ to graphene, each with a different strength: H+ physisorbing to negative puddles in graphene (e) and species forming covalent bonds with the graphene lattice (chemisorption). The diameter of the puddles in graphene on SiO2 is on the order of 20 nm, much smaller than the laser spot size (~1 μm diameter).
The scatter plots of the doping-dependent Raman parameters show that graphene on SiO2 exposed to air (black dots) is somewhat p-doped, conform with literature results showing charge-transfer curves characterized by a positive Dirac voltage VD22,34,35. The additional holes can come both from the oxygen in the air, as well as from charged impurities in the SiO2 substrate, evidenced by the fact that both suspended graphene36 and graphene in vacuum35 display values of GPOS as low as 1580 cm−1 and Dirac voltages closer to 0, indicative of virtually undoped graphene. Additionally, the data in Figure 2 indicate that when graphene is exposed to increasingly acidic solutions (deionized (DI) water, 10 mM HCl, 1 M HCl), it becomes increasingly doped, specifically hole-doped. Due to the absence of organic buffers and competing adsorbates this reversible p-doping can unambiguously be assigned to the adsorption of protonated hydrogen ions. Protons adsorbed to the graphene surface form a charge transfer complex with an electron in the lattice, thereby localizing this electron and p-doping graphene.
For alkaline solutions we expect the negatively charged hydroxide ions to increasingly n-dope the graphene. Experimentally we observe that with increasing pH, graphene becomes more and more undoped, as evidenced by the low GPOS and 2DPOS (Fig. 2(a)), high I2D/IG (Fig. 2(b)) and GFWHM (Fig. 2(c)). The n-doping effect of the hydroxide ions appears to offset the initial p-doping in graphene. At first glance, even 1 M NaOH does not appear strongly alkaline enough to create a net negative excess charge in graphene. Eventually, for even higher concentrations of NaOH, we do expect the graphene 2DPOS vs. GPOS scatter data to move up the doping trajectory again and then eventually move down the n-doping branch, characterized by a high GPOS but low 2DPOS. Similarly we expect that graphene that starts out more undoped (e.g. suspended graphene) exposed to a 1 M NaOH solution would present a net n-doping.
A discussion of the spread in the data in Figure 2(a–c) is included in Section I of the Supporting Information.
The average dopant concentration in graphene for each condition shown in Figure 2(a) is plotted as a function of pH in Figure 2(d). The data are also included in Table 1, which summarizes the doping effects different chemical environments have on graphene.
Of interest is the H+ equilibrium demonstrated by graphene, a fundamental material property, in the form of the acid dissociation constant (Ka) or constants. From the empirical data, two regimes can be distinguished: the acidic regime (pH <7) and the alkaline regime (pH>7), characterized by Ka,1 and Ka,2 respectively. In the former, graphene becomes more p-doped with increasing H+ concentration. Thus:

with

where θfree is the concentration of available (free) ‘sites’ on graphene, θP,H+ the fraction of ‘sites’ p-doped by H+ and θtot,P the total concentration of graphene sites available for p-doping by H+. The total graphene carbon atom density is 3.8 × 1015 cm−2. Of this, a portion (θinitial,P) is already hole-doped by the substrate and/or oxygen as discussed earlier. This implies θtot,P = 3.8 × 1015 cm−2 - θinitial,P.
Rearranging these equations gives the concentration of p-doped sites of graphene:

where θP = θP,H+ + θinitial,P.
The last step in eqn. (3) assumes that the bulk concentration of aqueous H+ does not decrease significantly as the graphene becomes p-doped. This is easily justified by comparing the highest amount of excess holes present on the ~1 cm2 graphene sample (~9 × 1012 at 1 M HCl, see Table 1) to the total amount of H+ present in solution: 150 μl of 1 M HCl contains  hydrogen ions, meaning only 0.00001% of these p-dope graphene. It is interesting to note that at this highest doping concentration, ~0.25% of all carbon atoms in the graphene lattice are p-doped.
hydrogen ions, meaning only 0.00001% of these p-dope graphene. It is interesting to note that at this highest doping concentration, ~0.25% of all carbon atoms in the graphene lattice are p-doped.
In the alkaline regime (pH >7) the data indicate the hydroxide ions (OH−) ‘neutralize’ the initially present hole-doping in graphene; thus:

with

where Kb is the ‘neutralization’ dissociation constant.
Solving for the hole concentration in the alkaline regime leads to:

As in the acidic regime we can assume that [OH−] ≈ 10−pOH with pOH = 14-pH. Moreover it should be noted that

Fitting the combined model (eqn. (3),(6) and (7)) to the data (dot-dash line in Fig. 2(d)) results in a value of pKa1 = 2.9, pKa2 = 14.2 and θinitial,P = 4.85 × 1012 cm−2. Note that the latter value is close to the experimentally observed hole concentration in pristine graphene on SiO2 exposed to air ( = 5.2 × 1012 cm−2, Table 1). The Hill coefficient n, originally developed to describe protein-ligand binding37,38, accounts for cooperativity and modifies eqns. (3) and (6) as follows:


A Hill coefficient <1 (negative cooperativity) implies that when one ‘ligand’ (in this case H+ or OH−) binds to graphene, the affinity for another binding event with that ‘ligand’ decreases. Similarly a Hill coefficient >1 (positive cooperativity) indicates binding affinity increases with the number of binding events. Finally a Hill coefficient of 1 signifies each binding event occurs independently (dot-dash line in Fig. 2(d)). The solid line in Fig. 2(d) yields n1 = n2 = 0.28, indicating negative cooperativity, with the other parameter values remaining the same. Based on Figure 2(d), we can say the maximum sensitivity equals −2.98 × 1012 dopants/cm2 per pH unit in the acid regime and −7.9 × 1011 dopants/cm2 per pH unit in the basic regime.
The fact that, in the pH range explored in these experiments, graphene displays two pKa's, implies it has two types of binding sites for H+, a weaker and a stronger one, represented by pKa1 and pKa2 respectively. Charged impurities in the graphene substrate (in this case SiO2) have been shown to cause spatial electron-density inhomogeneities (i.e. electron and hole puddles) in graphene itself39,40,41. One explanation is that the weaker type of binding is positive hydrogen ions physisorbing to the negative charge puddles in graphene. This is shown schematically in Figure 2(e), where blue (red) areas represent the positive (negative) charge puddles in graphene, with protons adsorbing the negative ones. The schematic is based on the surface potential of graphene measured via scanning tunneling microscopy/spectroscopy (STM/STS)41. The resulting electrostatic screening associated with ion adsorption anticipates negative cooperativity, as found above. In order to verify that the charge puddles in graphene are responsible for the attraction of charged species, we compared the pH sensitivity of graphene on SiO2 to that of graphene on a self-assembled monolayer (SAM) which can partially screen the charges of the underlying SiO224. As expected the graphene pH response is much reduced on the SAM (see Section II of the Supporting Information for details). The physisorption of these charged species is expected to be fully reversible, which we have experimentally verified by exposing graphene to multiple cycles of 10 mM NaOH-10 mM HCl (Figure S3 of Section III of the Supporting Information). The second pKa2 may be associated with covalent binding (chemisorption) of species to the graphene lattice. Though largely defect-free, CVD graphene has grain boundaries characterized by more strain, defects and dangling bonds, which provide an opportunity for charged species to interact with42,43,44. This type of covalent interaction would not introduce additional defects (and therefore would not increase the D/G ratio). Secondly highly reactive or radical species can introduce additional defects to the graphene lattice. For example, in the presence of oxygen at low pH, a hydroperoxide group (-OOH) could covalently bind to graphene, leaving the graphene p-doped. Although the presence of this structure in graphene has not yet been investigated to date, it is known to be present on and p-dope their one-dimensional equivalent, single walled carbon nanotubes (SWCNTs)45. It should be noted we only observe slight increases of the D/G intensity ratio for extreme alkaline (1 M NaOH) or acidic (1 M HCl) conditions, indicating the higher pKa is largely dominated by ions binding to existing dangling bonds.
Finally it is interesting to note that the graphene pH dependence, probed via Raman spectroscopy in this work, displays seemingly opposite trends compared to those reported using electrostatic gating of a graphene FET exposed to solutions with different pH values12,46,47. In FET experiments, a shift of the Dirac voltage VD towards more positive values with increasing pH has been interpreted as increased p-doping in several studies12,46,47. In contrast, our Raman results clearly indicate increased p-doping with decreasing pH. FET experiments for determining the pH response of graphene are complicated by several factors. First, contrary to our approach, a FET is operated with an applied bias voltage and thus does not investigate an equilibrium carrier population. Second, the application of a gate voltage results in a transverse electric field that can either attract or repel ionic charges away from the graphene surface, altering ionic adsorption and potentially giving ions enough energy to covalently bind to graphene. And third, to our knowledge, in all literature describing pH dependence using graphene FETs buffered pH solutions are used. These buffers (e.g. phthalate) contain groups that can themselves dope graphene.
Graphene as an Array of Addressable pH Sensors: a Single Cell Physiometer
One marker for cellular metabolism is the cellular acidification rate, forming the basis for established techniques such as the cytosensor micro-physiometer, but these typically require 104–106 cells48,49. Decreasing the sensor size to micron- and nano-meter dimensions has the potential to extend this analysis to single cells. Graphene can be considered a micro-array of sensor units, with the size of these units determined by the optical diffraction limit of the objective lens used in the Raman spectroscopy system. For example, when using a 50X objective with NA = 0.75, the diffraction limited spot size of the excitation laser on graphene is ~1.03 μm, implying a minimum sensor size of 0.84 μm2. A typical cell with a ~20–40 μm length scale would cover multiple sensors which can all be individually probed via Raman spectroscopy (Fig. 3(a)). We investigated if it was possible to spatially map the sub-cellular acidification rate of a single biological cell, placed on a graphene substrate.
(a) Schematic of graphene as a micro-array of information units, with each unit acting as a micro-sensor. Each unit can be probed via Raman spectroscopy and reveal information about the doping state of graphene. A cell placed on graphene is expected to leave a ‘footprint’ on the graphene which can be detected via Raman spectroscopy. Graphene lattice not drawn to scale.(b) optical micrograph of IgG-producing HEK-293F cell well-adhered to graphene. (c) Characteristic individual Raman spectra of graphene exposed to air (blue), exposed to growth medium (red), under the cell (green) showing the 3 main graphene peaks (D, G, 2D). All spectra are normalized with respect to G peak height. Inset zooms in on the G peak region. Stars denote additional peaks observed in graphene covered by the cell. (d) Spatial Raman footprint of the cell shown optically in (b), obtained by probing the graphene G peak position: higher values are found under neath the cell, indicative of more doping. Note this smoothed Raman map was obtained by masking the fast Fourier transform (FFT) image of the raw data and applying an inverse FFT to shift it back into the spatial domain. Raw data and details about the FFT are included in Section V of the Supporting Information.
Transducing human embryonic kidney (HEK-293F) cells with lentiviral vectors created a stable cell line expressing a murine IgG2a antibody (TA99). After passaging, the cells were resuspended and diluted in L-15 medium. An aliquot of 150 μl of this solution was deposited onto a monolayer of graphene supported by a SiO2/Si wafer. A coverslip was put on to prevent evaporation and after 3 hours of incubation (allowing the cells to adhere to graphene) the sample was examined with optical microscopy and Raman spectroscopy. More details on the experimental procedures can be found in the Methods section and Supporting Information Section IV.
Figure 3(b) shows an optical image of a typical well-adhered IgG-producing cell, after 3 hrs of incubation on the graphene lattice. Figure 3(c) shows representative Raman spectra of bare graphene exposed to air (blue) and of graphene covered in cellular growth medium (red) and underneath the cell (green). Having verified that the graphene underneath the cell exhibits a different Raman signal from that exposed to just medium, we recorded the spatial Raman footprint of an entire cell. To avoid evaporation of the growth medium, a rapid, spatial map of the GPOS was constructed for each cell, because this Raman feature displays the most pronounced changes to pH and other adsorbates (inset Fig. 3(c)). We find that other peak parameters, such as the 2DPOS and the I2D/IG ratio also show doping-dependent changes, but require an extended spectral window (1500 cm−1 –2800 cm−1 for both G and 2D peak vs. 1500 cm−1–1675 cm−1 for just the G peak), increasing the scan time. Figure 3(d) shows the G peak position of the cell and surrounding medium shown in Figure 3(b). The shape of the cell is remarkably well preserved in the spatial plot of G peak position (Fig. 3(d)). We note that some level of spatial inhomogeneity in these maps is likely caused by a multitude of factors: the cell, the cell medium but also the graphene itself and the SiO2 substrate, where the latter two have documented inhomogeneity arising from local variations in graphene quality, doping and electron/hole puddles. Central to this present study, however, is the observation that underneath and immediately around the cell the graphene exhibits higher values of GPOS compared to further away from the cell, indicating the cell is selectively p-doping the graphene within its immediate footprint22,23,25,33,34. In light of our earlier findings, we assign these observations to the cellular efflux of acidic products whose protons p-dope the graphene.
Similar spatial footprints were collected using a higher-sensitivity Raman system (see Methods Section for details) which enables significantly lower scan times while maintaining spatial resolution. Additionally, Raman maps were generated for IgG-producing cells after 30 minutes of incubation, resulting in less adhesion to graphene, as well as for both well- and less- adhered control (non IgG-producing) cells. Optical images and Raman footprints are shown in Figure S5. In suspension, the HEK293 cells are round. When fully adhered to a surface, they are spread, flat and spindle-shaped. The cells that were only allowed to spend 30 minutes in the incubator prior to data collection are still round, indicating they are not fully adhered yet. The observed doping effect is more pronounced for the more strongly adhered cells as well as for IgG-producing cells compared to non-IgG producing control cells.
Graphene as a Cytometer Based on Cell-Induced Graphene Doping
An alternative way to utilize graphene as a pH sensor array is to reduce the spatial resolution but dramatically increase the throughput by measuring the average local acidification under each single cell in a population. Such an experiment can profile hundreds of cells and again compare the results to established doping trajectories to yield information about the ensemble. This approach also allows us to verify that the data observed for single cells extends to representative populations.
Specifically we collected Raman spectra underneath 100 individual cells for all 4 cases: well-adhered IgG-producing cells, less-adhered IgG-producing cells, well-adhered control cells and less-adhered control cells. For each sample size of 100 cells we collected one full Raman spectrum in the center of each cell. Figure 4(a) and (b) are optical micrographs displaying typical cell densities used in these experiments, for both well-adhered and less-adhered cells respectively. Figure 4(c–f) shows scatter plots of 2DPOS vs. GPOS, for graphene underneath cells (red squares) and underneath growth medium far away from the cells (black dots). The insets display histograms of the G peak positions of the graphene sampled underneath the cells (red) and of the graphene covered in growth medium (black). The growth medium was sampled in various locations on the graphene, but always at least 100 micron away from a cell to minimize the presence of products excreted by the cells. Scatter plots showing I2D/IG and GFWHM versus GPOS are included in the Supporting Information (Fig. S6 and S7). The data in Figures 4, S6 and S7 confirm all the trends we uncovered based on the single cell Raman footprints shown in Figures 3 and S5:
Optical micrograph of well-adhered cells (allowed to adhere to graphene for 3 hours in the incubator prior to data collection) in growth medium on graphene (a) and of less adhered cells (spent only 30 minutes in the incubator prior to data collection) in medium on graphene (b).Scatter plots of 2D peak position vs. G peak position of graphene covered in medium (black dots) and 100 distinct cells (red squares) for IgG-producing cells ((c),(d)) and non-IgG-producing control cells ((e),(f)). Left panels ((a),(c),(e)) represent well-adhered cells (3 hrs of incubation prior to data collection); right panels ((b),(d),(f)) represent less-adhered cells (30 min of incubation prior to data collection). Dashed lines represent trend lines adapted from ref. 23 The trend lines are shifted downwards to account for the dependence of the 2D peak position on the excitation wavelength (a 532 nm laser was used in ref. 23, as opposed to a 633 nm laser in this work). Scales and axes are identical for all panels. Insets in (c),(d),(e) and (f) are histograms of the G peak position of graphene in medium (black) and under the cells (red). pG-position <0.01 for datasets in panel (c-e), indicating the distributions are significantly different; pG-position = 0.09 for datasets in panel (f) indicating the distributions are not significantly different (see also Supporting Information Section VII for more details).
‐ Graphene exposed to cell medium is virtually undoped; introducing cells then upshifts the peak positions of graphene locally, indicative of p-doping.
‐ Increased cell adhesion causes stronger graphene doping (compare left panels (c) and (e) in Figure 4 to right panels (d) and (f)). The stronger contact between the cells and the graphene likely allows for more trapping of proton efflux at the graphene surface and hence creates a lower local pH.
‐ IgG-producing cells dope graphene more strongly than non-IgG producing control cells (compare top panels (c) and (d) to bottom panels (e) and (f) in Figure 4).
A Graphene Cell Physiometer as a Tool to Measure Cellular Metabolic Rate
The fact that IgG-producing cells dope graphene to a greater extent than the non-IgG producing cells is consistent with the former having a higher proton efflux rate. The IgG-producing cells have been genetically manipulated to produce an additional antibody in comparison to the control cells (see Supporting Information Section IV); this additional task requires more energy from the cells, increasing their metabolism and hence the excretion of acidic products48,49. The difference between the measured graphene hole concentration underneath the cell and far away from the cell can be shown to be linearly proportional with the cell's steady state proton production rate (and thus, its metabolic activity). This derivation is included in Supporting Information Section IX. Based on this relationship and the data in Figure 4 we can compare the metabolic activity of entire cell populations. For example, the metabolic rate of IgG-producing well-adhered cells is four times higher than that of well-adhered control cells.
However, it should be noted that the IgG producing cells are not only more metabolically active compared to the control cells. Due to the presence of a signal peptide at the N-terminus of the IgG protein, the latter is being secreted by the cells (about 1 pg per cell per day). In Section X of the Supporting Information the effect of IgG on the graphene Raman signal is examined. An IgG concentration of 1 mg/ml in the medium n-dopes graphene with 1012 electrons per cm2. This n-doping is expected due to the presence of n-doping moieties in the protein. This implies the IgG excretion slightly offsets the effect of the increased proton secretion of the IgG-producing cells on the graphene Raman signal; this implies and that the IgG-producing cells are at least four times as metabolically active as the adhered control cells.
Graphene Raman-based sensing platform is extendable to other analytes
The response to IgG is particularly noteworthy, since antibody structure is largely conserved50 and the extension of this technique to antigen detection is highly compelling. The result of this control experiment also opens the door to monitor the presence of a variety of other chemical analytes with graphene, based on the sensitivity of its Raman signal to excess charge carriers presented by such analytes. Specifically, future work will aim to explore the mechanism of dopamine uptake and release in networks of fully differentiated mammalian neural cells. A lot of research efforts focus on spatially and temporally detecting exocytosis of neurotransmitters and neuromodulators across the membrane of single cells51,52,53,54,55,56. In preliminary work we show we can easily detect the potassium-triggered release of dopamine by neural progrenitor PC12 cells (see Figure 5 and Supporting Information Section XI for more details).
Effect of stimulated neural progenitor PC12 cells on the graphene Raman signal.
(a) Optical micrograph of a cluster of PC12 cells on graphene (45 μm × 45 μm). (b) Rayleigh scattering (i.e. confocal reflectance) map of the area shown in (a), in the central focal plane of the cell cluster. (c) Rayleigh scattering map of the area shown in (a), in the graphene focal plane, showing the part of the cell cluster that is well-adhered to the graphene. (d-f) Spatial map of Raman G peak position (d), G peak FWHM (e) and D to G intensity ratio (f) of the graphene shown in (a), indicating a clear footprint of the potassium-triggered dopamine-release can be detected (see also S.I. Section XI); 500 ms exposure was used.
Outlook
There is a pressing need for tools capable of single cell analysis of metabolic activity. Cancer cells typically have higher metabolic activity than non-cancerous ones57,58, leading to a stronger acidification of their environment. The proposed graphene physiometer could gauge the metabolic activity of such cells, correlate this with tumor malignancy59 and ultimately derive options for personalized medical treatments. Single-cell or population-based assessments of drug toxicity using pH changes of the cells are also compelling60. Graphene has advantages for monitoring biofilm growth, which is strongly pH-dependent61. Ongoing and future work will explore these applications as well as focus on increasing the sensor sensitivity and collection speed. Additionally, it is possible to extend the sensing platform to the detection of other analytes, for example IgG (Figure S9, S10), but also neuromodulators such as dopamine (see Figure 5 and Section XI of the Supporting Information).
Methods
Graphene synthesis and transfer
Copper substrates (Alfa Aesar, 25 μm thick, 99.8%, annealed, uncoated) were pretreated in hydrogen chloride for 5 minutes, then rinsed with water, acetone and isopropyl alcohol and dried on a 90°C hotplate for 5 minutes. The Cu foil was annealed in vacuum under hydrogen flow (30 sccm, 1000°C, 20 min.), after which 3 sccm methane was added for the next 30 minutes during which graphene synthesis occurred via chemical vapor deposition. The graphene grew on both sides of the Cu foil. One side was coated with poly(methyl methacrylate) (950PMMA, A4, MicroChem), via spincoating at 3000 rpm for 1 minute. The graphene on the other side was removed via reactive ion etching (Plasmatherm RIE, 100 W, 7mtorr oxygen, 5 min.). The remaining Cu-graphene-PMMA structure was placed on top of a Cu-etchant bath (1 M CuCl2 and 6 M HCl in water). After the copper was completely etched away (~30 min) the graphene-PMMA structure was scooped out and placed into 3 consecutive baths of deionized (DI) water for 10 minutes each to remove all residual ions from the Cu-etchant. The graphene-PMMA structure was scooped up with a ~2.5 × 2.5 cm Si/SiO2 wafer (300 nm oxide) that had been cleaned by sonication in acetone and isopropyl alcohol for 5 minutes each. The water was allowed to evaporate overnight, after which the sample was rinsed with copious amounts of acetone and isopropyl alcohol to remove the PMMA. The final sample was dried with nitrogen gas. The relative intensities and the widths of the G and 2D Raman peaks as well as the color contrast in the optical microscope confirm the growth of monolayer graphene25,33.
Raman spectroscopy and mapping
The Raman spectroscopic data in Figures 1b, 2, 3, 4, S1–S7 and S9–S10 was collected with a top-down Horiba Jobin Yvon LabRAM HR800 system with a 633 nm excitation laser and an exposure time of 5 seconds. The 100X objective was used to probe bare graphene (exposed to air). In order to limit evaporation whenever graphene was exposed to a liquid, a glass microscope cover slip was used. The thickness of the glass cover slip (~0.25 mm) and of the liquid layer increased the total distance between the graphene layer and the objective, requiring the use of the 50X objective (with a working distance of 0.38 mm instead of 0.21 mm for the 100X objective). The diffraction limited spot size (in nm) can be calculated with the following equation:

with λ the wavelength of the laser excitation and NA the numerical aperture of the objective. When using the 100X objective (NA = 0.9) the minimum spot size is ~0.86 μm in diameter, whereas it is ~1.03 μm for the 50X objective (NA = 0.75). Therefore when constructing a Raman map, we ensured that the distance between two points of the map is at least 1 μm to avoid overlap. The laser power was set at 10 mW.
Since the calibration of the Horiba Raman spectrometer may shift over time or with temperature changes, it is crucial to calibrate the instrument prior to and after collecting each dataset. Cyclohexane was used as a reference.
Moreover, in order to decrease data collection time, we gained access to a home-built high signal/noise ratio confocal bottom-up Raman setup in MIT's Laser Biomedical Research Center (referred to in the text as a ‘higher-sensitivity’ setup). A 60X objective with a high numerical aperture and a high near-infrared (nIR) transmission, a sensitive detector with a high quantum-efficiency in the nIR and galvanometers to direct the Raman laser excitation on the sample improve both spatial resolution and collection speed. An exposure time of 100–500 ms seconds is sufficient to achieve strong Raman signals. For example, to collect a map of Raman spectra in a 30 μm × 45 μm area, with 2 μm step size, with 500 ms exposure, ~3 minutes are required, which is more than an order of magnitude faster than when using the Horiba Jobin Yvon LabRAM HR800 setup. The home-built instrument is described in more detail in references62,63 and was used to collect the data in Figure 1(c), Fig. 5, the bottom panel of Fig. S5 and Figs. S11–S13. The laser power used equals 8 mW.
A custom peak-fitting algorithm fits the D, G and 2D Raman peaks to Lorentzians, after which values of the peak position, full width half maximum (FWHM) and total intensity (total area under the Lorentzian) were extracted and compared.
References
Novoselov, K. S. et al. Two-dimensional gas of massless Dirac fermions in graphene. Nature 438, 197–200 (2005).
Geim, A. K. & Novoselov, K. S. The rise of graphene. Nature Mater. 6, 183–191 (2007).
Geim, A. K. Graphene: status and prospects. Science 324, 1530–1534 (2009).
Feng, L. & Liu, Z. Graphene in biomedicine: opportunities and challenges. Nanomedicine 6, 317–324 (2011).
Huang, Y., Dong, X., Liu, Y., Li, L.-J. & Chen, P. Graphene-based biosensors for detection of bacteria and their metabolic activities. J. Mater. Chem. 21, 12358–12362 (2011).
Nguyen, P. & Berry, V. Graphene interfaced with biological cells: opportunities and challenges. J. Phys. Chem. Lett. 3, 1024–1029 (2012).
Mohanty, N. & Berry, V. Graphene-based single-bacterium resolution biodevice and DNA transistor: interfacing graphene derivatives with nanoscale and microscale biocomponents. Nano Lett. 8, 4469–4476 (2008).
Cohen-Karni, T., Qing, Q., Li, Q., Fang, Y. & Lieber, C. M. Graphene and nanowire transistors for cellular interfaces and electrical recording. Nano Lett. 10, 1098–1102 (2010).
Hess, L. H. et al. Graphene transistor arrays for recording action potentials from electrogenic cells. Adv. Mater. 23, 5045–5049 (2011).
Khatayevich, D. et al. Selective detection of target proteins by peptide-enabled graphene biosensor. Small 10, 1505–1513 (2014).
Chen, J. H. et al. Charged-impurity scattering in graphene. Nature Phys. 4, 377–381 (2008).
Ohno, Y., Maehashi, K., Yamashiro, Y. & Matsumoto, K. Electrolyte-gated graphene field-effect transistors for detecting pH and protein adsorption. Nano Lett. 9, 3318–3322 (2009).
Wheeler, A. R. et al. Microfluidic device for single-cell analysis. Anal. Chem. 75, 3581–3586 (2003).
Cornelison, D. D. & Wold, B. J. Single-cell analysis of regulatory gene expression in quiescent and activated mouse skeletal muscle satellite cells. Dev. Biol. 191, 270–283 (1997).
Reiter, M. et al. Quantification noise in single cell experiments. Nucleic Acids Res. 39, e124 (2011).
Spiller, D. G., Wood, C. D., Rand, D. A. & White, M. R. Measurement of single-cell dynamics. Nature 465, 736–745 (2010).
Jin, H. et al. Detection of single-molecule H2O2 signalling from epidermal growth factor receptor using fluorescent single-walled carbon nanotubes. Nature Nanotechnol. 5, 302–309 (2010).
Jin, H., Heller, D. A., Kim, J. H. & Strano, M. S. Stochastic analysis of stepwise fluorescence quenching reactions on single-walled carbon nanotubes: single molecule sensors. Nano Lett. 8, 4299–4304 (2008).
Varadarajan, N. et al. A high-throughput single-cell analysis of human CD8+ T cell functions reveals discordance for cytokine secretion and cytolysis. J. Clin. Invest. 121, 4322–4331 (2011).
Gupta, P. B. et al. Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell 146, 633–644 (2011).
Kou, P. M. & Babensee, J. E. Macrophage and dendritic cell phenotypic diversity in the context of biomaterials. J. Biomed. Mater. Res. A 96, 239–260 (2011).
Casiraghi, C. Doping dependence of the Raman peaks intensity of graphene close to the Dirac point. Phys. Rev. B 80, 233407 (2009).
Das, A. et al. Monitoring dopants by Raman scattering in an electrochemically top-gated graphene transistor. Nature Nanotechnol. 3, 210–215 (2008).
Wang, Q. H. et al. Understanding and controlling the substrate effect on graphene electron-transfer chemistry via reactivity imprint lithography. Nature Chem. 4, 724–732 (2012).
Ferrari, A. C. Raman spectroscopy of graphene and graphite: Disorder, electron-phonon coupling, doping and nonadiabatic effects. Solid State Commun. 143, 47–57 (2007).
Paulus, G. L. C. et al. Charge transfer at junctions of a single layer of graphene and a metallic single walled carbon nanotube. Small 9, 1954–1963 (2012).
Bishnoi, S. W. et al. All-optical nanoscale pH meter. Nano Lett. 6, 1687–1692 (2006).
Talley, C. E., Jusinski, L., Hollars, C. W., Lane, S. M. & Huser, T. Intracellular pH sensors based on surface-enhanced Raman scattering. Anal. Chem. 76, 7064–7068 (2004).
Basko, D. M., Piscanec, S. & Ferrari, A. C. Electron-electron interactions and doping dependence of the two-phonon Raman intensity in graphene. Phys. Rev. B 80, 165413 (2009).
Li, X. et al. Large-area synthesis of high-quality and uniform graphene films on copper foils. Science 324, 1312–1314 (2009).
Bae, S. et al. Roll-to-roll production of 30-inch graphene films for transparent electrodes. Nature Nanotechnol. 5, 574–578 (2010).
Reina, A. et al. Transferring and identification of single- and few-layer graphene on arbitrary substrates. J. Phys. Chem. C 112, 17741–17744 (2008).
Ferrari, A. C. et al. Raman spectrum of graphene and graphene layers. Phys. Rev. Lett. 97, 187401 (2006).
Casiraghi, C., Pisana, S., Novoselov, K. S., Geim, A. K. & Ferrari, A. C. Raman fingerprint of charged impurities in graphene. Appl. Phys. Lett. 91, 233108 (2007).
Guo, B. et al. Controllable N-doping of graphene. Nano Lett. 10, 4975–4980 (2010).
Berciaud, S., Ryu, S., Brus, L. E. & Heinz, T. F. Probing the intrinsic properties of exfoliated graphene: Raman spectroscopy of free-standing monolayers. Nano Lett. 9, 346–352 (2009).
Hill, A. The possible effects of the aggregation of the molecules of haemoglobin on its dissociation curves. J. Physiol. 40, 4–7 (1910).
Heck, H. A. Statistical theory of cooperative binding to proteins. Hill equation and the binding potential. J. Am. Chem. Soc. 93, 23–29 (1971).
Zhang, Y., Brar, V. W., Girit, C., Zettl, A. & Crommie, M. F. Origin of spatial charge inhomogeneity in graphene. Nature Phys. 5, 722–726 (2009).
Martin, J. et al. Observation of electron-hole puddles in graphene using a scanning single-electron transistor. Nature Phys. 4, 144–148 (2008).
Xue, J. et al. Scanning tunnelling microscopy and spectroscopy of ultra-flat graphene on hexagonal boron nitride. Nature Mater. 10, 282–285 (2011).
Yazyev, O. V. & Louie, S. G. Topological defects in graphene: Dislocations and grain boundaries. Phys. Rev. B 81, 195420 (2010).
Huang, P. Y. et al. Grains and grain boundaries in single-layer graphene atomic patchwork quilts. Nature 469, 389–392 (2011).
Malola, S., Häkkinen, H. & Koskinen, P. Structural, chemical and dynamical trends in graphene grain boundaries. Phys. Rev. B 81, 165447 (2010).
Dukovic, G. et al. Reversible surface oxidation and efficient luminescence quenching in semiconductor single-wall carbon nanotubes. J. Am. Chem. Soc. 126, 15269–15276 (2004).
Ang, P. K., Chen, W., Wee, A. T. S. & Loh, K. P. Solution-gated epitaxial graphene as pH sensor. J. Am. Chem. Soc. 130, 14392–14393 (2008).
Lei, N., Li, P., Xue, W. & Xu, J. Simple graphene chemiresistors as pH sensors: fabrication and characterization. Meas. Sci. Technol. 22, 107002 (2011).
Hafner, F. Cytosensor® Microphysiometer: technology and recent applications. Biosensors and Bioelectronics 15, 149–158 (2000).
Owicki, J. C. & Wallace Parce, J. Biosensors based on the energy metabolism of living cells: the physical chemistry and cell biology of extracellular acidification. Biosens. Bioelectron. 7, 255–272 (1992).
Wang, W., Singh, S., Zeng, D. L., King, K. & Nema, S. Antibody structure, instability and formulation. J. Pharm. Sci. 96, 1–26 (2007).
Mellander, L., Cans, A.-S. & Ewing, A. G. Electrochemical probes for detection and analysis of exocytosis and vesicles. ChemPhysChem 11, 2756–2763 (2010).
Chen, T. K., Luo, G. & Ewing, A. G. Amperometric monitoring of stimulated catecholamine release from rat pheochromocytoma (PC12) cells at the zeptomole level. Anal. Chem. 66, 3031–3035 (1994).
Heien, M. L. et al. Real-time measurement of dopamine fluctuations after cocaine in the brain of behaving rats. P. Natl. Acad. Sci. USA 102, 10023–10028 (2005).
Phillips, P. E., Stuber, G. D., Heien, M. L., Wightman, R. M. & Carelli, R. M. Subsecond dopamine release promotes cocaine seeking. Nature 422, 614–618 (2003).
He, Q. et al. Centimeter-long and large-scale micropatterns of reduced graphene oxide films: fabrication and sensing applications. ACS Nano 4, 3201–3208 (2010).
Koike, T. & Takashima, A. Cell cycle-dependent modulation of biosynthesis and stimulus-evoked release of catecholamines in PC12 pheochromocytoma cells. J. Neurochem. 46, 1493–1500 (1986).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Urano, Y. et al. Selective molecular imaging of viable cancer cells with pH-activatable fluorescence probes. Nature Med. 15, 104–109 (2008).
Bickis, I. J. & Henderson, I. W. Biochemical studies of human tumors. I. Estimation of tumor malignancy from metabolic measurements in vitro. Cancer 19, 89–102 (1966).
Ekwall, B. Toxicity to HeLa cells of 205 drugs as determined by the metabolic inhibition test supplemented by microscopy. Toxicology 17, 273–295 (1980).
Dunne, W. M. Bacterial adhesion: seen any good biofilms lately? Clin. Microbiol. Rev. 15, 155–166 (2002).
Kang, J. W. et al. Combined confocal Raman and quantitative phase microscopy system for biomedical diagnosis. Biomed. Opt. Express 2, 2484–2492 (2011).
Kang, J. W., Nguyen, F. T., Lue, N., Dasari, R. R. & Heller, D. A. Measuring uptake dynamics of multiple identifiable carbon nanotube species via high-speed confocal raman imaging of live cells. Nano Lett. 12, 6170–6174 (2012).
Acknowledgements
This work was supported in part by the U. S. Army Research Laboratory and the U. S. Army Research Office through the Institute for Soldier Nanotechnologies, under contract number W911NF-13-D-0001. Fluorescence activated cell sorting was partially funded by Cancer Center Support (core) Grant P30-CA14051 from the NCI. The higher-sensitivity Raman work was supported by the NIH National Institute of Biomedical Imaging and Bioengineering, grant P41EB015871-27 and by the MIT SkolTech initiative. We thank Professor Tomás Palacios and Benjamin Mailly-Giacchetti for useful discussions about the pH dependence of graphene.
Author information
Authors and Affiliations
Contributions
G.L.C.P., J.T.N. and M.S.S. designed the experiments. G.L.C.P. synthesized the graphene and performed UV-VIS spectroscopy. G.L.C.P., K.Y.L., Q.H.W., B.M., J.T.N., M.P.L., J.W.K. and R.R.D. performed Raman experiments. G.L.C.P., J.T.N. and Q.H.W. processed and analyzed the Raman data. C.F.O. and K.D.W. generated the cell lines and expressed and purified the IgG. N.F.R., J.Z., B.R.G., S.K., E.V.E., J.T.N. and G.L.C.P. passaged the cells. G.L.C.P. and J.Z. devised the model in Figure 2(d). G.L.C.P., J.T.N. and M.S.S. wrote the manuscript. All authors contributed to the discussion of the results and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Paulus, G., Nelson, J., Lee, K. et al. A graphene-based physiometer array for the analysis of single biological cells. Sci Rep 4, 6865 (2014). https://doi.org/10.1038/srep06865
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06865
This article is cited by
-
Glucose measurement via Raman spectroscopy of graphene: Principles and operation
Nano Research (2022)
-
Evaluation of a biosensor based on reduced graphene oxide and glucose oxidase enzyme on the monitoring of second-generation ethanol production
Journal of Solid State Electrochemistry (2020)
-
Single-molecule detection of protein efflux from microorganisms using fluorescent single-walled carbon nanotube sensor arrays
Nature Nanotechnology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.