Abstract
Few studies have investigated exercise-induced global gene expression responses in human skeletal muscle and these have typically focused at one specific mode of exercise and not implemented non-exercise control models. However, interpretation on effects of differentiated exercise necessitate direct comparison between essentially different modes of exercise and the ability to identify true exercise effect, necessitate implementation of independent non-exercise control subjects. Furthermore, muscle transcriptome data made available through previous exercise studies can be difficult to extract and interpret by individuals that are inexperienced with bioinformatics procedures. In a comparative study, we therefore; (1) investigated the human skeletal muscle transcriptome responses to differentiated exercise and non-exercise control intervention, and; (2) set out to develop a straightforward search tool to allow for easy access and interpretation of our data. We provide a simple-to-use spread sheet containing transcriptome data allowing other investigators to easily see how mRNA of their gene(s) of interest behave in skeletal muscle following exercise, both endurance, resistance and non-exercise, to better aid hypothesis-driven question in this field of research.
Design Type(s) | parallel group design • transcription profiling by array design • time series design • intervention design |
Measurement Type(s) | transcription profiling by array assay |
Technology Type(s) | DNA microarray |
Factor Type(s) | Intervention Group |
Sample Characteristic(s) | Homo sapiens • multicellular organism |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Metabolic stress inherent of endurance exercise versus mechanical stress inherent of resistance exercise is considered to comprise essentially different stimuli that drive training-induced muscle adaptations towards opposite ends of an adaptation continuum. According to this traditional contention, increased capability for oxidative metabolism is induced by endurance training, whereas increased capability for contractile force development is induced by resistance training1–3.
The underlying myocellular mechanism involved in promoting exercise-specific adaptations is receiving substantial research attention, not least because exercise provide a highly relevant model to investigate mechanisms involved in clinical conditions affecting muscle metabolism or muscle contractile function, as well as counteracting sarcopenia in aging.
Studies on muscle phenotype regulation primarily consist of hypothesis-driven studies on pre-selected myocellular pathways, in attempts to tie specific regulatory pathways to specific types of skeletal muscle adaptations4–12. However, such studies are limited by not being able to comprise the potential effects of simultaneously cross-talking mechanisms, redundant mechanisms and/or oppositely acting mechanisms.
To provide a more complete overview on exercise-induced myocellular responses, in more recent years, omic´s-based techniques have been utilized and data have become available on exercise-induced transcriptome responses from studies on endurance exercise13–16, as well as from studies on resistance exercise17–20.
Valuable information can be retrieved from such studies, but certain aspects still render interpretation difficult. Firstly, all of the studies conducted on acute transcriptome responses to exercise have focused only on one specific type of exercise. However, oppositely to an traditional contention, exercise-induced muscle phenotype adaptations are likely dispersed along a continuum, as dictated by the physical condition of included subjects and by the choice of intensity, duration, volume, frequency and/or interspaced recovery of a given exercise protocol1–3. If exercise-induced adaptations disperse along a continuum, the responses of underlying myocellular regulatory mechanisms likely do the same, with some mechanism potentially responding essentially different to differentiated exercise, whereas others may exhibit overlap in responses. Such information is important, in order to manipulate specific mechanisms, while simultaneously avoiding unintended effects. Thus, investigation of this requires comparative investigation of endurance exercise versus resistance exercise, practised by commonly practised principles, but has not yet been conducted in humans. Furthermore, previously conducted studies include either untrained or well-trained individuals13–20. However, unfamiliar exercise may comprise an unrealistic type and/or magnitude of stress and well-trained athletes may exhibit very low sensitivity to familiar exercise. Investigation on moderately trained individuals can therefore be argued to mimic a more probable situation.
Another challenge in interpretation of exercise-induced myocellular responses relate to whether they can be even be regarded as genuine effects of exercise or if they are rather related to other confounding factors inherent of an exercise trial. In accordance, factors such as dietary condition, clinical procedures (i.e. tissue collection) and/or circadian rhythm, may all obscure interpretation21. Such knowledge requires non-exercise control experiments, yet nearly all studies on exercised-induced transcriptome responses have not.
Finally, although transcriptome data are now in principle available from exercise studies, the formats of data sets, makes them difficult to extract and therefore interpret by researchers inexperienced with the procedures and the flaws of bioinformatics. For instance, reported data most often rely on highest ranking of response based on an arbitrary cut-off level. While this may serve to highlight molecules that are highly responsive, it is not necessarily telling of biological importance. Oppositely, potentially important molecules responding below a pre-set cut-off level are easily ignored. Presentations based on degree of responsiveness are not optimal to illustrate effects on related molecular family members or molecules related to the same pathway. Pathway tools exist to obtain such information, but can still be difficult to interpret. Development of procedures to allow more simplified unravelling of omic´s data can therefore be of great value.
Based on these challenges, one major aim of the current study was to compare exercise-induced human skeletal muscle transcriptome responses to differentiated exercise in training-accustomed individuals as well as towards non-exercise control intervention. This approach involved 10 weeks of prior endurance or resistance training to familiarise subjects before conducting the actual single-bout exercise trial to examine exercise-induced responses. Another major aim of the study was to develop a simplified procedure to aid search and interpretation of our data.
Methods
Subjects
The subjects have previously been described in details22–26. In brief, fourteen young untrained healthy male subjects were randomly divided into two groups; an endurance training group and a resistance training group. A further six subjects composed the non-exercising control group. All subjects were informed about the purpose and the risks related to the study and gave written, informed consent to participate. The study was approved by the ethics committee of Region Midtjylland (j.no. M-20080177) and performed in accordance with the Declaration of Helsinki.
Experimental design and training protocol
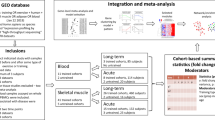
The design and training protocols have previously been described in details22–26. In brief, prior to a single-bout exercise trial aiming to study acute gene expression responses, subjects were habituated through a prior 10 week training phase. At least three day before commencing this training phase, a biopsy was harvested (i.e. termed ‘Pre’, identical to pre training and used as pre value for investigation of acute transcriptome responses to single-bout exercise—see Figure 1 and further below in text). Exercise groups completed either 10 weeks of progressive endurance training (ET) on a cycle ergometers or 10 weeks of progressive conventional resistance training (ST) for lower extremity muscle groups. ET was performed on stationary bicycles (Kettler Ergoracer GT, Kettler, Ense-parsit, Germany). 3 weekly sessions consisted of one session each of continuous cycling of 30 to 45 min at 60–75% of Watt-max, a second session consisting of two intervals of 20 min at 70–80% of Watt-max, and a third session consisting of 8×4 min intervals at 80–90% of Watt-max. The RT group completed a conventional progressive overload training program, similar to that previously described27, consisting of three leg exercises each performed as 3–5 sets of 10→6 repetitions with repetitions corresponding to RM loading. All training sessions were supervised to ensure proper progression for both groups.
Three individual groups of human subjects had a biopsy taken prior to commencing 10 weeks of intervention (Pre). They then completed 10 weeks of intervention as either endurance, resistance or no training. After several days of recovery, subjects reported at the same time point of the day to the laboratory for±single-bout exercise, in which they conducted either endurance exercise (i.e., 2 h of cycling at 60% aerobic intensity), resistance exercise (i.e., rested for 1 h followed by a 30 min session including 4 sets x 12 repetitions for three leg exercises) or no exercise (i.e., the subjects rested for 2 h). Biopsies were harvested at the same absolute time points prior to (i.e. similar to post training) and at 0, 2, 5 and 22 h post±exercise. RNA was extracted and RNA from time points corresponding to Pre, 2 and 5 h (time points for array analysis marked in red in the graph) were hybridized to GeneChip arrays followed by bioinformatics procedures and development of a search tool to allow simplified data access. In parallel, RNA from all time points were included in cDNA synthesis to be used for validation of array data by Real-time PCR.
Following three days of recovery from the final exercise session of the training phase, subjects completed a single-bout trial including either the type of exercise they were habituated to through prior training or non-exercise for the control subjects. During this trial, muscle biopsies were harvested and used for transcriptome analysis.
Single-bout exercise protocol
A schematic overview of the single-bout trial design is seen in Figure 1. Subjects arrived after an overnight fast at 8 am on day 1 for pre-exercise measurements. The subjects rested in the supine position for approximately 30 min. A pre-exercise muscle biopsy (i.e. termed ‘Post’, identical to post training) was then taken from the vastus lateralis muscle. At 8.30 am, subjects of the endurance group commenced 120 min of bicycle exercise at 60% post training Vo2 peak. Subjects in the resistance group remained resting until 10 am before commencing 4 x post training 12 RM of three thigh muscle exercises with 1 minutes rest between sets, thereby allowing them to finish at 10.30 am (i.e. same absolute time point as for subjects of the endurance group). The three thigh muscle exercises were leg press, knee extension, and hamstring curl. Minor weight adjustments were allowed to ensure performance of 12 RM as precise as possible. Non-exercise control subjects rested from 8.30 am to 10.30 am. The non-exercise control group was included, to control for potential non-exercise related effects of fasting conditions and the invasive procedures inherent of the protocol. To control for potential effects of circadian rhythm, it was attempted to strictly equalise absolute daily time points as well as time resolution of the protocol, regardless of which group the subjects were assigned to. After±exercise, all subjects rested under fasting conditions (with water offered ad libitum) until 3.30 pm (corresponding to 5 h post exercise). They were allowed normal dietary intake for the remainder of day 1 and instructed to fast overnight before arriving in fasting condition to the laboratory on the morning of day 2 of the trial. During post-exercise recovery, muscle samples were harvested at 0, 2, 5 and 22 h after exercise. All biopsies were harvested from separate incision holes and dispersed between legs.
Preparation of muscle biopsies and RNA isolation
Biopsy samples harvested during the single-bout exercise trial were rinsed from visible fat and connective tissue and frozen in liquid nitrogen as quickly as possible. All samples were then stored at −80 °C until further investigation. Total RNA (from approximately 20 mg of muscle) was extracted using the guanidinium thiocyanate–phenol–chloroform extraction method as previously described28 and the concentration was determined spectrophotometrically using a Nanodrop 1000 (Thermo Fischer Scientific, Wilmington, DE, USA). RNA integrity was ensured by gel electrophoresis.
Microarray
From six subjects in each of the three exercise groups, samples from the three time points, Pre (i.e. pre training), 2 h and 5 h, were selected for microarray analysis (54 samples). 10 μl total RNA (2.5—8 μg) was purified further using the RNeasy MinElute Cleanup kit (Qiagen). The purified RNA was amplified and labeled using the Ambion WT Expression Kit (Applied Biosystems) according to manufactures instructions. 100 ng total RNA was used as input. The labeled samples were hybridized to the Human Gene 1.0 ST GeneChip array (Affymetrix, Santa Clara, CA, USA). The arrays were washed and stained with phycoerytrin conjugated streptavidin (SAPE) using the Affymetrix Fluidics Station® 450, and the arrays were scanned in the Affymetrix GeneArray® 3000 scanner to generate fluorescent images, as described in the Affymetrix GeneChip® protocol. Cell intensity files (CEL files) were generated in the GeneChip® Command Console® Software (AGCC) (Affymetrix, Santa Clara, CA, USA).The CEL files were analyzed using Partek Genomic Suite v6.6 (Partek), using the default Partek settings (GC content pre-adjustment, RMA backgrounds correction, quantile normalization, mean probe set summarization) to generate transcript expression levels (Data Citation 1). To test for unintended group differences, an ANOVA for Exercise was performed on the Pre samples only. No transcripts came out with a FDR<0.05. To facilitate comparison of changes over time all transcript expression values were normalized to their respective pre value for each subject. A repeated measures two-way ANOVA for Time, Exercise and Time x Exercise (REML for Variance Component Estimation) was performed to generate p-values for overall effects as well as between individual subgroups (Contrasts). Transcripts exclusively induced in a single group were identified as follows. The transcript level has to be significantly different from both Pre and the two other groups for at least one of the two time points. Furthermore, the transcript must not be significantly different from Pre at any time in the other groups. Time regulated transcripts were identified as follows: Time x Group (raw P>0.1) and Time (FDR<0.05) at 2.5 h or 5 h and all three groups should be significantly different from Pre (raw P<0.05) at the same time point. FDR (False discovery rate) of less than 0.05 is used as significance level unless stated otherwise.
Real-time PCR
500 ng total RNA was converted into cDNA in 20 μl using the OmniScript reverse transcriptase (Qiagen, California, USA) and 1 μM poly-dT (Invitrogen, Naerum, Denmark) according to the manufacture's protocol (Qiagen). For each target mRNA, 0.25 μl cDNA was amplified in a 25 μl SYBR Green polymerase chain reaction (PCR) containing 1×Quantitect SYBR Green Master Mix (Qiagen) and 100 nM of each primer (Table 1). The amplification was monitored real time using the MX3005P Real-time PCR machine (Stratagene, California, USA) with the amplification protocol; 95°,15′->{95°,15′′->58°,30′′->63°,90′′(signal)}x50->melt curve analysis. The Ct values were related to a standard curve made with known concentrations of cloned PCR products or DNA oligonucleotides (UltramerTM oligos, Integrated DNA Technologies, Inc., Leuven, Belgium) with a DNA sequence corresponding to the sequence of the expected PCR product. The specificity of the PCR products was confirmed by melting curve analysis after amplification. The large ribosomal protein P0 (RPLP0) mRNA was chosen as internal control.
Data Records
Identification of regulated transcripts
To identify exercise induced mRNA, a microarray comparison was performed using the Pre samples and the two acute samples obtained during recovery from the single-bout trial, 2 h and 5 h. Both acute changes within the first hours after exercise as well as long-term training changes would be caught by comparing the 2 h and 5 h Post expression values with the Pre value. Six subjects from each exercise group were analysed using the Affymetrix Gene ST 1.0 arrays (28869 transcripts) (Data Citation 1). As the values of interest here are the changes from Pre, all expression values were normalized to the respective pre value before further analyses. Two-way repeated measures ANOVA shows that around a thousand mRNA were significantly changed in both the resistance and the endurance groups, whereas a much lower number were changed in the control group (Table 2). Comparisons between the groups within the two Post exercise time points also show massive differences between the groups, including endurance versus resistance, indicating large and different impact of endurance and resistance exercise on the transcriptome (Tables 3 and 4).
Several hundred transcripts can be identified as specifically responsive to a single exercise type, especially in the resistance training group (Table 5).
About one hundred transcripts can be identified as generally responding to exercise, independent of the exercise type, whereas, using very stringent criteria, 36 can be found to respond to both exercise type, but in different ways (Table 6).
The finding of some changes in the control group indicates a general time dependent effect as well, e.g. from circadian rhythm or dietary status. In principle such transcripts are those that display a significant time effect, but not an interaction, in the ANOVA. However, since 2/3 of the samples are exercise samples, a general exercise effect might erroneously also give such an effect, due to the relative low number of control samples compared to the exercise samples. To exclude those, a stringent criterion was added, such that furthermore, all three groups should have a raw P-value of less than 0.05 compared to Pre at the specific time point. By these criteria, 192 time regulated transcripts were identified, 131 at 2.5 h and 103 at 5 h (Table 7).
Technical Validation
To validate the microarray data, we tested the expression of a few differentially expressed mRNA by another technique, real-time RT-PCR. In this case we included all time points to also test time points which were not included in the statistical analysis of the microarrays, i.e. outside potential selection bias.
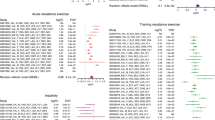
As can be seen in Figure 2 RT–PCR, there is a good correspondence between the results obtained with the microarray data and the results using real-time RT-PCR. Furthermore, the pattern is replicated in some of the time points not included in the microarray analysis, indicating that this is not a statistical artifact (type II error).
Comparison of results obtained from the microarray data with results obtained using standard real-time RT–PCR. Only time-points Pre and 2 and 5 h Post final exercise session are measured with microarrays (Left). All time-points were measured with real-time RT-PCR (Right). MYOG (Myogenin) and EIF2AK3 (eukaryotic translation initiation factor 2-alpha kinase 3) were identified as strength specific and JUND (Jun D) as exercise specific, but dependent on exercise type. Data are shown as geometric mean±back-transformed s.e.m. on a logarithmic axis (log2).
Usage Notes
The data from this transcriptome analysis is summarized in an Excel file which can be downloaded from the GEO database (Data Citation 1).
The file contains five sheets containing instructions, statistical analysis and mean data, individual data, and two presentation sheets for mean and individual data, respectively.
The intention is that the reader can generate a list of all mRNA regulated by a specific condition or investigate the regulation of one or more specific mRNA by using the filtering function on the statistics or the names, respectively. Bar graphs or dot plots can then be easily generated by tapping in the unique ID’s in the two graph sheets.
Figure 3 Example shows an example of the graphs generated for the VEGFA (Vascular Endothelial Growth Factor A) mRNA simply by writing ‘VEGFA’ in the two graph sheets.
Illustration of the graphs that can easily be generated for a specific mRNA using the Excel file. In this case the VEGFA mRNA. Besides getting the graphs, the statistical result is also provided, here showing that VEGFA mRNA is differentially regulated by exercise (Differential Exercise) and that within the Endurance group both 2.5 and 5 h are significantly different from Pre (E2.5h and E5h), whereas for the Strength group only 5 h is significant (S5h). Furthermore, within 2.5 h Endurance is different from both Control (EvC2.5h) and Strength (SvE2.5h), whereas at 5 h Endurance is only significantly different from Control (EvC5h) but not Strength (no SvE5h). On the other hand also Strength is different from Control at 5 h (SvC5h).
Additional information
How to cite this article: Vissing, K. & Schjerling, P. Simplified data access on human skeletal muscle transcriptome responses to differentiated exercise. Sci. Data 1:140041 doi: 10.1038/sdata.2014.41 (2014).
References
References
Spiering, B. A. et al. Resistance exercise biology: manipulation of resistance exercise programme variables determines the responses of cellular and molecular signalling pathways. Sports Med. 38, 527–540 (2008).
Wernbom, M., Augustsson, J. & Thomee, R. The influence of frequency, intensity, volume and mode of resistance training on whole muscle cross-sectional area in humans. Sports Med. 37, 225–264 (2007).
Hawley, J. A. Adaptations of skeletal muscle to prolonged, intense endurance training. Clin. Exp. Pharmacol. Physiol. 29, 218–222 (2002).
Chin, E. R. et al. A calcineurin-dependent transcriptional pathway controls skeletal muscle fiber type. Genes Dev. 12, 2499–2509 (1998).
Wu, H. et al. MEF2 responds to multiple calcium-regulated signals in the control of skeletal muscle fiber type. EMBO J. 19, 1963–1973 (2000).
Atherton, P. J. et al. Selective activation of AMPK-PGC-1alpha or PKB-TSC2-mTOR signaling can explain specific adaptive responses to endurance or resistance training-like electrical muscle stimulation. FASEB J. 19, 786–788 (2005).
Goodman, C. A. et al. A phosphatidylinositol 3-kinase/protein kinase B-independent activation of mammalian target of rapamycin signaling is sufficient to induce skeletal muscle hypertrophy. Mol. Biol. Cell 21, 3258–3268 (2010).
Sartori, R. et al. BMP signaling controls muscle mass. Nat. Genet. 45, 1309–1318 (2013).
Vissing, K. et al. Differentiated mTOR but not AMPK signaling after resistance vs endurance exercise in training-accustomed individuals. Scand. J. Med. Sci. Sports 23, 355–366 (2013).
von Maltzahn, J., Chang, N. C., Bentzinger, C. F. & Rudnicki, M. A. Wnt signaling in myogenesis. Trends Cell Biol. 22, 602–609 (2012).
Leger, B. et al. Akt signalling through GSK-3beta, mTOR and Foxo1 is involved in human skeletal muscle hypertrophy and atrophy. J. Physiol. 576, 923–933 (2006).
Stefanetti, R. J. et al. Influence of divergent exercise contraction mode and whey protein supplementation on atrogin-1, MuRF1 and FOXO1/3A in human skeletal muscle. J. Appl. Physiol. (1985) 116 (11): 1491–1502 (2014).
Catoire, M. et al. Pronounced effects of acute endurance exercise on gene expression in resting and exercising human skeletal muscle. PLoS ONE 7, e51066 (2012).
Busso, T. & Fluck, M. A mixed-effects model of the dynamic response of muscle gene transcript expression to endurance exercise. Eur. J. Appl. Physiol. 113, 1279–1290 (2013).
Rowlands, D. S. et al. Transcriptome and translational signaling following endurance exercise in trained skeletal muscle: impact of dietary protein. Physiol. Genomics 43, 1004–1020 (2011).
Mahoney, D. J., Parise, G., Melov, S., Safdar, A. & Tarnopolsky, M. A. Analysis of global mRNA expression in human skeletal muscle during recovery from endurance exercise. FASEB J. 19, 1498–1500 (2005).
Murton, A. J. et al. Transient transcriptional events in human skeletal muscle at the outset of concentric resistance exercise training. J. Appl. Physiol. (1985) 116, 113–125 (2014).
Thalacker-Mercer, A. et al. Cluster analysis reveals differential transcript profiles associated with resistance training-induced human skeletal muscle hypertrophy. Physiol. Genom. 45, 499–507 (2013).
Gordon, P. M. et al. Resistance exercise training influences skeletal muscle immune activation: a microarray analysis. J. Appl. Physiol. (1985) 112, 443–453 (2012).
MacNeil, L. G., Melov, S., Hubbard, A. E., Baker, S. K. & Tarnopolsky, M. A. Eccentric exercise activates novel transcriptional regulation of hypertrophic signaling pathways not affected by hormone changes. PLoS ONE 5, e10695 (2010).
Vissing, K., Andersen, J. L. & Schjerling, P. Are exercise-induced genes induced by exercise? FASEB J. 19, 94–96 (2005).
Farup, J. et al. Muscle morphological and resistance adaptations to endurance vs. resistance training. J. Resistance Cond. Res. 26, 398–407 (2012).
Vissing, K. et al. Effect of resistance exercise contraction mode and protein supplementation on members of the STARS signalling pathway. J. Physiol. 591, 3749–3763 (2013).
Lamon, S. et al. Regulation of the STARS signaling pathway in response to endurance and resistance exercise and training. Pflugers Arch. 465, 1317–1325 (2013).
Møller, A. B. et al. Resistance exercise, but not endurance exercise, induces IKKβ phosphorylation in human skeletal muscle of training-accustomed individuals. Pflugers Arch. 465, 1785–1795 (2013).
Stefanetti, R. J. et al. Regulation of ubiquitin proteasome pathway molecular markers in response to endurance and resistance exercise and training. Pflugers Arch. doi:10.1007/s00424-014-1587-y (2014).
Vissing, K. et al. Muscle adaptations to plyometric vs. resistance training in untrained young men. J. Resistance Cond. Res. 22, 1799–1810 (2008).
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159 (1987).
Data Citations
Schjerling, P., & Vissing, K. Gene Expression Omnibus GSE59088 (2014)
Acknowledgements
We thank the subjects for their participation in the project. Mikkel H. Vendelbo is thanked for clinical assistance. Jean Farup and Tue Kjølhede are thanked for assisting with testing and training procedures. Anja Jokipii-Nielsen and Ida Trondhjem Haakansson are thanked for excellent technical assistance. Furthermore, Rehannah Borup and the RH Microarray Center at Rigshospitalet for running the microarrays. Funding is gratefully acknowledged from Antidoping Danmark, the Nordea Foundation (Healthy Aging grant), the Danish Medical Research Council and the NovoNordiskFoundation. Kristian Vissing was supported by the NovoNordiskFoundation. K.V. and P.S. had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Contributions
Conception and design of the experiments: K.V. and P.S. Analysis and interpretation of data: K.V. and P.S. Drafting the article: K.V. and P.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Vissing, K., Schjerling, P. Simplified data access on human skeletal muscle transcriptome responses to differentiated exercise. Sci Data 1, 140041 (2014). https://doi.org/10.1038/sdata.2014.41
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2014.41
This article is cited by
-
Skeletal muscle phenotyping of Hippo gene-mutated mice reveals that Lats1 deletion increases the percentage of type I muscle fibers
Transgenic Research (2022)
-
Genomic predictors of testosterone levels are associated with muscle fiber size and strength
European Journal of Applied Physiology (2022)
-
Time trajectories in the transcriptomic response to exercise - a meta-analysis
Nature Communications (2021)
-
Impact of protein supplementation during endurance training on changes in skeletal muscle transcriptome
BMC Genomics (2020)
-
Transcriptomic profiling of skeletal muscle adaptations to exercise and inactivity
Nature Communications (2020)