Abstract
Understanding how soil ecosystem responds to transgenic Bacillus thuringiensis (Bt) rice is necessary for environmental risk assessment. While the influences of short-term cultivation of Bt rice on soil properties have been reported previously, little is known about the long-term effects of Bt rice on soil ecosystems. In this study, soil samples were taken from a long-term rice cultivation site in Fujian Province, China, where transgenic Bt rice (Kefeng-6) and its non-Bt parent breed (Minghui-86) had been continuously cultivated for 8 years. Soil Bt protein concentration and a total of 16 variables were analyzed to assess potential risks of soil health under Bt rice cultivation. The results revealed that soil Bt protein is unlikely to accumulate after Bt rice cultivated in the field, and no consistently significant changes were observed in soil enzymatic activities (catalase, dehydrogenase, acid phosphatase, and urease), microbial biomass (microbial carbon and nitrogen), total organic carbon, decomposition (soil respiration, Q 10, and qCO2), soil nitrogen and phosphorus contents. Due to a local tradition that aboveground biomass was removed after harvest, the increased net primary productivity by Bt rice cultivation did not significantly change soil C cycling. Results of this study suggested that on the aspects of soil microbial functioning and C cycling, long-term cultivation of Bt rice is unlikely to result in significant effects on soil health.
Similar content being viewed by others
Introduction
Transgenic Bacillus thuringiensis (Bt) crops are genetically modified to express certain larvicidal toxins to kill target pests. These insect-resistant crops have a great advantage in agriculture by reducing pesticide application1. Transgenic Bt crops, however, release Bt protein into environments through above- and below-ground debris and root exudates, which may lead to profound impacts on agro-ecosystems. Previous studies have reported that transgenic Bt crops may beget compound effects on agro-ecosystems, such as gene flow between plant species2, developing more resistance to Bt protein in target insects3,4,5, and potential impacts on non-target insects6,7,8. Soil is the core of agro-ecosystem, supporting crop growth, serving as a nutrient reservoir and providing the habitat of belowground organisms that mediate terrestrial nutrient cyclings. It is imperative to understand whether soil health has been significantly influenced by Bt crops.
It was reported that Bt protein from Bt crops could persist in the soil, mainly through binding to humus or clay9,10,11, and remained functioning for a relatively long period12. On the aspect of soil health, it is important to understand whether Bt crops can significantly change soil biota, microbial activities and ecological processes (e.g., soil carbon and nutrient cyclings). A large number of previous studies have been conducted to explore possible effects of Bt crops on soil ecosystem but obtained inconsistent results. Li and Liu13 found that the richness and diversity of nematode were not significantly altered by Bt cotton. Similar results were reported on herbivory nematode14, earthworm and protozoa15. Besides, a few studies pointed out that Bt crops did not significantly change soil microbial biomass16 or soil enzymatic activities17,18,19. Soil total organic carbon (TOC) and total nitrogen (TN) pools were insensitive to cultivation of Bt crops20,21,22. In situ soil respiration rates did not significantly differ between Bt and non-Bt crops18, 23. On the other hand, it was reported that growing Bt crops might cause some significant changes in soil variables related to soil health. For instance, Bt cotton could increase the diversity of fungi and bacteria24. Bt crops enhanced dehydrogenase17, 25 and urease activities26. Fliessbach et al.25 found that Bt maize increased the dehydrogenase activity by 6%. In addition, soil respiration could be significantly increased27, 28 or decreased25 by transgenic Bt maize in different studies. Most of these reported studies were based on short-term field experiments, up to two or three years13, 29, 30, enabling us to learn what and how Bt crops affect soil microbial organisms and soil eco-processes. However, short-term cultivation experiments encountered temporal variations in soil environment and crop growth, were not able to reflect the accumulative effects of Bt protein along with the food web in the soil, it is still poorly understood whether long-term cultivation of Bt crops will result in irreversible impacts on soil health.
Furthermore, previous studies almost centered on Bt-corn and Bt-cotton18, 25, 26 and it is indispensable to evaluate soil health under Bt rice cultivation according to the case-by-case principle of biosafety assessment. Rice is one of major food sources in the world, and rice planting area covers more than 150 million hectares31. In China, many breeds of transgenic Bt rice have been reported to promote food production and to reduce pesticide applications1, but none has been commercially released to date. Transgenic Bt rice is likely to be commercially released in near future, to meet increasing food demand. It is an urgent task to understand potential threats to soil health of long-term cultivation of transgenic Bt rice. Under the canopy of Bt rice, soil microbial biomass declined by 7%32; dehydrogenase activity increased by 95% at the initial stage of straw decomposition but decreased by 47% at subsequent stages33; soil respiration rate increased by 25%34. All of these studies were based on short-term experiments (1–3 years), mainly due to the lack of long-term experimental platform of transgenic Bt rice cultivation.
Using knowledge from short-term cultivation of Bt rice may not reliably predict the long-term effects of Bt rice on soil ecosystem, because that significant shifts in soil traits might occur after lasting accumulation of slight changes caused by Bt rice, or that these slight changes might offset each other, resulting in no significant long-term effects on soil ecosystem. It is necessary and urgent to assess the long-term effects of Bt rice on soil ecosystem before Bt rice is commercially released and widely planted. Because of the lack of long-term experimental platform, assessing the potential environment risk of Bt rice is still a difficult task. In this study, we took advantage of an experimental platform for breeding Bt rice Kefeng-6 (encoding Cry1Ac protein), the longest one of Bt rice cultivation in China where Bt rice had been continuously cultivated for 8 years when we conducted this study, to assess the possible effects of Bt rice on soil health. The objectives of this study were to investigate: (1) whether long-term cultivation of Bt rice causes the accumulation of Bt protein in soil; (2) whether there are irreversibly negative effects of Bt rice cultivation on soil properties mediating soil carbon and nitrogen cycling and maintaining ecosystem health, such as enzymatic activities, microbial biomass, and dynamics of C pool and decomposition.
Results
Plant biomass, C/N ratio, and Bt protein concentration
The straw biomass of Bt rice was significantly higher than that of non-Bt rice (t = 2.85, p < 0.05; Fig. 1a). However, no significant differences were observed in root biomass between Bt and non-Bt rice, either in soil layer of 0–10 cm (t = 1.14, p = 0.29) or 10–20 cm (t = −2.03, p = 0.08). The C/N ratios of root, stem and leaf were similar between Bt rice and its parental breed (t = 0.87, p = 0.43; t = 0.19, p = 0.10; t = 0.68, p = 0.53, respectively; Fig. 1b). Bt protein was not detected out using enzyme-linked immunosorbent assay (ELISA) approach in soil samples collected from Bt treatment.
Enzymatic activities
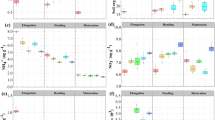
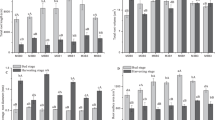
In general, there were no consistently significant differences in soil (0–20 cm) catalase activity among all three treatments (Fig. 2a). During summer time when rice was growing, catalase activity was significantly higher in Non-GM than those in other treatments. In autumn after rice harvest, Half-GM had a lower soil catalase activity than GM and Non-GM, but the differences were not statistically significant (F = 3.55, p = 0.07).
Differences in dehydrogenase activity among treatments varied over time (Fig. 2b). During spring and summer, dehydrogenase activities in Non-GM were significantly lower than those in GM and Half-GM (F = 7.61, p = 0.01; F = 16.56, p < 0.001). After rice harvest, soil samples of three treatments exhibited similar level of dehydrogenase activities (F = 0.66, p = 0.54).
In spring before rice planting, differences in acid phosphatase activity among treatments were not significant (F = 1.58, p = 0.26; Fig. 2c). During summer and autumn, Non-GM had significantly lower acid phosphatase activity than Half-GM and GM (F = 15.92, p < 0.001; F = 4.15, p = 0.05). No consistently significant differences in phosphatase activity were observed between GM and Half-GM treatments throughout the year.
Although Half-GM had the lowest urease activity among the three treatments through the year (Fig. 2d), the differences were significant only between Half-GM and GM treatments.
Soil respiration
Figure 3a shows soil respiration variation along with incubation temperature for soil samples (0–20 cm) taken in summer time. Soil respiration rate against temperature in other seasons was similar (data not shown). The reference respiration rates (R 20) were 0.0168 ± 0.0003, 0.0165 ± 0.0003, and 0.0173 ± 0.0006 μmol C g−1 dry soil h−1 for GM, Half-GM and Non-GM, respectively. Estimated Q 10 values of soil respiration were 2.34 ± 0.04, 2.24 ± 0.03 and 2.38 ± 0.10 for GM, Half-GM and Non-GM treatment, respectively, suggesting that the temperature sensitivity of TOC decomposition was not significantly influenced by treatments (F = 0.94, p = 0.41). Similarly, soil microbial metabolic quotients (qCO2) were similar among treatments (F = 2.08, p = 0.18), indicating that the overall microbial activity was not significantly changed by different treatments (Fig. 3b).
Soil carbon and nutrients
No consistent differences in soil microbial biomass carbon (MBC) and nitrogen (MBN) were observed among Bt treatments along with time (Fig. 4a and b). Although MBC in Half-GM was the lowest, the differences were statistically significant only in autumn after rice harvest (F = 14.09, p = 0.001). MBN in Half-GM was the highest in summer (F = 5.10, p = 0.02), but the lowest in autumn (F = 14.09, p = 0.001). In spring before rice planting, the ratio of MBC/TOC in GM was higher than those in Half-GM and Non-GM. However, Non-GM had the highest ratios in summer and autumn, significantly greater than those in other two treatments (F = 10.48, p = 0.002; F = 56.34, p < 0.001; Fig. 4c).
TOC and TN were at similar levels among the three treatments, except in summer when TOC and TN were significantly higher in Half-GM than those in Non-GM (F = 8.26, p = 0.005). The annual mean C/N ratios were 10.87, 11.01 and 10.81 for GM, Half-GM and Non-GM, respectively, suggesting no systematically significant effect by Bt treatments (Fig. 5).
In spring before rice planting, soils had the highest phosphorous availability, with no significant differences among Bt treatments (Fig. 6). In general, soil phosphorus availability among three treatments was similar during summer and autumn. However, after rice harvest, available phosphorus concentration in Non-GM was significantly lower than those in Half-GM and GM (p = 0.016).
The NMDS ordinations, based on year-round C and nutrient pools (Fig. 7a), and soil enzymatic activities and microbial biomass (Fig. 7b), show close overlaps among soils from different treatments, indicating that Bt rice does not bring about obvious changes on soil properties throughout the year.
Nonmetric multidimensional scaling (NMDS) analyses of similarities among Bt treatments. (a) year-round carbon and nutrient pools (TOC, TN, available phosphorus); (b) soil enzymatic activities and microbial biomass (catalase, dehydrogenase, acid phosphatase, urease, MBC and MBN). Circles stand for soil samples from Non-GM, triangles for Half-GM and squares for GM. Ellipses were the 95% confidence limits of multivariate means.
Discussion
Bt protein in the soil
In general, Bt protein concentration in the soil under transgenic Bt rice is relatively low, often lower than the minimal detecting level of ELISA method35. Higher levels of Bt protein concentration, 0.82–2.13 ng g−1 dry soil in rhizosphere soils, were reported under cultivation of Bt-Minghui 63 and Bt-Shanyou 63 36. In this study, no Bt protein was detected from the soil under Bt rice. Based on the amount of root biomass at the time of harvest and Bt protein concentration of roots, the annual input of Bt protein into the soil through root turnover was estimated at about 1.0 ng g−1 in 0–10 cm and 0.24 ng g−1 in 10–20 cm depth. The input of Bt protein through exudation from live root was difficult to be estimated. Bt protein from either root turnover or root exudation should concentrate in rhizosphere rather than in bulk soil. The low concentration of Bt protein may render us unable to detect out Bt protein in root-free soil samples using ELISA in our study. Another possible reason might be partly related to the short residence time of Bt protein in soil. Feng et al. pointed out that the half-life of Bt protein in the soil varied from 0.75 to 10.89 days, and only 0.02–1.51% remained in the soil after 180 days37. Commonly, Bt protein cannot be detected in soils at the time of next growing season starting38. Our study suggested that long-term cultivation of Bt rice is unlikely to result in the accumulation of Bt protein in the soil.
Effects of Bt rice on C cycling
The major input of organic matter into the soil ecosystem is from plants through above-ground litterfall, root turnover and root exudation. Changes in crop traits and the quantity and/or quality of C input into the soil may affect soil carbon cycling39, 40. It has been reported that Bt rice had a different tiller number41 and net photosynthetic rate42, leading to differences in crop biomass between Bt and non-Bt crops. Saxena and Stotzky43 pointed out that lignin concentration of Bt maize was 33–97% higher than those of non-Bt ones, partly due to a greater diameter of vascular bundle and surrounding lignin cell. Bt gene may also change organic carbon and nitrogen concentrations in Bt crops relative to non-Bt crops44, 45. By contrast, Bt rice (Kefeng-6) did not have significantly different root biomass, root distributions between surface and sub-surface soil layers, or root C:N ratio compared with its non-Bt parental breed (Fig. 1). Although the aboveground biomass of Bt rice was significantly higher, it did not result in a higher C input to the soil as most part of aboveground straw was removed or burned after harvest. For a long term and large scale commercial release of Bt rice, increased aboveground biomass may lead to a greater amount of C accompanied with Bt protein input to the soil.
In situ determination of root exudations is still a challenge to ecological research46. It was estimated that about 30–60% of photosynthetically fixed carbon was allocated into belowground47,48,49, which then turned into root biomass increment, root respiration and exudation. Under relatively stable environmental conditions, root exudation may be positively correlated with root biomass50. Recently, it was reported that root exudation was sensitive to the level of available phosphorus in the soil51, 52. Our experimental results of similar root biomass and soil phosphorus availability among Bt treatments (Figs 1a and 6) may suggest no significant change in root exudation between Bt and non-Bt rice.
The reference respiration (e.g. R 20 in this study) of root-free soil samples defined soil organic matter decomposition rate under given conditions and Q 10 value defined the dependence of TOC decomposition on temperature. As soil water content was kept at 60% of field capacity during soil incubation, R 20 and Q 10 reflected the essential characteristics of TOC decomposition in soils of different Bt treatmetns. Meanwhile, due to similar TOC contents in different soil samples, R 20 could also indicate the relative TOC decomposibility, i.e. the quality of TOC. Our results suggested that after 8 years cultivation of transgenic Bt rice Kefeng-6, TOC decomposition has not been significantly changed (Fig. 3). In situ soil respiration under Bt crops has been observed increased27, 53 or decreased25. Fang et al. reported that Bt rice (Huachi-B6) could significantly reduce anaerobic soil respiration by 22.4% at the heading stage but only 2.78% at filling stage29. Field measured soil respiration includes both root respiration and TOC decomposition, regulated by many other factors other than Bt protein. The possible mechanisms underlying altered in situ soil respiration under Bt crops are complicated and still unclear. Changed crop debris input might be one of the reasons that changed soil respiration under Bt maize54.
Enzymatic activities and microbial biomass
Bt rice did not induce consistently significant changes in soil enzymatic activities after 8-year cultivation, although significant differences among treatments were observed in summer time when Half-GM and GM treatments reduced catalase activities and GM increased dehydrogenase and acid phosphatase activities (Fig. 2). These results from Bt rice seemed to be different from previous studies on other Bt crops. It was reported that soil enzymatic activities including dehydrogenase and acid phosphatase were lowered after Bt cotton consecutively planted for 3–5 years55. Sarkar et al. found that Bt cotton issued in higher enzymatic activities, MBC, MBN, and MBC/TOC21. There may be two possible reasons, i.e. Bt protein concentration and the amount of crop debris input to soil ecosystem, accounting for this divergence. Bt concentrations under Bt rice cultivations were much lower than other Bt crops where Bt protein concentration could be up to 56.14 ng g−1 under Bt cotton22. Besides, due to aboveground litter removed, there were no significant differences in crop debris input to soil between Bt rice and non-Bt rice in this study. Our results were consistent with reported short-term studies on Bt rice19. Eight years cultivation of Bt rice (Kefeng-6) has not significantly nor consistently changed soil enzymatic activities. The results of MBC and MBN in this study revealed that there were no consistently significant differences among treatments after 8 years cultivation (Fig. 4). Similar results were observed in other short-term studies21, 30 or long-term studies on Bt maize36, 56. The fact that microbial metabolic quotients among Bt rice treatments were close to one another also suggested that soil microbial biomass was not affected by Bt rice Kefeng-6.
Effects of long-term Bt rice cultivation on soil ecosystem
Agricultural soil is a complex system under intensive human disturbances. Some components of the system, such as enzymatic or microbial activities, respond instantaneously and sensitively to environmental changes. In most cases, these responses are reversible if changed environmental factors return to previous situation. Furthermore, sensitive components of soil system are often highly heterogeneous and variable57, 58. The other components of soil ecosystem are relatively insensitive to environmental changes, such as TOC. The amount of TOC reflects an accumulative result of changed influencing factors over a relatively long period. The amount of TOC may stay stable even if changed influencing factors return to previous stituation. So far, the effects of transgenic Bt crops on soil ecosystem have been assessed seperately on enzymatic activities19, 29, TOC21, 40, soil respiration56 and microbial biomass21, 26 in a short term and significant differences were occasionally observed. However, taking into consideration all these arguments in this study, no consistent changes have been observed in soil enzymatic activities, microbial biomass, C cycling after 8 years cultivation of transgenic Bt rice Kefeng-6, suggested that soil ecosystem had not been irreversibly changed by Bt rice (Fig. 7). With a large scale commercial release of Kefeng-6, increased aboveground biomass may increase organic matter input to the soil, resulting in an increase in TOC and some irreversible changes of soil ecosystem, but these possible changes are unlikely to be negative.
The experimental site of this study was initially constructed for Bt rice breeding, a fully random design of soil sampling procedure at this site was practically impossible. The pseudo-replication of sampling procedure may bring in some uncertainties to results due to possibly heterogenous background among different treatments. Indeed, pseudo-replication design cannot statistically factor out the influences of background heterogeneity from measured differences among treatments. However, this pseudo-replication was unlikely to cause a serious bias to measured results and conclusion that there was no significant differences among different Bt treatments. In this study, coefficients of variation (CV) of measured soil variables showed no consistent tendences in different seasons, as well as in different treatments (data not shown). Taking the activities of catalase for example, the greatest CV was observed in Half-GM (0.15), and Non-GM and GM was at the similar level (approximately 0.10) before rice planting. Whereas, the greatest value apeared in GM and the lowest one in Non-GM after rice harvest. In addition, the greatest CV of MBC appeared in Non-GM in spring (0.11), GM in summer (0.08), and Half-GM in autumn (0.06), respectively. No systematical or significant differences in soil variables was observed within and between treatments. This suggested that either the influence of background heterogenity was relatively small, or the agro-ecosystem has a capacity to buffer slight differences in soil background.
Conclusions
After 8 years cultivation of transgenic Bt rice Kefeng-6, no Bt protein was detected out from the soil. In addition, soil enzymatic activities, microbial biomass and metabolic quotient, soil organic carbon pools and decomposition have not been significantly nor consistently changed. Although the net primary productivity of Bt rice was greater than that of non-Bt rice, C allocation to belowground were similar. Due to above-ground straw was removed after rice harvest, increased net primary productivity by Bt rice has not resulted in significant changes in soil C cycling. From the aspects of soil microbial functioning and C cycling, long-term cultivation of transgenic Bt rice Kefeng-6 is unlikely to cause irreversibly effects on soil ecosystem.
Materials and Methods
Site and experimental treatments
Field experiment was estabished in 2002 in an authorized and confined Biosafety Experimental Field at Wufeng Village of Fuzhou City, Southeastern China. Three experimental blocks (Fig. 8) were set up for: (1) GM treatment, continuously growing transgenic Bt rice (Kefeng-6, encoding Cry1Ac protein with lepidopteron being the target insect); (2) Half-GM treatment, transgenic Bt rice (Kefeng-6) and non-transgenic rice (Minghui-86, the parent breed of Kefeng-6) being alternatively cultivated in subsequent years, and (3) Non-GM control, continuously growing non-transgenic rice Minghui-86. The dimension of each block was about 60 × 25 m. Each block was separated from others by a hard dam and had an independent irrigation system. Half-GM block was divided into 3 × 3 m plots. Bt rice (Kefeng-6) and non-Bt rice (Minghui-86) were planted in adjacent plots. In the consecutive year, Bt plots were changed to non-Bt plots, and vice versa. Each block was split into four sub-blocks, which could be considered as pseudo-replications. All treatments were under traditional management.
The schematic diagram of experimental design. Non-GM treatment was always planted non-Bt rice (Minghui-86); GM was planted Bt rice (Kefeng-6); and Half-GM was a treatment where Bt rice was planted in one plot and non-Bt rice was planted in all adjacent plots. In consecutive year, Bt plot was changed to non-Bt plot, and vice versa. Reservation area was always planted non-Bt rice.
Soil and plant roots sampling
Soil samples were collected from 0–20 cm depth in spring (before rice planting), summer (rice growing season), and autumn (after rice harvest). A soil auger of 5 cm inside diameter was used to collect soil. For GM and Non-GM treatments, 5 locations were randomly selected in each sub-block. Collected soils from these 5 locations were composited into a soil sample. For Half-GM treatment, one location was randomly selected in each plot, and collected soils were then composited to obtain one soil sample. After visible roots removed, all soil samples were transported to laboratory within 24 h. A subsample was immediately kept at 4 °C for enzymatic activity analyses (except for catalase) and the remaining part was milled to 2 mm and also kept at 4 °C for chemical, biological analyses and soil incubation. For each soil sample, a sub-sample of roughly 10 g was air-dried and milled to 0.15 mm for TOC and TN measurements. Soil properties of the three treatments are presented in Table 1.
Aboveground rice biomass and roots were collected just before rice harvest. For aboveground biomass sampling, five quadrats of 50 × 50 cm2 were randomly selected in each treatment site. Aboveground biomass was clipped to ground level and separated into stem, leaf and seed. Surface litter was then carefully removed. A soil cube of 25 × 25 × 10 cm was taken from 0–10 and 10–20 cm depths in each quadrat. Soil cubes were washed manually over a 2-mm mesh size nylon net using clear water. All plant samples were then washed with deionized water and dried at 60 °C to constant weight. Subsamples of root, stem and leaf were milled to 0.15 mm for measurement of TOC and TN. These analyses were conducted within 6 months after samplings.
Chemical and biological analyses
Soil pH was determined at a soil: water ratio of 1:5 (w/v). Available soil phosphorus content was colorimetrically determined59. TOC and TN were analyzed on a NC soil analyzer (Flash EA 1112 series; Thermo Finnigan, Elk Grove Village, IL), with combustion at 625 °C and 900 °C, respectively. Bt protein concentration in the soil was determined using ELISA approach with the phosphate-buffered saline Tween 20 and the detailed procedures was in accord with Xiao et al.60, which has a generally minimal detecting level of 0.5 ng g−1 dry soil35.
Soil MBC and MBN were analyzed using the chloroform fumigation-extraction method61. After pre-incubated at 25 °C for 15 days, two fumigated and two non-fumigated soil samples, 20 g dry weight equivalent, were extracted with 80 ml 0.5 M K2SO4 and horizontally shaken at 250 rev min−1 for 45 min. Carbon and nitrogen concentrations in the extracts were analyzed with a TOC analyzer (Multi N/C 3100, Jena, Germany). MBC and MBN were estimated following Wu et al.61, taking K EC = 0.45.
Enzymatic activities
Catalase was determined by titration with KMnO4 62. Forty milliliter deionized water and 5.0 ml 0.3% hydrogen peroxide solution were added to 2 g air-dried soil sample (milled to 1 mm). The mixture was shaken at 150 rev min−1 for 20 min, followed by adding 5.0 ml H2SO4 (1.5 mol L−1) to end the reaction. The solution was filtered and titrated using 0.02 mol L−1 KMnO4. Catalase activity was finally expressed as 20 min KMnO4 0.02 mol L−1.
Acid phosphatase activity was determined using P-nitrophenyl phosphate disodium as substrates63. One gram moist soil was put into a 50 ml centrifugal tube, and then 0.25 ml toluene, 4.00 ml MUB buffer (pH 6.5) and 15 mmol L−1 P-nitrophenyl phosphate disodium were added. The mixture was incubated at 37 °C for 1 hour. Chemical reaction was ended by adding 4.00 ml CaCl2 (0.5 mol L−1) and 4.00 ml NaOH (0.5 mol L−1). The soil suspension was filtered and the resulting solution was measured at 400 nm using a UV-VIS spectrometer (T6, Beijing Purkingje General Instrument, China). The activity of acid phosphatase was expressed as μg PNP g−1 dry soil h−1.
Dehydrogenase activity was measured by the reduction of triphenyl tetrazolium chloride (TTC) to 2, 3, 5-triphenyl formazan (TPF)64. Ten gram moist soil was throughly mixed with 0.1 g CaCO2, and then 3 g mixture was dispensed to a 50 ml centrifugal tube. Soil sample was added with 0.5 ml 3% TTC and 1.25 ml deionized water, and incubated at 37 °C for 24 hours. Reaction was ended by adding 5.0 ml toluene. Mixture in the tube was centrifuged at 4000 rev min−1 for 5 min to extract TPF. The extraction was then added to 50.0 ml using toluene. The optical density of extracted supernatant was measured at 485 nm on a UV-VIS spectrometer (T6, Beijing Purkingje General Instrument, China). Soil dehydrogenase activity was expressed as μg TPF g−1 dry soil.
To analyze urease activity, approximate 5 g soil was placed into a 50 ml volumetric flask mixed with 1 ml phenol. Fifteen minutes later, 5 ml urea solution (100 g urea L−1) and 5 ml citrate solution (pH 6.7) were added to soil sample which was then incubated at 38 °C for 3 hours. The mixture was then diluted to 50 ml using deionized water (38 °C). The suspension was filtered, and a aliquot (1.0 ml) filtered solution was moved into a 50 ml volumetric flask and diluted to 10 ml with deionized water; 4 ml sodium phenol and 3 ml sodium hypochlorite were then added and mixed; 20 min later, the mixture was diluted to 50 ml with deionized water. The final solution was analyzed on a spectrophotometer (T6, Beijing Purkingje General Instrument, China) at 578 nm within 1 hour. Urease activity was presented as mg NH4 + -N g−1 dry soil h−1 65, 66.
All soil samples were conducted with 3 replicates and enzymatic activities were represented as the average of replicates.
Soil incubation and respiration measurement
Soil incubations followed Chen et al.67. Fresh soil samples, equivalent to 30 g dry weight, were incubated in 300 ml jars under changing temperature. Soil moisture was adjusted to and maintained at 60% of field holding capacity during the whole incubation by adding deionized water every two days to recover weight loss67, 68. All samples were pre-incubated at 25 °C for 48 hours to minimize initial disturbances. Fresh air was continuously passed through the head space of incubation jar at a flow rate of 0.75 L min−1. Incubation temperature was increased from 10 °C to 31 °C with a step length of 3 °C and then backed to the minimum69. Soils were kept at each temperature for 4–10 hours, of which the first 2 hours were allowed for achieving a new equilibrium after temperature changed and the following hours for respiration measurement.
Five milliliter gas was sampled from the head space immediately after incubation jars were closed. Whereafter, a same volume of CO2-free air was immediately injected into jars to balance pressure. After a certain period, depending on CO2 concentration change in the head space, a second gas sample of 5 ml was taken and incubation jar was then opened to allow fresh air flowing through. The actual volume of head space was measured by water replacement at the end of incubation. Soil respiration rate was calculated as CO2 concentration differences between the first and the second gas samples, based on the volume of head space and the period of jar closure.
To measure the temperature sensitivity of soil C mineralization, soil respiration rates were fitted to temperature by an exponential function as below:
where R is soil respiration rate in μmol g−1 dry soil h−1, t is temperature (°C), a and b are fitting parameters, respectively. A reference soil respiration rate, R 20, was defined as the respiration rate at 20 °C.
The temperature sensitivity of soil respiration, measured by Q 10 value, was calculated by:
Soil microbial metabolic quotient, qCO2, was defined as:
Data analysis
All data were analyzed using a software of SPSS for Windows, 21.0 version. The t-test were used to compare biomass between Bt rice and non-Bt rice. The other data were analyzed by one-way factorial analysis of variance. Post hoc pairwise multiple comparisons were conducted using the Duncan tests. To test whether there were changes in each type of soil enzymatic activity, carbon and nutrient pools, a nonmetric multidimensional scaling (NMDS) based on Bray-Curtis distances was performed to graphically visualize the organization of samples in two-dimensional space, using a R multivariate statistical analysis package: Vegan70.
References
Huang, J. K., Hu, R. F., Rozelle, S. & Pray, C. Insect-resistant GM rice in farmers’ fields: Assessing productivity and health effects in China. Science. 308, 688–690 (2005).
Rong, J. et al. Modelling pollen‐mediated gene flow in rice: Risk assessment and management of transgene escape. Plant Biotechnol. J 8, 452–464 (2010).
Gassmann, A. J. Field-evolved resistance to Bt maize by western corn rootworm: predictions from the laboratory and effects in the field. J. Invertebr. Pathol. 110, 287–293 (2012).
Janmaat, A. F. & Myers, J. Rapid evolution and the cost of resistance to Bacillus thuringiensis in greenhouse populations of cabbage loopers, Trichoplusia ni. Proc. R. Soc. London, Ser. A 270, 2263–2270 (2003).
Tabashnik, B. E. et al. Suppressing resistance to Bt cotton with sterile insect releases. Nat. Biotechnol. 28, 1304–1307 (2010).
Bernal, C. C., Aguda, R. M. & Cohen, M. B. Effect of rice lines transformed with Bacillus thuringiensis toxin genes on the brown planthopper and its predator Cyrtorhinus lividipennis. Entomol. Exp. Appl. 102, 21–28 (2002).
Lu, Y., Wu, K., Jiang, Y., Guo, Y. & Desneux, N. Widespread adoption of Bt cotton and insecticide decrease promotes biocontrol services. Nature. 487, 362–365 (2012).
Marvier, M., McCreedy, C., Regetz, J. & Kareiva, P. A meta-analysis of effects of Bt cotton and maize on nontarget invertebrates. Science. 316, 1475–1477 (2007).
Lee, L., Saxena, D. & Stotzky, G. Activity of free and clay-bound insecticidal proteins from Bacillus thuringiensis subsp. israelensis against the mosquito Culex pipiens. Appl. Environ.Microb 69, 4111–4115 (2003).
Stotzky, G. Persistence and biological activity in soil of insecticidal proteins from Bacillus thuringiensis and of bacterial DNA bound on clays and humic acids. J. Environ. Qual 29, 691–705 (2000).
Zhou, X., Huang, Q., Chen, S. & Yu, Z. Adsorption of the insecticidal protein of Bacillus thuringiensis on montmorillonite, kaolinite, silica, goethite and red soil. Appl. Clay Sci. 30, 87–93 (2005).
Sims, S. R. & Ream, J. E. Soil inactivation of the Bacillus thuringiensis subsp kurstaki CryIIA insecticidal protein within transgenic cotton tissue: Laboratory microcosm and field studies. J. Agr. Food Chem. 45, 1502–1505 (1997).
Li, X. & Liu, B. A 2-year field study shows little evidence that the long-term planting of transgenic insect-resistant cotton affects the community structure of soil nematodes. Plos One. 8, doi:10.1371/journal.pone.0061670 (2013).
Liu, Y., Li, J., Stewart, C. N. Jr., Luo, Z. & Xiao, N. The effects of the presence of Bt-transgenic oilseed rape in wild mustard populations on the rhizosphere nematode and microbial communities. Sci. Total Environ. 530, 263–270 (2015).
Saxena, D. & Stotzky, G. Bacillus thuringiensis (Bt) toxin released from root exudates and biomass of Bt corn has no apparent effect on earthworms, nematodes, protozoa, bacteria, and fungi in soil. Soil Biol. Biochem. 33, 1225–1230 (2001).
Devare, M. H., Jones, C. M. & Thies, J. E. Effect of Cry3Bb transgenic corn and tefluthrin on the soil microbial community: Biomass, activity, and diversity. J. Environ. Qual. 33, 837–843 (2004).
Mina, U., Chaudhary, A. & Kamra, A. Effect of Bt cotton on enzymes activity and microorganisms in rhizosphere. J. Agr. Sci. 3, 96–104 (2011).
Shen, R. F., Cai, H. & Gong, W. H. Transgenic Bt cotton has no apparent effect on enzymatic activities or functional diversity of microbial communities in rhizosphere soil. Plant Soil 285, 149–159 (2006).
Wei, M. et al. Impact of Bt-transgenic rice (SHK601) on soil ecosystems in the rhizosphere during crop development. Plant Soil Environ 58, 217–223 (2012).
Al-Kaisi, M. M. & Guzman, J. G. Effects of tillage and nitrogen rate on decomposition of transgenic Bt and near-isogenic non-Bt maize residue. Soil Till. Res. 129, 32–39 (2013).
Sarkar, B., Patra, A. K., Purakayastha, T. & Megharaj, M. Assessment of biological and biochemical indicators in soil under transgenic Bt and non-Bt cotton crop in a sub-tropical environment. Environ. Monit. Assess. 156, 595–604 (2009).
Yang, W., Zhang, M. & Ding, G. Effect of transgenic Bt cotton on bioactivities and nutrients in rhizosphere soil. Commun. Soil Sci. Plan 43, 689–700 (2012).
Yanni, S. F., Whalen, J. K., Simpson, M. J. & Janzen, H. H. Plant lignin and nitrogen contents control carbon dioxide production and nitrogen mineralization in soils incubated with Bt and non-Bt corn residues. Soil Biol. Biochem. 43, 63–69 (2011).
Yang, B., Liu, X., Chen, H. & Ge, F. The specific responses of Acari community to Bt cotton cultivation in agricultural soils in northern China. Appl. Soil Ecol. 66, 1–7 (2013).
Fliessbach, A., Nietlispach, B., Messmer, M., Rodriguez-Romero, A.-S. & Maeder, P. Microbial response of soils with organic and conventional management history to the cultivation of Bacillus thuringiensis (Bt)-maize under climate chamber conditions. Biol. Fert. Soils 49, 829–837 (2013).
Velmourougane, K. et al. Soil dehydrogenase activity in agro-ecological sub regions of black soil regions in India. Geoderma. 197, 186–192 (2013).
Devare, M., Londono-R, L. M. & Thies, J. E. Neither transgenic Bt maize (MON863) nor tefluthrin insecticide adversely affect soil microbial activity or biomass: A 3-year field analysis. Soil Biol. Biochem. 39, 2038–2047 (2007).
Raubuch, M., Behr, K., Roose, K. & Joergensen, R. G. Specific respiration rates, adenylates, and energy budgets of soil microorganisms after addition of transgenic Bt-maize straw. Pedobiologia. 53, 191–196 (2010).
Fang, H. et al. Effect of vegetation of transgenic Bt rice lines and their straw amendment on soil enzymes, respiration, functional diversity and community structure of soil microorganisms under field conditions. J. Environ. Sci-China 24, 1259–1270 (2012).
Singh, R. J., Ahlawat, I. P. S. & Singh, S. Effects of transgenic Bt cotton on soil fertility and biology under field conditions in subtropical inceptisol. Environ. Monit. Assess. 185, 485–495 (2013).
FAO. Statistical yearbook 2013: World food and agriculture. Food and Agriculture Organization of the United Nations, Rome 289 (2013).
Liu, W., Wu, W., Lu, H. H. & Chen, Y. X. Carbon flux from decomposing C-13-labeled transgenic and nontransgenic parental rice straw in paddy soil. J. Soil Sediment 14, 1659–1668 (2014).
Wu, W. X., Ye, Q. F., Min, H., Duan, X. J. & Jin, W. M. Bt-transgenic rice straw affects the culturable microbiota and dehydrogenase and phosphatase activities in a flooded paddy soil. Soil Biol. Biochem. 36, 289–295 (2004).
Liu, W. et al. Transgenic Bt rice does not affect enzyme activities and microbial composition in the rhizosphere during crop development. Soil Biol. Biochem. 40, 475–486 (2008).
Wang, H., Ye, Q., Wang, W., Wu, L. & Wu, W. Cry1Ab protein from Bt transgenic rice does not residue in rhizosphere soil. Environ. Pollut. 143, 449–455 (2006).
Wang, Y. et al. Determination of the movement and persistence of Cry1Ab/1Ac protein released from Bt transgenic rice under field and hydroponic conditions. Soil Biol. Biochem. 58, 107–114 (2013).
Feng, Y. et al. Effects of temperature, water content and pH on degradation of Cry1Ab protein released from Bt corn straw in soil. Soil Biol. Biochem. 43, 1600–1606 (2011).
Gruber, H., Paul, V., Meyer, H. H. & Müller, M. Determination of insecticidal Cry1Ab protein in soil collected in the final growing seasons of a nine-year field trial of Bt-maize MON810. Transgenic Res 21, 77–88 (2012).
Bending, G. D., Turner, M. K. & Jones, J. E. Interactions between crop residue and soil organic matter quality and the functional diversity of soil microbial communities. Soil Biol. Biochem. 34, 1073–1082 (2002).
Hao, X., Indraratne, S. P. & Blackshaw, R. E. Will genetically engineered crop production affect soil carbon? Can. J. Soil Sci. 92, 841–844 (2012).
Wang, F. et al. Effects of N treatments on the yield advantage of Bt-SY63 over SY63 (Oryza sativa) and the concentration of Bt protein. Field Crop. Res. 129, 39–45 (2012).
Liu, W., Wang, S., Chen, Y. & Wu, W. Microbial diversity in rhizosphere soil of transgenic Bt rice based on the characterization of phospholipids fatty acids (in chinese). Chinese J. Appl. Ecol. 22, 727–733 (2011).
Saxena, D. & Stotzky, G. Bt corn has a higher lignin content than non-Bt corn. Am. J. Bot. 88, 1704–1706 (2001).
Karuri, H., Amata, R., Amugune, N. & Waturu, C. Nematode diversity in soil from a field trial with decomposing Bt cotton expressing Cry1Ac and Cry2Ab2 protein. Span. J. Agric. Res. 11, 968–979 (2013).
Tarkalson, D. D., Kachman, S. D., Knops, J. M., Thies, J. E. & Wortmann, C. S. Decomposition of Bt and non-Bt corn hybrid residues in the field. Nutr. Cycl. Agroecosys 80, 211–222 (2008).
Wassmann, R. & Aulakh, M. S. The role of rice plants in regulating mechanisms of methane missions. Biol. Fert. Soils 31, 20–29 (2000).
Dennis, P. G., Miller, A. J. & Hirsch, P. R. Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbiol. Ecol 72, 313–327 (2010).
Kuzyakov, Y. & Domanski, G. Carbon input by plants into the soil- review. J. Soil Sci. Plant Nut 163, 421–431 (2000).
Vranova, V., Rejsek, K., Skene, K. R., Janous, D. & Formanek, P. Methods of collection of plant root exudates in relation to plant metabolism and purpose: a review. J. Plant Nutr. Soil Sc 176, 175–199 (2013).
Aulakh, M., Wassmann, R., Bueno, C., Kreuzwieser, J. & Rennenberg, H. Characterization of root exudates at different growth stages of ten rice (Oryza sativa L.) cultivars. Plant Biology 3, 139–148 (2001).
Bhattacharyya, P., Das, S. & Adhya, T. Root exudates of rice cultivars affect rhizospheric phosphorus dynamics in soils with different phosphorus statuses. Commun. Soil Sci. Plan 44, 1643–1658 (2013).
Tawaraya, K. et al. Metabolite profiling of shoot extracts, root extracts, and root exudates of rice plant under phosphorus deficiency. J. Plant Nutr. 36, 1138–1159 (2013).
Velmourougane, K. & Sahu, A. Impact of transgenic cottons expressing Cry1Ac on soil biological attributes. Plant Soil Environ 59, 108–114 (2013).
Londono-R, L. M., Tarkalson, D. & Thies, J. E. In-field rates of decomposition and microbial communities colonizing residues vary by depth of residue placement and plant part, but not by crop genotype for residues from two Cry1Ab Bt corn hybrids and their non-transgenic isolines. Soil Biol. Biochem. 57, 349–355 (2013).
Tarafdar, J. C. & Shiva, V. Effect of Bt-transgenic cotton on soil biological health. Appl. Biol. Res. 14, 15–23 (2012).
Kravchenko, A. N., Hao, X. & Robertson, G. P. Seven years of continuously planted Bt corn did not affect mineralizable and total soil C and total N in surface soil. Plant Soil 318, 269–274 (2009).
Boerner, R. E. J., Brinkman, J. A. & Smith, A. Seasonal variations in enzyme activity and organic carbon in soil of a burned and unburned hardwood forest. Soil Biol. Biochem. 37, 1419–1426 (2005).
Wallenstein, M. D., McMahon, S. K. & Schimel, J. P. Seasonal variation in enzyme activities and temperature sensitivities in Arctic tundra soils. Global Change Biol 15, 1631–1639 (2009).
Olsen, S. R. & Sommers, L. E. Phosphorus. In D. Buxton, ed., Method of Soil Analysis: Chemical and Microbiological Properties (second ed.), Soil Science Society of American, Madison, Wisconsin, 403–427(1982).
Xiao, M. Q. et al. Effects of water management practices on residue decomposition and degradation of Cry1Ac protein from crop-wild Bt rice hybrids and parental lines during winter fallow season. Ecotox. Environ. Safe 122, 275–289 (2015).
Wu, J., Joergensen, R., Pommerening, B., Chaussod, R. & Brookes, P. Measurement of soil microbial biomass C by fumigation-extraction—an automated procedure. Soil Biol. Biochem. 22, 1167–1169 (1990).
Johnson, J. L. & Temple, K. L. Some variables affecting the measurement of “catalase activity” in soil. Soil Sci. Soc. Am. J 28, 207–209 (1964).
Tabatabai, M. & Bremner, J. Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1, 301–307 (1969).
Casida, L. Jr., Klein, D. & Santoro, T. Soil dehydrogenase activity. Soil Sci. 98, 371–376 (1964).
Kizilkaya, R. & Dengiz, O. Variation of land use and land cover effects on some soil physico-chemical characteristics and soil enzyme activity. Zemdirbyste 97, 15–24 (2010).
Zhang, C., Liu, G., Xue, S. & Song, Z. Rhizosphere soil microbial activity under different vegetation types on the Loess Plateau, China. Geoderma. 161, 115–125 (2011).
Chen, X. et al. Evaluating the impacts of incubation procedures on estimated Q 10 values of soil respiration. Soil Biol. Biochem. 42, 2282–2288 (2010).
Singh, J., Saggar, S., Giltrap, D. & Bolan, N. S. Decomposition of dicyandiamide (DCD) in three contrasting soils and its effect on nitrous oxide emission, soil respiratory activity, and microbial biomass—an incubation study. Soil Res 46, 517–525 (2008).
Fang, C. & Moncrieff, J. The dependence of soil CO2 efflux on temperature. Soil Biol. Biochem. 33, 155–165 (2001).
Oksanen, J. et al. Package ‘vegan’. R packag ver 254, 20–28 (2013).
Acknowledgements
This study was supported by the Jiangsu Provincial Natural Science Foundation of China (No. BK20130421), the Natural Science Foundation of the Jiangsu Higher Education Institutions of China (No. 13KJB180026), the National Natural Science Foundation of China (No. 31300443). The Key Project for Breeding Genetically Modified Organisms (2009 ZX08011-012B) and the National Natural Science Foundation of China, China (31170505).
Author information
Authors and Affiliations
Contributions
L.Z.L. performed experiments, data analysis and wrote the manuscript. F.C.M., with assistance from S.Z.P. and W.F., was in charge of experiment design and project management. X.M.Q. helped in taking samples in fields. B.N.S., C.J., and C.X.P. helped to measure soil arguments. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhaolei, L., Naishun, B., Jun, C. et al. Effects of long-term cultivation of transgenic Bt rice (Kefeng-6) on soil microbial functioning and C cycling. Sci Rep 7, 4647 (2017). https://doi.org/10.1038/s41598-017-04997-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04997-8
This article is cited by
-
Soil nutrient cycling and microbiome responses to Bt rice cultivation
Plant and Soil (2024)
-
Combined metagenomic and metabolomic analyses reveal that Bt rice planting alters soil C-N metabolism
ISME Communications (2023)
-
Impact of transgenic birch with modified nitrogen metabolism on soil properties, microbial biomass and enzymes in 4-year study
Plant and Soil (2023)
-
An insight to rhizosphere bacterial community composition and structure of consecutive winter-initiated sugarcane ratoon crop in Southern China
BMC Plant Biology (2022)
-
Evaluating the effects of transgenic Bt rice cultivation on soil stability
Environmental Science and Pollution Research (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.