Abstract
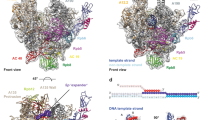
RNA polymerase III (Pol III) synthesizes structured, essential small RNAs, such as transfer RNA, 5S ribosomal RNA and U6 small nuclear RNA. Pol III, the largest nuclear RNA polymerase, is composed of a conserved core region and eight constitutive regulatory subunits, but how these factors jointly regulate Pol III transcription remains unclear. Here, we present cryo-EM structures of human Pol III in both apo and elongating states, which unveil both an orchestrated movement during the apo-to-elongating transition and an unexpected apo state in which the RPC7 subunit tail occupies the DNA–RNA-binding cleft of Pol III, suggesting that RPC7 plays important roles in both autoinhibition and transcription initiation. The structures also reveal a proofreading mechanism for the TFIIS-like subunit RPC10, which stably retains its catalytic position in the secondary channel, explaining the high fidelity of Pol III transcription. Our work provides an integrated picture of the mechanism of Pol III transcription regulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM 3D maps of human Pol III in the apo form and in complex with the scaffold have been deposited in the EMDB database with accession codes EMD-30578 and EMD-30577, respectively. The atomic models of human Pol III in the apo form and in complex with the scaffold have been deposited in the PDB database with accession codes 7D59 and 7D58, respectively. Source data are provided with this paper.
References
Roeder, R. G. & Rutter, W. J. Multiple forms of DNA-dependent RNA polymerase in eukaryotic organisms. Nature 224, 234–237 (1969).
Dieci, G., Fiorino, G., Castelnuovo, M., Teichmann, M. & Pagano, A. The expanding RNA polymerase III transcriptome. Trends Genet. 23, 614–622 (2007).
Roeder, R. G. 50+ years of eukaryotic transcription: an expanding universe of factors and mechanisms. Nat. Struct. Mol. Biol. 26, 783–791 (2019).
Turowski, T. W. & Tollervey, D. Transcription by RNA polymerase III: insights into mechanism and regulation. Biochem. Soc. Trans. 44, 1367–1375 (2016).
Moir, R. D. & Willis, I. M. Regulation of Pol III transcription by nutrient and stress signaling pathways. Biochim. Biophys. Acta 1829, 361–375 (2013).
Arimbasseri, A. G. & Maraia, R. J. RNA polymerase III advances: structural and tRNA functional views. Trends Biochem. Sci. 41, 546–559 (2016).
White, R. J. RNA polymerases I and III, non-coding RNAs and cancer. Trends Genet. 24, 622–629 (2008).
Dauwerse, J. G. et al. Mutations in genes encoding subunits of RNA polymerases I and III cause Treacher Collins syndrome. Nat. Genet. 43, 20–22 (2011).
Yeganeh, M. & Hernandez, N. RNA polymerase III transcription as a disease factor. Genes Dev. 34, 865–882 (2020).
Vannini, A. & Cramer, P. Conservation between the RNA polymerase I, II and III transcription initiation machineries. Mol. Cell 45, 439–446 (2012).
Arimbasseri, A. G. & Maraia, R. J. Mechanism of transcription termination by RNA polymerase III utilizes a non-template strand sequence-specific signal element. Mol. Cell 58, 1124–1132 (2015).
Dumay-Odelot, H., Durrieu-Gaillard, S., El Ayoubi, L., Parrot, C. & Teichmann, M. Contributions of in vitro transcription to the understanding of human RNA polymerase III transcription. Transcription 5, e27526 (2014).
Kornberg, R. D. The molecular basis of eucaryotic transcription. Cell Death Differ. 14, 1989–1997 (2007).
Murakami, K. et al. Architecture of an RNA polymerase II transcription pre-initiation complex. Science 342, 1238724 (2013).
Engel, C., Neyer, S. & Cramer, P. Distinct mechanisms of transcription initiation by RNA polymerases I and II. Annu. Rev. Biophys. 47, 425–446 (2018).
Cramer, P. Organization and regulation of gene transcription. Nature 573, 45–54 (2019).
Carter, R. & Drouin, G. The increase in the number of subunits in eukaryotic RNA polymerase III relative to RNA polymerase II is due to the permanent recruitment of general transcription factors. Mol. Biol. Evol. 27, 1035–1043 (2010).
Khatter, H., Vorlander, M. K. & Muller, C. W. RNA polymerase I and III: similar yet unique. Curr. Opin. Struct. Biol. 47, 88–94 (2017).
Kassavetis, G. A., Prakash, P. & Shim, E. The C53/C37 subcomplex of RNA polymerase III lies near the active site and participates in promoter opening. J. Biol. Chem. 285, 2695–2706 (2010).
Landrieux, E. et al. A subcomplex of RNA polymerase III subunits involved in transcription termination and reinitiation. EMBO J. 25, 118–128 (2006).
Brun, I., Sentenac, A. & Werner, M. Dual role of the C34 subunit of RNA polymerase III in transcription initiation. EMBO J. 16, 5730–5741 (1997).
Kenneth, N. S., Marshall, L. & White, R. J. Recruitment of RNA polymerase III in vivo. Nucleic Acids Res. 36, 3757–3764 (2008).
Lefevre, S. et al. Structure–function analysis of hRPC62 provides insights into RNA polymerase III transcription initiation. Nat. Struct. Mol. Biol. 18, 352–358 (2011).
Ayoubi, L. E. et al. The hRPC62 subunit of human RNA polymerase III displays helicase activity. Nucleic Acids Res. 47, 10313–10326 (2019).
Hoffmann, N. A. et al. Molecular structures of unbound and transcribing RNA polymerase III. Nature 528, 231–236 (2015).
Abascal-Palacios, G., Ramsay, E. P., Beuron, F., Morris, E. & Vannini, A. Structural basis of RNA polymerase III transcription initiation. Nature 553, 301–306 (2018).
Vorlander, M. K., Khatter, H., Wetzel, R., Hagen, W. J. H. & Muller, C. W. Molecular mechanism of promoter opening by RNA polymerase III. Nature 553, 295–300 (2018).
Han, Y., Yan, C., Fishbain, S., Ivanov, I. & He, Y. Structural visualization of RNA polymerase III transcription machineries. Cell Discov. 4, 40 (2018).
Thuillier, V., Stettler, S., Sentenac, A., Thuriaux, P. & Werner, M. A mutation in the C31 subunit of Saccharomyces cerevisiae RNA polymerase III affects transcription initiation. EMBO J. 14, 351–359 (1995).
Bartholomew, B., Durkovich, D., Kassavetis, G. A. & Geiduschek, E. P. Orientation and topography of RNA polymerase III in transcription complexes. Mol. Cell. Biol. 13, 942–952 (1993).
Gamba, P. & Zenkin, N. Transcription fidelity and its roles in the cell. Curr. Opin. Microbiol. 42, 13–18 (2018).
Alic, N. et al. Selectivity and proofreading both contribute significantly to the fidelity of RNA polymerase III transcription. Proc. Natl Acad. Sci. USA 104, 10400–10405 (2007).
Ruan, W., Lehmann, E., Thomm, M., Kostrewa, D. & Cramer, P. Evolution of two modes of intrinsic RNA polymerase transcript cleavage. J. Biol. Chem. 286, 18701–18707 (2011).
Chedin, S., Riva, M., Schultz, P., Sentenac, A. & Carles, C. The RNA cleavage activity of RNA polymerase III is mediated by an essential TFIIS-like subunit and is important for transcription termination. Genes Dev. 12, 3857–3871 (1998).
Vassylyev, D. G., Vassylyeva, M. N., Perederina, A., Tahirov, T. H. & Artsimovitch, I. Structural basis for transcription elongation by bacterial RNA polymerase. Nature 448, 157–162 (2007).
Neyer, S. et al. Structure of RNA polymerase I transcribing ribosomal DNA genes. Nature 540, 607–610 (2016).
Bernecky, C., Herzog, F., Baumeister, W., Plitzko, J. M. & Cramer, P. Structure of transcribing mammalian RNA polymerase II. Nature 529, 551–554 (2016).
Blombach, F. et al. Archaeal TFEα/β is a hybrid of TFIIE and the RNA polymerase III subcomplex hRPC62/39. Elife 4, e08378 (2015).
Hu, P. et al. Characterization of human RNA polymerase III identifies orthologues for Saccharomyces cerevisiae RNA polymerase III subunits. Mol. Cell. Biol. 22, 8044–8055 (2002).
Werner, F. & Grohmann, D. Evolution of multisubunit RNA polymerases in the three domains of life. Nat. Rev. Microbiol. 9, 85–98 (2011).
Tafur, L. et al. Molecular structures of transcribing RNA polymerase I. Mol. Cell 64, 1135–1143 (2016).
Fernandez-Tornero, C. et al. Crystal structure of the 14-subunit RNA polymerase I. Nature 502, 644–649 (2013).
Wang, D. et al. Structural basis of transcription: backtracked RNA polymerase II at 3.4-Å resolution. Science 324, 1203–1206 (2009).
Cheung, A. C. & Cramer, P. Structural basis of RNA polymerase II backtracking, arrest and reactivation. Nature 471, 249–253 (2011).
Tafur, L. et al. The cryo-EM structure of a 12-subunit variant of RNA polymerase I reveals dissociation of the A49-A34.5 heterodimer and rearrangement of subunit A12.2. Elife 8, e43204 (2019).
Nielsen, S., Yuzenkova, Y. & Zenkin, N. Mechanism of eukaryotic RNA polymerase III transcription termination. Science 340, 1577–1580 (2013).
Mishra, S. & Maraia, R. J. RNA polymerase III subunits C37/53 modulate rU:dA hybrid 3′ end dynamics during transcription termination. Nucleic Acids Res. 47, 310–327 (2019).
Iben, J. R. et al. Point mutations in the Rpb9-homologous domain of Rpc11 that impair transcription termination by RNA polymerase III. Nucleic Acids Res. 39, 6100–6113 (2011).
Willis, I. M. & Moir, R. D. Signaling to and from the RNA polymerase III transcription and processing machinery. Annu. Rev. Biochem. 87, 75–100 (2018).
Vorländer, M. K. et al. Structural basis for RNA polymerase III transcription repression by Maf1. Nat. Struct. Mol. Biol. 27, 229–232 (2020).
Teichmann, M., Wang, Z. & Roeder, R. G. A stable complex of a novel transcription factor IIB- related factor, human TFIIIB50, and associated proteins mediate selective transcription by RNA polymerase III of genes with upstream promoter elements. Proc. Natl Acad. Sci. USA 97, 14200–14205 (2000).
Schramm, L., Pendergrast, P. S., Sun, Y. & Hernandez, N. Different human TFIIIB activities direct RNA polymerase III transcription from TATA-containing and TATA-less promoters. Genes Dev. 14, 2650–2663 (2000).
Schramm, L. & Hernandez, N. Recruitment of RNA polymerase III to its target promoters. Genes Dev. 16, 2593–2620 (2002).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
The PyMOL Molecular Graphics System v2.4 (Schrödinger, 2002).
Acknowledgements
We thank the staff members of the Electron Microscopy Facility and Mass Spectrometry Facility at the Shanghai Institute of Precision Medicine for providing technical support and assistance with data collection. This work was supported by grants from the National Key R&D Program of China (2018YFA0107000 to M.L.), the National Natural Science Foundation of China (31525007 and 31930063 to M.L., 31900929 and U1932114 to P.L., 31900855 to Q.W., 81902085 to Y.X.), the Shanghai Municipal Education Commission Gaofeng Clinical Medicine Grant Support (20181711 to J.W.), the Shanghai Pujiang Program (19PJ1407200 to Q.W.), the Innovative Research Team of High-level Local University in Shanghai (SSMU-ZDCX20180702 to Q.W., SSMU-ZDCX20180401 to P.L.) and the Fund of Excellent Young Scholars of Shanghai Ninth People’s Hospital, Shanghai JiaoTong University School of Medicine (to P.L.).
Author information
Authors and Affiliations
Contributions
Q.W. and F.W. purified the human Pol III complex. Q.W., F.W., M.C. and S.L. prepared cryo-EM specimens, collected datasets and determined the structure. J.W. and Y.X. carried out model building and refinement. Z.W. and F.W. carried out yeast functional analysis. All authors contributed to data interpretation and the writing of the manuscript. M.L., Q.W. and P.L. wrote the manuscript. M.L., Q.W. and J.W. initiated and orchestrated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Extended data
Extended Data Fig. 1 Domain organization of all 17 subunits of human Pol III.
Black lines above the domain bar denote the modeled regions with clear EM densities.
Extended Data Fig. 2 Purification and characterization of human Pol III.
a, SDS-PAGE analysis of the purified human Pol III by silver staining. b, A representative negative−staining image of human Pol III. c, 2D class average views of the negative-staining images of human Pol III. Uncropped image for panel a is available as source data.
Extended Data Fig. 3 Cryo-EM analysis of human Pol III EC.
a, Left: A representative raw cryo-EM image of human Pol III EC. Right: 2D class average views of human Pol III EC. b, Image processing flowchart for human Pol III EC. c, Local resolution map for human Pol III EC. d, Angular distribution of particles for final reconstruction of human Pol III EC. e, Fourier Shell Correlation curve of constructed map of human Pol III EC.
Extended Data Fig. 4 Cryo-EM density maps of human Pol III EC and the active center.
a, Cryo-EM density map of human Pol III EC is shown in two different views. Protein subunits and the DNA-RNA scaffold are color coded as in Fig. 1b. b, Representative cryo-EM density maps for the selected elements of the human Pol III protein subunits. c, Close-up views of the active center of bacterial RNA polymerase (PDB: 2o5i), bovine Pol II (PDB: 5flm), yeast Pol I (PDB: 5m3f) and human Pol III. The lid loop, the fork loop 1 and the rudder are colored in yellow, magenta and blue, respectively.
Extended Data Fig. 5 Cryo-EM analysis of apo human Pol III.
a, Left: A representative raw cryo-EM image of the apo human Pol III. Right: 2D class average views of apo human Pol III. b, Image processing flowchart for apo human Pol III. The data processing details are described in the method session. c, Local resolution map for apo human Pol III. d, Angular distribution of particles for final reconstruction of apo human Pol III. e, Fourier Shell Correlation curve of constructed map of apo human Pol III.
Extended Data Fig. 6 Cryo-EM density maps of apo human Pol III.
a, Cryo-EM density map of the apo human Pol III complex is displayed in two different views. Protein subunits are color coded as in Fig. 1b. b, Cryo-EM density map of human Pol III in transcribing state with endogenous DNA in the catalytic cleft. The Pol III EC structure was fit into the density map (semi-transparent). The density corresponding to the double-stranded DNA is highlighted in yellow.
Extended Data Fig. 7 Structural comparison of the yeast and human RPC6 C-terminus and the location of WH domains 1 and 2 of RPC5 in human Pol III.
a, Multiple sequence alignment of the C-terminus of RPC6 from representative species reveals that yeast Pol III lacks the [4Fe-4S]-binding motif (Homo sapiens; Pan troglodytes; Mus musculus; Rattus norvegicus; Sus scrofa; Bos taurus; Xenopus laevis; Danio rerio and Saccharomyces cerevisiae). Four cystine residues in the [4Fe-4S]-binding motif are highlighted by red magenta arrow heads. b, Close-up comparison of the C-terminus of C34 and RPC6 in human (left) and yeast (right) Pol III structures, respectively. c, Left: Cryo-EM density map of apo Pol III with focused mask refinement for RPC5 WH domains 1 and 2. Right: Pol III EC at a lower threshold for visualization of RPC5 WH domains 1 and 2.
Extended Data Fig. 8 Multiple sequence alignment RPC7, BRF1, RPC10 and RPC1.
a, Multiple sequence alignment of the stalk-interaction motif of RPC7 from representative species. b, Multiple sequence alignment of RPC7CT. Two observed regions of RPC7CT in the EM density map are highlighted in red and magentas boxes, respectively. Two regions of RPC7CT analyzed by yeast tetrad analysis are highlighted in the blue box and cyan arrowheads, respectively. c, Multiple sequence alignment of the N-terminal Zn-ribbon motif of BRF1. Amino acids involved in the contact with RPC7CT are highlighted in blue box. d, Multiple sequence alignment of the anchor region of RPC10. RPC10 Leu53 is highlighted by a gold arrowhead. e, Multiple sequence alignment of the RPC10-anchor-interacting region of RPC1. Amino acids involved in the hydrophobic interactions with the RPC10 anchor around RPC10Leu35 are highlighted in green arrowheads.
Extended Data Fig. 9 Superposition analysis of the C-Zn-ribbon motif of RPC10 in human Pol III and TFIIS in yeast backtracked Pol II.
a, Superposition of the C-Zn-ribbon motif of RPC10 in human Pol III elongation complex (orange) and TFIIS in yeast backtracked Pol II structures containing mismatched DNA-RNA hybrid (3po3 and 3gtm in cyan and gray, respectively). b, Top: Close-up view of the C-Zn-ribbon motif of RPC10 modeled onto yeast Pol II backtracked DNA-RNA hybrid (3gtm). Bottom: Close-up view of the C-Zn-ribbon motif of TFIIS in yeast backtracked Pol II (3gtm).
Extended Data Fig. 10 Superimposition of yeast Maf1 with human Pol III.
Structure of yeast Maf1 (PDB:6tut) modeled onto the apo structure of human Pol III in two views reveals that MAF1 binding is not compatible with the autoinhibited state of Pol III.
Supplementary information
Supplementary Information
Supplementary Table 1.
Supplementary Video 1
A movie of human Pol III with structural information of the RPC7 C terminus, the [4Fe–4S], the transition between the apo and EC states of Pol III, as well as the constant surveillance factor RPC10.
Source data
Source Data Fig. 2
Unprocessed images of yeast growth on plates.
Source Data Fig. 3
Unprocessed images of yeast growth on plates.
Source Data Extended Data Fig. 2
Unprocessed SDS–PAGE gel.
Rights and permissions
About this article
Cite this article
Wang, Q., Li, S., Wan, F. et al. Structural insights into transcriptional regulation of human RNA polymerase III. Nat Struct Mol Biol 28, 220–227 (2021). https://doi.org/10.1038/s41594-021-00557-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-021-00557-x
This article is cited by
-
Structural basis of Ty1 integrase tethering to RNA polymerase III for targeted retrotransposon integration
Nature Communications (2023)
-
Regulation of ribosomal RNA gene copy number, transcription and nucleolus organization in eukaryotes
Nature Reviews Molecular Cell Biology (2023)
-
Structures of transcription preinitiation complex engaged with the +1 nucleosome
Nature Structural & Molecular Biology (2023)
-
A cancer-associated RNA polymerase III identity drives robust transcription and expression of snaR-A noncoding RNA
Nature Communications (2022)
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases
Nature Reviews Molecular Cell Biology (2022)