Abstract
Protein lysine 2-hydroxyisobutyrylation (Khib) has recently been shown to play a critical role in the regulation of cellular processes. However, the mechanism and functional consequence of Khib in prokaryotes remain unclear. Here we report that TmcA, an RNA acetyltransferase, functions as a lysine 2-hydroxyisobutyryltransferase in the regulation of transcription. We show that TmcA can effectively catalyze Khib both in vitro and intracellularly, and that R502 is a key site for the Khib catalytic activity of TmcA. Using quantitative proteomics, we identified 467 endogenous candidates targeted by TmcA for Khib in Escherichia coli. Interestingly, we demonstrate that TmcA can specifically modulate the DNA-binding activity of H-NS, a nucleoid-associated protein, by catalysis of Khib at K121. Furthermore, this TmcA-targeted Khib regulates transcription of acid-resistance genes and enhances E. coli survival under acid stress. Our study reveals transcription regulation mediated by TmcA-catalyzed Khib for bacterial acid resistance.

This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
All data required to evaluate the conclusions are present in the paper and in Supplementary information. The proteomics MS data have been deposited with the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository, with the dataset identifiers PXD024897 and PXD024901. RNA-seq data can be obtained at the Gene Expression Omnibus database under accession no. GSE169499. Source data are provided with this paper.
References
Park, J. et al. SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol. Cell 50, 919–930 (2013).
Rardin, M. J. et al. SIRT5 regulates the mitochondrial lysine succinylome and metabolic networks. Cell Metab. 18, 920–933 (2013).
Choudhary, C., Weinert, B. T., Nishida, Y., Verdin, E. & Mann, M. The growing landscape of lysine acetylation links metabolism and cell signalling. Nat. Rev. Mol. Cell Biol. 15, 536–550 (2014).
Zhao, S. et al. Regulation of cellular metabolism by protein lysine acetylation. Science 327, 1000–1004 (2010).
Harmel, R. & Fiedler, D. Features and regulation of non-enzymatic post-translational modifications. Nat. Chem. Biol. 14, 244–252 (2018).
Sreelatha, A. et al. Protein AMPylation by an evolutionarily conserved pseudokinase. Cell 175, 809–821 (2018).
Chambers, K. A. & Scheck, R. A. Bacterial virulence mediated by orthogonal post-translational modification. Nat. Chem. Biol. 16, 1043–1051 (2020).
Sabari, B. R., Zhang, D., Allis, C. D. & Zhao, Y. Metabolic regulation of gene expression through histone acylations. Nat. Rev. Mol. Cell Biol. 18, 90–101 (2017).
Wagner, E. J. & Carpenter, P. B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 13, 115–126 (2012).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Semer, M. et al. DNA repair complex licenses acetylation of H2AZ1 by KAT2A during transcription. Nat. Chem. Biol. 15, 992–1000 (2019).
Browning, D. F., Grainger, D. C. & Busby, S. J. Effects of nucleoid-associated proteins on bacterial chromosome structure and gene expression. Curr. Opin. Microbiol. 13, 773–780 (2010).
Dillon, S. C. & Dorman, C. J. Bacterial nucleoid-associated proteins, nucleoid structure and gene expression. Nat. Rev. Microbiol. 8, 185–195 (2010).
Sakatos, A. et al. Posttranslational modification of a histone-like protein regulates phenotypic resistance to isoniazid in mycobacteria. Sci. Adv. 4, eaao1478 (2018).
Carabetta, V. J., Greco, T. M., Cristea, I. M. & Dubnau, D. YfmK is an N(ε)-lysine acetyltransferase that directly acetylates the histone-like protein HBsu in Bacillus subtilis. Proc. Natl Acad. Sci. USA 116, 3752–3757 (2019).
Dai, L. et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nat. Chem. Biol. 10, 365–370 (2014).
Huang, H. et al. p300-Mediated lysine 2-hydroxyisobutyrylation regulates glycolysis. Mol. Cell 70, 663–678 (2018).
Huang, H. et al. Landscape of the regulatory elements for lysine 2-hydroxyisobutyrylation pathway. Cell Res. 28, 111–125 (2018).
Huang, J. et al. 2-Hydroxyisobutyrylation on histone H4K8 is regulated by glucose homeostasis in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 114, 8782–8787 (2017).
Dong, H. et al. Systematic identification of lysine 2-hydroxyisobutyrylated proteins in Proteus mirabilis. Mol. Cell. Proteomics 17, 482–494 (2018).
Dong, H. et al. Protein lysine de-2-hydroxyisobutyrylation by CobB in prokaryotes. Sci. Adv. 5, eaaw6703 (2019).
Sabari, B. R. et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol. Cell 58, 203–215 (2015).
Dancy, B. M. & Cole, P. A. Protein lysine acetylation by p300/CBP. Chem. Rev. 115, 2419–2452 (2015).
Christensen, D. G. et al. Mechanisms, detection, and relevance of protein acetylation in prokaryotes. mBio 10, e02708-18 (2019).
Christensen, D. G. et al. Identification of novel protein lysine acetyltransferases in Escherichia coli. mBio 9, e01905-18 (2018).
Ikeuchi, Y., Kitahara, K. & Suzuki, T. The RNA acetyltransferase driven by ATP hydrolysis synthesizes N4-acetylcytidine of tRNA anticodon. EMBO J. 27, 2194–2203 (2008).
Chimnaronk, S. et al. RNA helicase module in an acetyltransferase that modifies a specific tRNA anticodon. EMBO J. 28, 1362–1373 (2009).
Dorman, C. J. H-NS: a universal regulator for a dynamic genome. Nat. Rev. Microbiol. 2, 391–400 (2004).
Tendeng, C. & Bertin, P. N. H-NS in Gram-negative bacteria: a family of multifaceted proteins. Trends Microbiol. 11, 511–518 (2003).
Weinert, B. T. et al. Lysine succinylation is a frequently occurring modification in prokaryotes and eukaryotes and extensively overlaps with acetylation. Cell Rep. 4, 842–851 (2013).
Weinert, B. T. et al. Acetyl-phosphate is a critical determinant of lysine acetylation in E. coli. Mol. Cell 51, 265–272 (2013).
Dilweg, I. W. & Dame, R. T. Post-translational modification of nucleoid-associated proteins: an extra layer of functional modulation in bacteria? Biochem. Soc. Trans. 46, 1381–1392 (2018).
Hommais, F. et al. Large-scale monitoring of pleiotropic regulation of gene expression by the prokaryotic nucleoid-associated protein, H-NS. Mol. Microbiol. 40, 20–36 (2001).
Gao, X. et al. Engineered global regulator H-NS improves the acid tolerance of E. coli. Microb. Cell Fact. 17, 118 (2018).
Krin, E., Danchin, A. & Soutourina, O. Decrypting the H-NS-dependent regulatory cascade of acid stress resistance in Escherichia coli. BMC Microbiol. 10, 273 (2010).
Wang, W., Li, G. W., Chen, C., Xie, X. S. & Zhuang, X. Chromosome organization by a nucleoid-associated protein in live bacteria. Science 333, 1445–1449 (2011).
Dame, R. T., Rashid, F. Z. M. & Grainger, D. C. Chromosome organization in bacteria: mechanistic insights into genome structure and function. Nat. Rev. Genet. 21, 227–242 (2020).
Zhao, S., Zhang, X. & Li, H. Beyond histone acetylation-writing and erasing histone acylations. Curr. Opin. Struct. Biol. 53, 169–177 (2018).
Kaczmarska, Z. et al. Structure of p300 in complex with acyl-CoA variants. Nat. Chem. Biol. 13, 21–29 (2017).
Liu, X. et al. NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2. EMBO Rep. 17, 349–366 (2016).
Ito, S. et al. Human NAT10 is an ATP-dependent RNA acetyltransferase responsible for N4-acetylcytidine formation in 18 S ribosomal RNA (rRNA). J. Biol. Chem. 289, 35724–35730 (2014).
Dame, R. T., Noom, M. C. & Wuite, G. J. Bacterial chromatin organization by H-NS protein unravelled using dual DNA manipulation. Nature 444, 387–390 (2006).
Krulwich, T. A., Sachs, G. & Padan, E. Molecular aspects of bacterial pH sensing and homeostasis. Nat. Rev. Microbiol. 9, 330–343 (2011).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
This work was supported by funding from the National Natural Science Foundation of China to K.Z. (nos. 21874100 and 22074103), to H.D. (no. 32101023), to G.Z. (no. 21904097) and to X.B. (no. 22004091); and from the Talent Excellence Program from Tianjin Medical University, to K.Z.
Author information
Authors and Affiliations
Contributions
K.Z. supervised experiments. H.D. and K.Z. designed experiments, analyzed the data and wrote the manuscript. H.D., J.Z. and Y.Z. carried out cell culture, enzymatic activity assay and molecular biological experiments. H.D., G.Z. and X.B. carried out proteomic survey. C.B. and H.D. carried out chemical synthesis. H.D., Y.H., S.T., D.H., L.X. and K.Z. carried out data collection, analysis and interpretation. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification TmcA as potential lysine 2-hyroxyisobutyryltransferase, not a lysine acetyltransferase.
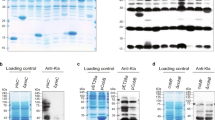
a, Overexpression of TmcA and YiaC increase Khib in E. coli. E. coli BL21 were transferred with empty vector (pET28a) as control and GNAT-pET28a vectors for GNAT overexpressing strains, the Khib level of whole cell lysates were analyzed by western blotting. b, Overexpression of TmcA has no obvious effect on Kac in E. coli. E. coli BL21 were transferred with empty vector (pET28a) as control and TmcA-pET28a vector overexpressing strains, the Kac level of whole cell lysates were analyzed by western blotting. c, TmcA KO has no obvious effect on Kac in E. coli. Kac levels of TmcA KO E. coli MG1655 were analyzed by western blotting, WT as control. And each group had three biological repetitions. All western experiments had three biological repetitions with similar result.
Extended Data Fig. 2 TmcA can bind with 2-hydroxyisobutyryl-CoA.
a, Structural modeling of TmcA combining acetyl-CoA. Exhibition the crystal structure of TmcA combining acetyl-CoA by UCSF Chimera. Ac-CoA: acetyl-CoA. Related to Fig. 2a. b, Structural modeling of TmcA combining 2-hydroxyisobutyryl-CoA. Using Autodock modes the binding of 2-hydroxyisobutyryl-CoA with TmcA and exhibits the crystal structure by UCSF Chimera. Hib-CoA: 2-hydroxyisobutyryl-CoA. c, Structural modeling of TmcA combining Hib-CoA under different setting of Box. Small box:100 Å × 60 Å × 60 Å, large box:120 Å × 80 Å × 80 Å. d, The hydrogen bonds and hydrophobic interaction between TmcA and Hib-CoA are shown by LigPlot+ with large box. e-f, ITC analysis of the affinity of recombinant W504A TmcA mutant with Ac-CoA and Hib-CoA separately, and each group had three biological repetitions, each group had three biological repetitions with similar result.
Extended Data Fig. 3 LC-MS/MS detection ac4C intracellularly and in vitro.
a, ac4C levels of wild type E.coli (WT), TmcA KO (KO), complementary TmcA (TmcA) and complementary TmcA R502 (TmcA R502) strains were detected by LC-MS/MS, each with three biological repetitions (data are presented as means ± SEM). b, The synthetic tRNAMet incubated with TmcA or TmcA R502 and detected the ac4C levels by LC-MS/MS. Each group had three biological repetitions with similar result.
Extended Data Fig. 4 Profiling the the endogenous substrate proteins for acetylation by TmcA in E. coli.
a, Schematic representation of experimental workflow for the SILAC quantification of Khib in WT and TmcA KO E. coli. b, The histograms show experimentally determined relative protein abundance distributions for the samples used to analyze acetylation.
Extended Data Fig. 5 TmcA-mediated Khib specifically related to transcription and translation.
Shown is an interaction network of the TmcA-regulated Khib proteome based on the STRING database (v10). The network is visualized in Cytoscape 3.8.
Extended Data Fig. 6 The MS/MS spectrum of the intracellular tryptic peptide bearing Khib and its counterpart synthetic peptide.
a, MS/MS spectrum of ASDVK(hib)DSSIR (sythesis, intracellular and both mixture) and the extracted ion chromatograms. b, Multiple protein sequences alignment of H-NS from different species.
Extended Data Fig. 7 Volcano plots showing differentially expressed genes in different E. coli MG1655 strains.
WT: wild type E. coli MG1655, O: H-NS KO E. coli MG1655, K: H-NS KO strain harboring H-NS, Q: H-NS KO strain harboring H-NS K121Q, R: H-NS KO strain harboring H-NS K121R. FDR ANOVA < 0.01, n = 3 biological repetitions.
Extended Data Fig. 8 Heatmap of acid resistance related gene expression values of different H-NS related strains.
Rows show Z scores calculated for each group. Wt: wild type E. coli MG1655, O: H-NS KO E. coli MG1655, K: H-NS KO strain harboring H-NS, Q: H-NS KO strain harboring H-NS K121Q, R: H-NS KO strain harboring H-NS K121R.
Extended Data Fig. 9 Acid resistance related genes were increased in Q strain compared with K strain.
a, AR2, AR3 and AR4 are the different mechanisms for acid resistance in E. coli, red numbers represent the difference multiple of gene expression compared Q versus K. The red numbers represent the multiple of gene up regulation. b-e, RT-qPCR detecting mRNA levels of acid resistance related genes in WT, TmcA KO strain, overexpressing TmcA strain and overexpressing TmcA R502A strain, and each group had three biological repetitions and each biological sample had three repetitions. (data are presented as means ± SEM, Two-tailed t-test, n = 3 biological repetitions).
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Tables 1, 2, 5 and 6.
Supplementary Tables
Supplementary Table 3. Quantification results for 2-hydroisobutyrylated proteomes with TmcA KO E. coli and WT E. coli. Supplementary Table 4. Quantification results for acetylated proteomes with TmcA KO E. coli and WT E. coli.
Source data
Source Data Fig. 4
Unprocessed immunoblots and gels for Fig. 4.
Source Data Fig. 5
Unprocessed immunoblots and gels for Fig. 5.
Rights and permissions
About this article
Cite this article
Dong, H., Zhao, Y., Bi, C. et al. TmcA functions as a lysine 2-hydroxyisobutyryltransferase to regulate transcription. Nat Chem Biol 18, 142–151 (2022). https://doi.org/10.1038/s41589-021-00906-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00906-3
This article is cited by
-
Spatiotemporal and direct capturing global substrates of lysine-modifying enzymes in living cells
Nature Communications (2024)
-
Proteome-wide analysis of lysine 2-hydroxyisobutyrylation in Frankliniella occidentalis
BMC Genomics (2022)