Abstract
Extrachromosomal DNA (ecDNA) amplification has been observed in at least 30 different cancer types and is associated with worse patient outcomes. This has been linked to increased oncogene dosage because both oncogenes and associated enhancers can occupy ecDNA. New data challenge the view that only oncogene dosage is affected by ecDNA, and raises the possibility that ecDNA could disrupt genome-wide gene expression. Recent investigations suggest that ecDNA localizes to specialized nuclear bodies (hubs) in which they can act in trans as ectopic enhancers for genes on other ecDNA or chromosomes. Moreover, ecDNA can reintegrate into the genome, possibly further disrupting the gene regulatory landscape in tumor cells. In this Perspective, we discuss the emerging properties of ecDNA and highlight promising avenues to exploit this new knowledge for the development of ecDNA-directed therapies for cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Nowell, P. C. & Hungerford, D. A. Chromosome studies on normal and leukemic human leukocytes. J. Natl Cancer Inst. 25, 85–109 (1960).
Campbell, P. J. et al. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Mitelman, F. et al. Mitelman database of chromosome aberrations and gene fusions in cancer. https://www.scienceopen.com/document?vid=9169110a-2e31-415d-8e0a-40946decb158 (2014).
Biedler, J. & Spengler, B. Metaphase chromosome anomaly: association with drug resistance and cell-specific products. Science 191, 185–187 (1976).
Mitelman, F. & Levan, G. Clustering of aberrations to specific chromosomes in human neoplasms. II. A survey of 287 neoplasms. Hereditas 82, 167–174 (1976).
Cox, D., Yuncken, C. & Spriggs, A. Minute chromatin bodies in malignant tumours of childhood. Lancet 286, 55–58 (1965).
van Roy, N. et al. Translocation-excision-deletion-amplification mechanism leading to nonsyntenic coamplification of MYC and ATBF1. Genes Chromosomes Cancer 45, 107–117 (2006).
Kohl, N. E. et al. Transposition and amplification of oncogene-related sequences in human neuroblastomas. Cell 35, 359–367 (1983).
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat. Genet. 52, 891–897 (2020).
Sakhdari, A. et al. Homogeneously staining region (hsr) on chromosome 11 is highly specific for KMT2A amplification in acute myeloid leukemia (AML) and myelodysplastic syndrome (MDS). Cancer Genet. 238, 18–22 (2019).
Moreau, L. A. et al. Does MYCN amplification manifested as homogeneously staining regions at diagnosis predict a worse outcome in children with neuroblastoma? A Children’s Oncology Group study. Clin. Cancer Res. 12, 5693–5697 (2006).
Christiansen, J. H. et al. Extrachromosomal DNA (ecDNA) carrying amplified oncogenes as a biomarker for insensitivity to pembrolizumab treatment in gastric cancer patients. J. Clin. Oncol. 38, 3123–3123 (2020).
Ruiz, J. C., Choi, K. H., von Hoff, D. D., Roninson, I. B. & Wahl, G. M. Autonomously replicating episomes contain mdr1 genes in a multidrug-resistant human cell line. Mol. Cell. Biol. 9, 109–115 (1989).
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Xing, J. et al. Progress on the role of extrachromosomal DNA in tumor pathogenesis and evolution. Clin. Genet. 99, 503–512 (2020).
Verhaak, R. G. W., Bafna, V. & Mischel, P. S. Extrachromosomal oncogene amplification in tumour pathogenesis and evolution. Nat. Rev. Cancer 19, 283–288 (2019).
Lange, J. T. et al. Principles of ecDNA random inheritance drive rapid genome change and therapy. Preprint at https://www.biorxiv.org/content/10.1101/2021.06.11.447968v1 (2021).
Wahl, G. M. The importance of circular DNA in mammalian gene amplification. Cancer Res. 49, 1333–1340 (1989).
Bailey, C., Shoura, M. J., Mischel, P. S. & Swanton, C. Extrachromosomal DNA—relieving heredity constraints, accelerating tumour evolution. Ann. Oncol. 31, 884–893 (2020).
Kuttler, F. & Mai, S. Formation of non-random extrachromosomal elements during development, differentiation and oncogenesis. Semin. Cancer Biol. 17, 56–64 (2007).
Carroll, S. M. et al. Double minute chromosomes can be produced from precursors derived from a chromosomal deletion. Mol. Cell. Biol. 8, 1525–1533 (1988).
Oettinger, M. A., Schatz, D., Gorka, C. & Baltimore, D. RAG-1 and RAG-2, adjacent genes that synergistically activate V(D)J recombination. Science 248, 1517–1523 (1990).
Tonegawa, S. Somatic generation of antibody diversity. Nature 302, 575–581 (1983).
Paulsen, T., Malapati, P., Eki, R., Abbas, T. & Dutta, A. EccDNA formation is dependent on MMEJ, repressed by c-NHEJ pathway, and stimulated by DNA double-strand break. Preprint at https://www.biorxiv.org/content/10.1101/2020.12.03.410480v1 (2020).
Møller, H. D., Parsons, L., Jørgensen, T. S., Botstein, D. & Regenberg, B. Extrachromosomal circular DNA is common in yeast. Proc. Natl Acad. Sci. USA 112, 3114–3122 (2015).
Storlazzi, C. T. et al. Gene amplification as doubleminutes or homogeneously staining regions in solid tumors: origin and structure. Genome Res. 20, 1198–1206 (2010).
Storlazzi, C. T. et al. MYC-containing double minutes in hematologic malignancies: evidence in favor of the episome model and exclusion of MYC as the target gene. Hum. Mol. Genet. 15, 933–942 (2006).
Gibaud, A. et al. Extrachromosomal amplification mechanisms in a glioma with amplified sequences from multiple chromosome loci. Hum. Mol. Genet. 19, 1276–1285 (2010).
Shoshani, O. et al. Chromothripsis drives the evolution of gene amplification in cancer. Nature 591, 137–141 (2020).
Ly, P. & Cleveland, D. W. Rebuilding chromosomes after catastrophe: emerging mechanisms of chromothripsis. Trends Cell Biol. 27, 917–930 (2017).
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013).
Zhao, X.-K. et al. Extrachromosomal DNA is associated with chromothripsis events and diverse prognoses in gastric cardia adenocarcinoma. Preprint at https://www.biorxiv.org/content/10.1101/2021.07.02.450861v1 (2021).
Ly, P. et al. Selective Y centromere inactivation triggers chromosome shattering in micronuclei and repair by non-homologous end joining. Nat. Cell Biol. 19, 68–75 (2017).
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
L’abbate, A. et al. Genomic organization and evolution of double minutes/homogeneously staining regions with MYC amplification in human cancer. Nucleic Acids Res. 42, 9131–9145 (2014).
Liao, Z. et al. Classification of extrachromosomal circular DNA with a focus on the role of extrachromosomal DNA (ecDNA) in tumor heterogeneity and progression. Biochim. Biophys. Acta Rev. Cancer 1874, 188392 (2020).
Tanaka, H. & Watanabe, T. Mechanisms underlying recurrent genomic amplification in human cancers. Trends Cancer 6, 462–477 (2020).
Coquelle, A., Rozier, L., Dutrillaux, B. & Debatisse, M. Induction of multiple double-strand breaks within an hsr by meganucleaseI-SceI expression or fragile site activation leads to formation of double minutes and other chromosomal rearrangements. Oncogene 21, 7671–7679 (2002).
Shimizu, N., Hashizume, T., Shingaki, K. & Kawamoto, J.-K. Amplification of plasmids containing a mammalian replication initiation region is mediated by controllable conflict between replication and transcription. Cancer Res. 63, 5281–5290 (2003).
Shimizu, N. Extrachromosomal double minutes and chromosomal homogeneously staining regions as probes for chromosome research. Cytogenet. Genome Res. 124, 312–326 (2009).
Oobatake, Y. & Shimizu, N. Double-strand breakage in the extrachromosomal double minutes triggers their aggregation in the nucleus, micronucleation, and morphological transformation. Genes Chromosomes Cancer 59, 133–143 (2020).
Koche, R. P. et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat. Genet. 52, 29–34 (2019).
Shimizu, N., Shingaki, K., Kaneko-Sasaguri, Y., Hashizume, T. & Kanda, T. When, where and how the bridge breaks: anaphase bridge breakage plays a crucial role in gene amplification and HSR generation. Exp. Cell. Res. 302, 233–243 (2005).
McClintock, B. The stability of broken ends of chromosomes in Zea mays. Genetics 26, 234–282 (1941).
Windle, B. E. & Wahl, G. M. Molecular dissection of mammalian gene amplification: new mechanistic insights revealed by analyses of very early events. Mutat. Res. 276, 99–224 (1992).
Ruiz, J. C. & Wahl, G. M. Chromosomal destabilization during gene amplification. Mol. Cell. Biol. 10, 3056–3066 (1990).
Helmsauer, K. et al. Enhancer hijacking determines extrachromosomal circular MYCN amplicon architecture in neuroblastoma. Nat. Commun. 11, 5823 (2020).
Morton, A. R. et al. Functional enhancers shape extrachromosomal oncogene amplifications. Cell 179, 1330–1341 (2019).
Gilbert, F., Balaban, G., Brangman, D., Herrmann, N. & Lister, A. Homogeneously staining regions and tumorigenicity. Int. J. Cancer 31, 765–768 (1983).
Garcillán-Barcia, M. P., Alvarado, A. & de la Cruz, F. Identification of bacterial plasmids based on mobility and plasmid population biology. FEMS Microbiol. Rev. 35, 936–956 (2011).
Shimizu, N., Itoh, N., Utiyama, H. & Wahl, G. M. Selective entrapment of extrachromosomally amplified DNA by nuclear budding and micronucleation during S phase. J. Cell Biol. 140, 1307–1320 (1998).
Ambros, I. M. et al. Neuroblastoma cells can actively eliminate supernumerary MYCN gene copies by micronucleus formation—sign of tumour cell revertance? Eur. J. Cancer 33, 2043–2049 (1997).
von Hoff, D. D. et al. Elimination of extrachromosomally amplified MYC genes from human tumor cells reduces their tumorigenicity. Proc. Natl Acad. Sci. USA 89, 8165–8169 (1992).
Itoh, N. & Shimizu, N. DNA replication-dependent intranuclear relocation of double minute chromatin. J. Cell Sci. 111, 3275–3285 (1998).
Shimizu, N., Ochi, T. & Itonaga, K. Replication timing of amplified genetic regions relates to intranuclear localization but not to genetic activity or G/R band. Exp. Cell. Res. 268, 201–210 (2001).
Leibowitz, M. L. et al. Chromothripsis as an on-target consequence of CRISPR–Cas9 genome editing. Nat. Genet. 53, 895–905 (2021).
Nevaldine, B. H., Rizwana, R. & Hahn, P. J. Differential sensitivity of double minute chromosomes to hydroxyurea treatment in cultured methotrexate-resistant mouse cells. Mutat. Res. Genomics 406, 55–62 (1999).
Prochazka, P., Hrabeta, J., Vícha, A. & Eckschlager, T. Expulsion of amplified MYCN from homogenously staining chromosomal regions in neuroblastoma cell lines after cultivation with cisplatin, doxorubicin, hydroxyurea, and vincristine. Cancer Genet. Cytogenet. 196, 96–104 (2010).
Narath, R. et al. Induction of senescence in MYCN amplified neuroblastoma cell lines by hydroxyurea. Genes Chromosomes Cancer 46, 130–142 (2007).
Shimizu, N. et al. Loss of amplified c-myc genes in the spontaneously differentiated HL-60 cells. Cancer Res. 54, 3561–3567 (1994).
Eckhardt, S. G. et al. Induction of differentiation in HL60 cells by the reduction of extrachromosomally amplified c-myc. Proc. Natl Acad. Sci. USA 91, 6674–6678 (1994).
Wrighton, K. W. Genome organization: zooming in on nuclear organization. Nat. Rev. Genet. 18, 269 (2017).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Solovei, I. et al. Topology of double minutes (dmins) and homogeneously staining regions (HSRs) in nuclei of Human neuroblastoma cell lines. Genes Chromosomes Cancer 46, 130–140 (2000).
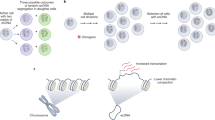
Zhu, Y. et al. Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription. Cancer Cell 39, 694–707 (2021).
Hung, K. L. et al. EcDNA hubs drive cooperative intermolecular oncogene expression. Nature 600, 731–736 (2021).
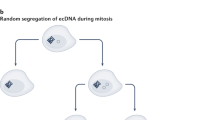
Yi, E. et al. Live-cell imaging shows uneven segregation of extrachromosomal DNA elements and transcriptionally active extrachromosomal DNA clusters in cancer. Cancer Discov. 12, 1–16 (2021).
Kanda, T. & Wahl, G. M. The dynamics of acentric chromosomes in cancer cells revealed by GFP-based chromosome labeling strategies. J. Cell. Biochem. Suppl. 35, 107–114 (2000).
Kanda, T., Otter, M. & Wahl, G. Mitotic segregation of viral and cellular acentric extrachromosomal molecules by chromosome tethering. J. Cell Sci. 114, 49–58 (2001).
Choi, L. M. et al. Telomerase activity by TRAP assay and telomerase RNA (hTR) expression are predictive of outcome in neuroblastoma. Med. Pediatr. Oncol. 35, 647–650 (2000).
Adelman, K. & Martin, B. J. E. ecDNA party bus: bringing the enhancer to you. Mol. Cell 81, 1866–1867 (2021).
Nathanson, D. A. et al. Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science 343, 72–76 (2014).
Decarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Schulte, A. et al. Glioblastoma stem-like cell lines with either maintenance or loss of high-level EGFR amplification, generated via modulation of ligand concentration. Clin. Cancer Res. 18, 1901–1913 (2012).
Song, K. et al. Abstract 2036: plasticity and vulnerability associated with extrachromosomal and intrachromosomal BRAF amplifications. Cancer Res. 81, 2036–2036 (2021).
Raymond, E. et al. Effects of hydroxyurea on extrachromosomal DNA in patients with advanced ovarian carcinomas. Clin. Cancer Res. 7, 1171–1180 (2001).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Winter, G. E. et al. BET bromodomain proteins function as master transcription elongation factors independent of CDK9 recruitment. Mol. Cell 67, 5–19 (2017).
Henssen, A. et al. Targeting MYCN-driven transcription by BET-bromodomain inhibition. Clin. Cancer Res. 22, 2470–2481 (2016).
Acknowledgements
We apologize to colleagues whose work was not discussed or cited due to space constraints. We thank G. Wahl, K. Astrahantseff, J. Glaser, K. Helmsauer and R. Koche for fruitful discussions. A.G.H. is supported by Deutsche Forschungsgemeinschaft (no. 398299703) and the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant agreement no. 949172).
Author information
Authors and Affiliations
Contributions
A.G.H wrote the manuscript. E.V.L. designed the figures and contributed to the manuscript text. L.B. contributed to the text and the reference list.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Peter Scacheri and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
van Leen, E., Brückner, L. & Henssen, A.G. The genomic and spatial mobility of extrachromosomal DNA and its implications for cancer therapy. Nat Genet 54, 107–114 (2022). https://doi.org/10.1038/s41588-021-01000-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-01000-z
This article is cited by
-
Scrambling the genome in cancer: causes and consequences of complex chromosome rearrangements
Nature Reviews Genetics (2024)
-
Genetic determinants of micronucleus formation in vivo
Nature (2024)
-
Oncogenic structural aberration landscape in gastric cancer genomes
Nature Communications (2023)
-
MGA-seq: robust identification of extrachromosomal DNA and genetic variants using multiple genetic abnormality sequencing
Genome Biology (2023)
-
Parallel sequencing of extrachromosomal circular DNAs and transcriptomes in single cancer cells
Nature Genetics (2023)